Abstract
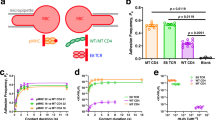
How the engagement of a T-cell receptor to antigenic peptide-loaded major histocompatibility complex on antigen-presenting cells (APCs) initiates intracellular signalling cascades in T cells is not well understood. In particular, the dimension of the cellular contact zone is regarded as a determinant, but its influence remains controversial. This is due to the need for appropriate strategies for manipulating intermembrane spacing between the APC–T-cell interfaces without involving protein modification. Here we describe a membrane-anchored DNA nanojunction with distinct sizes to extend, maintain and shorten the APC–T-cell interface down to 10 nm. Our results suggest that the axial distance of the contact zone is critical in T-cell activation, presumably by modulating protein reorganization and mechanical force. Notably, we observe the promotion of T-cell signalling by shortening the intermembrane distance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the article and its Supplementary Information. Source data are provided with this paper and available via Zenodo at https://doi.org/10.5281/zenodo.7424802.
Code availability
The code designed for data collection and analysis of this study is available from the corresponding authors upon reasonable request.
Change history
19 September 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41565-023-01505-0
References
Weiss, A. & Dan, R. L. Signal transduction by lymphocyte antigen receptors. Cell 76, 263–274 (1994).
Brownlie, R. J. & Zamoyska, R. T cell receptor signalling networks: branched, diversified and bounded. Nat. Rev. Immunol. 13, 257–269 (2013).
Chakraborty, A. K. & Weiss, A. Insights into the initiation of TCR signaling. Nat. Immunol. 15, 798–807 (2014).
Xu, X., Li, H. & Xu, C. Structural understanding of T cell receptor triggering. Cell. Mol. Immunol. 17, 193–202 (2020).
Schamel, W. W., Alarcon, B. & Minguet, S. The TCR is an allosterically regulated macromolecular machinery changing its conformation while working. Immunol. Rev. 291, 8–25 (2019).
Lee, M. S. et al. A mechanical switch couples T cell receptor triggering to the cytoplasmic juxtamembrane regions of CD3ζζ. Immunity 43, 227–239 (2015).
Feng, Y., Reinherz, E. L. & Lang, M. J. αβ T cell receptor mechanosensing forces out serial engagement. Trends Immunol. 39, 596–609 (2018).
Mckeithan, T. W. Kinetic proofreading in T-cell receptor signal transduction. Proc. Natl Acad. Sci. USA 92, 5042–5046 (1992).
Rabinowitz, J. D., Beeson, C., Lyons, D. S. & Mcconnell, D. H. M. Kinetic discrimination in T-cell activation. Proc. Natl Acad. Sci. USA 93, 1401–1405 (1996).
Huang, J. et al. The kinetics of two-dimensional TCR and pMHC interactions determine T-cell responsiveness. Nature 464, 932–936 (2010).
Huppa, J. B. et al. TCR–peptide–MHC interactions in situ show accelerated kinetics and increased affinity. Nature 463, 963–967 (2010).
Springer, T. A. Adhesion receptors of the immune system. Nature 346, 425–434 (1990).
Choudhuri, K., Wiseman, D., Brown, M. H., Gould, K. G. & Der Merwe, P. A. V. T-cell receptor triggering is critically dependent on the dimensions of its peptide-MHC ligand. Nature 436, 578–582 (2005).
Cai, H. et al. Full control of ligand positioning reveals spatial thresholds for T-cell receptor triggering. Nat. Nanotechnol. 13, 610–617 (2018).
Sun, L. et al. DNA-edited ligand positioning on red blood cells to enable optimized T-cell activation for adoptive immunotherapy. Angew. Chem. Int. Ed. 59, 14842–14853 (2020).
Garcia, K. C. et al. An alphabeta T-cell receptor structure at 2.5 Å and its orientation in the TCR–MHC complex. Science 274, 209–219 (1996).
Birnbaum, M. E. et al. Deconstructing the peptide–MHC specificity of T-cell recognition. Cell 157, 1073–1087 (2014).
McCall, M. N., Shotton, D. M. & Barclay, A. N. Expression of soluble isoforms of rat CD45. Analysis by electron microscopy and use in epitope mapping of anti-CD45R monoclonal antibodies. Immunology 76, 310–317 (1992).
Carbone, C. B. et al. In vitro reconstitution of T-cell receptor-mediated segregation of the CD45 phosphatase. Proc. Natl Acad. Sci. USA 114, E9338–E9345 (2017).
Chang, V. T. et al. Initiation of T cell signaling by CD45 segregation at ‘close contacts’. Nat. Immunol. 17, 574–582 (2016).
Hermiston, M. L., Xu, Z. & Weiss, A. CD45: a critical regulator of signaling thresholds in immune cells. Annu. Rev. Immunol. 21, 107–137 (2003).
Davis, S. J. & van der Merwe, P. A. The kinetic-segregation model: TCR triggering and beyond. Nat. Immunol. 7, 803–809 (2006).
Razvag, Y., Neve-Oz, Y., Sajman, J., Reches, M. & Sherman, E. Nanoscale kinetic segregation of TCR and CD45 in engaged microvilli facilitates early T cell activation. Nat. Commun. 9, 732 (2018).
