Abstract
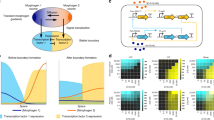
Spontaneous pattern formation in Turing systems relies on feedback. But patterns in cells and tissues seldom form spontaneously—instead they are controlled by regulatory biochemical interactions that provide molecular guiding cues. The relationship between these guiding cues and feedback in controlled biological pattern formation remains unclear. Here, we explore this relationship during cell-polarity establishment in the one-cell-stage Caenorhabditis elegans embryo. We quantify the strength of two feedback systems that operate during polarity establishment: feedback between polarity proteins and the actomyosin cortex, and mutual antagonism among polarity proteins. We characterize how these feedback systems are modulated by guiding cues from the centrosome, an organelle regulating cell cycle progression. By coupling a mass-conserved Turing-like reaction–diffusion system for polarity proteins to an active-gel description of the actomyosin cortex, we reveal a transition point beyond which feedback ensures self-organized polarization, even when cues are removed. Notably, the system switches from a guide-dominated to a feedback-dominated regime well beyond this transition point, which ensures robustness. Together, these results reveal a general criterion for controlling biological pattern-forming systems: feedback remains subcritical to avoid unstable behaviour, and molecular guiding cues drive the system beyond a transition point for pattern formation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data generated or analysed in this study are available from the corresponding author upon request.
References
Motegi, F. & Seydoux, G. The PAR network: redundancy and robustness in a symmetry-breaking system. Philos. Trans. Royal Soc. B 368, 20130010 (2013).
Hoege, C. & Hyman, A. A. Principles of PAR polarity in Caenorhabditis elegans embryos. Nat. Rev. Mol. Cell Biol. 14, 315–322 (2013).
Lang, C. F. & Munro, E. The PAR proteins: from molecular circuits to dynamic self-stabilizing cell polarity. Development 144, 3405–3416 (2017).
Goldstein, B. & Macara, I. G. The PAR proteins: fundamental players in animal cell polarization. Dev. Cell. 13, 609–622 (2007).
Motegi, F. et al. Microtubules induce self-organization of polarized PAR domains in Caenorhabditis elegans zygotes. Nat. Cell Biol. 13, 1361–1367 (2011).
Goehring, N. W. et al. Polarization of PAR proteins by advective triggering of a pattern-forming system. Science 334, 1137–1141 (2011).
Trong, P. K., Nicola, E. M., Goehring, N. W., Kumar, K. V. & Grill, S. W. Parameter-space topology of models for cell polarity. New J. Phys. 16, 065009 (2014).
Munro, E., Nance, J. & Priess, J. R. Cortical flows powered by asymmetrical contraction transport PAR proteins to establish and maintain anterior-posterior polarity in the early C. elegans embryo. Dev. Cell. 7, 413–424 (2004).
Wang, S.-C. et al. Cortical forces and CDC-42 control clustering of PAR proteins for Caenorhabditis elegans embryonic polarization. Nat. Cell Biol. 19, 988–995 (2017).
Rodriguez, J. et al. aPKC cycles between functionally distinct PAR protein assemblies to drive cell polarity. Dev. Cell. 42, 400–415 (2017).
Dickinson, D. J., Schwager, F., Pintard, L., Gotta, M. & Goldstein, B. A single-cell biochemistry approach reveals PAR complex dynamics during cell polarization. Dev. Cell. 42, 416–434 (2017).
Schneider, S. Q. & Bowerman, B. Cell polarity and the cytoskeleton in the Caenorhabditis elegans zygote. Annu. Rev. Genet. 37, 221–249 (2003).
Cheeks, R. J. et al. C. elegans PAR proteins function by mobilizing and stabilizing asymmetrically localized protein complexes. Curr. Biol. 14, 851–862 (2004).
Beatty, A., Morton, D. G. & Kemphues, K. PAR-2, LGL-1 and the CDC-42 GAP CHIN-1 act in distinct pathways to maintain polarity in the C. elegans embryo. Development 140, 2005–2014 (2013).
Zonies, S., Motegi, F., Hao, Y. & Seydoux, G. Symmetry breaking and polarization of the C. elegans zygote by the polarity protein PAR-2. Development 137, 1669–1677 (2010).
Nance, J. & Zallen, J. A. Elaborating polarity: PAR proteins and the cytoskeleton. Development 138, 799–809 (2011).
Gross, P., Kumar, K. V. & Grill, S. W. How active mechanics and regulatory biochemistry combine to form patterns in development. Annu. Rev. Biophys. 46, 337–356 (2017).
Tse, Y. C. et al. RhoA activation during polarization and cytokinesis of the early Caenorhabditis elegans embryo is differentially dependent on NOP-1 and CYK-4. Mol. Biol. Cell. 23, 4020–4031 (2012).
Motegi, F. & Sugimoto, A. Sequential functioning of the ECT-2 RhoGEF, RHO-1 and CDC-42 establishes cell polarity in Caenorhabditis elegans embryos. Nat. Cell Biol. 8, 978–985 (2006).
Mayer, M., Depken, M., Bois, J. S., Jülicher, F. & Grill, S. W. Anisotropies in cortical tension reveal the physical basis of polarizing cortical flows. Nature 467, 617–621 (2010).
Sailer, A., Anneken, A., Li, Y., Lee, S. & Munro, E. Dynamic opposition of clustered proteins stabilizes cortical polarity in the C. elegans zygote. Dev. Cell. 35, 131–142 (2015).
Goehring, N. W., Chowdhury, D., Hyman, A. A. & Grill, S. W. FRAP analysis of membrane-associated proteins: lateral diffusion and membrane-cytoplasmic exchange. Biophys. J. 99, 2443–2452 (2010).
Goehring, N. W., Hoege, C., Grill, S. W. & Hyman, A. A. PAR proteins diffuse freely across the anterior-posterior boundary in polarized C. elegans embryos. J. Cell Biol. 193, 583–594 (2011).
Nishikawa, M., Naganathan, S. R., Jülicher, F. & Grill, S. W. Controlling contractile instabilities in the actomyosin cortex. eLife 6, e19595 (2017).
Wu, J. Q. & Pollard, T. D. Counting cytokinesis proteins globally and locally in fission yeast. Science 310, 310–314 (2005).
David, D. J. V., Tishkina, A. & Harris, T. J. C. The PAR complex regulates pulsed actomyosin contractions during amnioserosa apical constriction in Drosophila. Development 137, 1645–1655 (2010).
Turing, A. M. The chemical basis of morphogenesis. Philos. Trans. Royal Soc. B 237, 37–72 (1952).
Kondo, S. & Miura, T. Reaction-diffusion model as a framework for understanding biological pattern formation. Science 329, 1616–1620 (2010).
Marcon, L. & Sharpe, J. Turing patterns in development: what about the horse part? Curr. Opin. Genet. Dev. 22, 578–584 (2012).
Mikhailov, A. S. & Showalter, K. Control of waves, patterns and turbulence in chemical systems. Phys. Rep. 425, 79–194 (2006).
Prokopenko, M. Guided self-organization. HFSP J. 3, 287–289 (2009).
Otsuji, M. et al. A mass conserved reaction-diffusion system captures properties of cell polarity. PLoS Comput. Biol. 3, 1040–1054 (2007).
Jilkine, A. & Edelstein-Keshet, L. A comparison of mathematical models for polarization of single eukaryotic cells in response to guided cues. PLoS Comput. Biol. 7, e1001121 (2011).
Wu, F. et al. Multistability and dynamic transitions of intracellular Min protein patterns. Mol. Syst. Biol. 12, 873 (2016).
Raspopovic, J., Marcon, L., Russo, L. & Sharpe, J. Digit patterning is controlled by a bmp-sox9-wnt Turing network modulated by morphogen gradients. Science 345, 566–570 (2014).
Corson, F., Couturier, L., Rouault, H., Mazouni, K. & Schweisguth, F. Self-organized notch dynamics generate stereotyped sensory organ patterns in Drosophila. Science 356, 501–508 (2017).
Zagorski, M. et al. Decoding of position in the developing neural tube from antiparallel morphogen gradients. Science 356, 1379–1383 (2017).
Lee, S. S. & Shibata, T. Self-organization and advective transport in the cell polarity formation for asymmetric cell division. J. Theor. Biol. 382, 1–14 (2015).
Halatek, J. & Frey, E. Rethinking pattern formation in reaction–diffusion systems. Nat. Phys. 14, 507–514 (2018).
Bois, J. S., Jülicher, F. & Grill, S. W. Pattern formation in active fluids. Phys. Rev. Lett. 106, 028103 (2011).
Arata, Y. et al. Cortical polarity of the RING protein PAR-2 is maintained by exchange rate kinetics at the cortical-cytoplasmic boundary. Cell Reports 16, 2156–2168 (2016).
Saha, A. et al. Determining physical properties of the cell cortex. Biophys. J. 110, 1421–1429 (2016).
Robin, F. B., McFadden, W. M., Yao, B. & Munro, E. M. Single-molecule analysis of cell surface dynamics in Caenorhabditis elegans embryos. Nat. Methods 11, 677–682 (2014).
Munro, E. & Bowerman, B. Cellular symmetry breaking during Caenorhabditis elegans development. Cold Spring Harb. Perspect. Biol. 1, a003400 (2009).
Cowan, C. R. & Hyman, A. A. Centrosomes direct cell polarity independently of microtubule assembly in C. elegans embryos. Nature 431, 92–96 (2004).
Kamath, R. S. & Ahringer, J. Genome-wide RNAi screening in Caenorhabditis elegans. Methods 30, 313–321 (2003).
Sedzinski, J. et al. Polar actomyosin contractility destabilizes the position of the cytokinetic furrow. Nature 476, 462–466 (2011).
Blanchoud, S., Busso, C., Naef, F. & Goenczy, P. Quantitative analysis and modeling probe polarity establishment in C. elegans embryos. Biophys. J. 108, 799–809 (2015).
Ruettinger, S. et al. Comparison and accuracy of methods to determine the confocal volume for quantitative fluorescence correlation spectroscopy. J. Microsc. 232, 343–352 (2008).
Jaqaman, K. et al. Robust single-particle tracking in live-cell time-lapse sequences. Nat. Methods 5, 695–702 (2008).
Thielicke, W. & Stamhuis, E. J. PIVlab - towards user-friendly, affordable and accurate digital particle imagevelocimetry in MATLAB. J. Open Res. Softw. 2, e30 (2014).
Acknowledgements
P.G. acknowledges a EMBO Long-Term Fellowship for funding. The research of K.V.K. is supported by the Department of Biotechnology, India through a Ramalingaswami Re-entry Fellowship, and the Max Planck Society and the Department of Science and Technology, India through a Max Planck Partner Group at ICTS-TIFR. N.W.G. was supported by the Francis Crick Institute, which receives its core funding from Cancer Research UK (FC001086), the UK Medical Research Council (FC001086) and the Wellcome Trust (FC001086), and is a member of the GENiE network supported by COST Action BM1408 and EMBO. S.W.G was supported by the DFG (SPP 1782, GSC 97, GR 3271/2, GR 3271/3, GR 3271/4), the European Research Council (grants 281903 and 742712), ITN grants 281903 and 641639 from the EU, the Max-Planck-Society as a Max-Planck-Fellow, and the Human Frontier Science Program (RGP0023/2014). J.S.B. acknowledges the Human Frontier Science Program for funding. We thank D. Dickinson, B. Goldstein, F. Motegi and G. Seydoux for sharing C. elegans strains. We thank P. Gönczy, L. Hubatsch, T. Hyman, K. Kruse and M. Labouesse for discussion and insightful comments on the manuscript.
Author information
Authors and Affiliations
Contributions
P.G. performed experiments and K.V.K. developed the theory, with help from all authors. Data were analysed together with input from all authors. P.G., K.V.K., F.J. and S.W.G. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Discussion, Supplementary Equations, Supplementary Figures 1–15, Supplementary Tables 1–4, Supplementary References 1–27
Supplementary Video 1
Concentration field of PAR-2::GFP (blue, N = 8) and PAR-6::mCherry (red, N = 8), during mlc-4 RNAi, over time. Error bands represent the standard error of the mean. The solid line shows the best fit, using equations as described in Supplementary Table 1, with parameters shown in Supplementary Tables 2 and 3.
Supplementary Video 2
Ensemble-averaged concentration field of NMY-2::GFP (grey, N = 8) and the ensemble-averaged NMY-2 flow field (green, N = 10), during par-2 and par-6 double RNAi, over time. Error bands represent the standard error of the mean. The solid line shows the best fit, using equations as described in Supplementary Table 1, with parameters shown in Supplementary Tables 2 and 3.
Supplementary Video 3
Ensemble-averaged concentration field of PAR-2-MT-::GFP (blue, N = 9) and PAR-6::mCherry (red, N = 9) NMY-2::mKate2 (grey, N = 6) and the ensemble-averaged NMY-2 flow field (green, N = 9) for the PAR-2 MT- condition, over time. Error bands represent the standard error of the mean. The solid line shows the model prediction, using equations as described in Supplementary Table 1, with parameters shown in Supplementary Tables 2 and 3.
Supplementary Video 4
Ensemble-averaged concentration field of PAR-2::GFP (blue, N = 6) and PAR-6::mCherry (red, N = 6) NMY-2::GFP (grey, N = 8) and the ensemble-averaged NMY-2 flow field (green, N = 12) for the unperturbed condition, over time. Error bands represent the standard error of the mean. The solid line shows the model prediction, using equations as described in Supplementary Table 1, with parameters shown in Supplementary Tables 2 and 3.
Supplementary Video 5
Ensemble-averaged concentration field of PAR-2::GFP (blue, N = 6) and PAR-6::mCherry (red, N = 6) NMY-2::GFP (grey, N = 8) and the ensemble-averaged NMY-2 flow field (green, N = 12) for the unperturbed condition, over time. Error bands represent the standard error of the mean. The solid line shows a fit to the model, using equations as described in Supplementary Table 1, as shown in Supplementary Fig. 9
Rights and permissions
About this article
Cite this article
Gross, P., Kumar, K.V., Goehring, N.W. et al. Guiding self-organized pattern formation in cell polarity establishment. Nat. Phys. 15, 293–300 (2019). https://doi.org/10.1038/s41567-018-0358-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41567-018-0358-7
This article is cited by
-
Self-organized intracellular twisters
Nature Physics (2024)
-
Friction forces determine cytoplasmic reorganization and shape changes of ascidian oocytes upon fertilization
Nature Physics (2024)
-
Directing Min protein patterns with advective bulk flow
Nature Communications (2023)
-
Control of protein-based pattern formation via guiding cues
Nature Reviews Physics (2022)
-
Forced and spontaneous symmetry breaking in cell polarization
Nature Computational Science (2022)