Li, Y.-C. et al. Cutting edge: mechanical forces acting on T cells immobilized via the TCR complex can trigger TCR signaling. J. Immunol. 184, 5959–5963 (2010).
James, J. R. & Vale, R. D. Biophysical mechanism of T-cell receptor triggering in a reconstituted system. Nature 487, 64–69 (2012).
Irles, C. et al. CD45 ectodomain controls interaction with GEMs and Lck activity for optimal TCR signaling. Nat. Immunol. 4, 189–197 (2003).
Chen, B. et al. The affinity of elongated membrane-tethered ligands determines potency of T cell receptor triggering. Front. Immunol. 8, 793 (2017).
Choudhuri, K. & van der Merwe, P. A. Molecular mechanisms involved in T cell receptor triggering. Semin. Immunol. 19, 255–261 (2007).
van der Merwe, P. A. & Dushek, O. Mechanisms for T cell receptor triggering. Nat. Rev. Immunol. 11, 47–55 (2011).
Malissen, B. & Bongrand, P. Early T cell activation: integrating biochemical, structural, and biophysical cues. Annu. Rev. Immunol. 33, 539–561 (2015).
Courtney, A. H., Lo, W.-L. & Weiss, A. TCR signaling: mechanisms of initiation and propagation. Trends Biochem. Sci. 43, 108–123 (2018).
Goodman, R. P. et al. Rapid chiral assembly of rigid DNA building blocks for molecular nanofabrication. Science 310, 1661–1665 (2005).
Lin, M. et al. Programmable engineering of a biosensing interface with tetrahedral DNA nanostructures for ultrasensitive DNA detection. Angew. Chem. Int. Ed. 54, 2151–2155 (2015).
Li, J. et al. Cell-membrane-anchored DNA nanoplatform for programming cellular interactions. J. Am. Chem. Soc. 141, 18013–18020 (2019).
Jung, Y. et al. Three-dimensional localization of T-cell receptors in relation to microvilli using a combination of superresolution microscopies. Proc. Natl Acad. Sci. USA 113, E5916–E5924 (2016).
Yi, J. C. & Samelson, L. E. Microvilli set the stage for T-cell activation. Proc. Natl Acad. Sci. USA 113, 11061–11062 (2016).
Du, Y. et al. Ligand dilution analysis facilitates aptamer binding characterization at the single-molecule level. Angew. Chem. Int. Ed. 7, e202215387 (2022).
Huse, M. et al. Spatial and temporal dynamics of T cell receptor signaling with a photoactivatable agonist. Immunity 27, 76–88 (2007).
Armstrong, J. K., Wenby, R. B., Meiselman, H. J. & Fisher, T. C. The hydrodynamic radii of macromolecules and their effect on red blood cell aggregation. Biophys. J. 87, 4259–4270 (2004).
Zehn, D., Lee, S. Y. & Bevan, M. J. Complete but curtailed T-cell response to very low-affinity antigen. Nature 458, 211–214 (2009).
Law, C. C. et al. Expression and characterization of recombinant soluble human CD3 molecules: presentation of antigenic epitopes defined on the native TCR–CD3 complex. Int. Immunol. 14, 389–400 (2002).
Cohen, S. & Milstein, C. Structure of antibody molecules. Nature 214, 449–452 (1967).
Mosayebi, M., Louis, A. A., Doye, J. P. K. & Ouldridge, T. E. Force-induced rupture of a DNA duplex: from fundamentals to force sensors. ACS Nano 9, 11993–12003 (2015).
Furukawa, T., Itoh, M., Krueger, N. X., Streuli, M. & Saito, H. Specific interaction of the CD45 protein-tyrosine phosphatase with tyrosine-phosphorylated CD3 zeta chain. Proc. Natl Acad. Sci. USA 91, 10928–10932 (1994).
Hegedus, Z. et al. Contribution of kinases and the CD45 phosphatase to the generation of tyrosine phosphorylation patterns in the T cell receptor complex ζ chain. Immunol. Lett. 67, 31–39 (1999).
Straus, D. B. & Weiss, A. The CD3 chains of the T cell antigen receptor associate with the ZAP-70 tyrosine kinase and are tyrosine phosphorylated after receptor stimulation. J. Exp. Med. 178, 1523–1530 (1993).
Chan, A. C., Iwashima, M., Turck, C. W. & Weiss, A. ZAP-70: a 70 kd protein-tyrosine kinase that associates with the TCR ζ chain. Cell 71, 649–662 (1992).
Sherman, E. et al. Functional nanoscale organization of signaling molecules downstream of the T cell antigen receptor. Immunity 35, 705–720 (2011).
Yokosuka, T. et al. Newly generated T cell receptor microclusters initiate and sustain T cell activation by recruitment of Zap70 and SLP-76. Nat. Immunol. 6, 1253–1262 (2005).
Zhang, D. Y. & Seelig, G. Dynamic DNA nanotechnology using strand-displacement reactions. Nat. Chem. 3, 103–113 (2011).
Stone, J. D., Chervin, A. S. & Kranz, D. M. T cell receptor binding affinities and kinetics: impact on T cell activity and specificity. Immunology 126, 165–176 (2009).
Valitutti, S., Müller, S., Cella, M., Padovan, E. & Lanzavecchia, A. Serial triggering of many T-cell receptors by a few peptide–MHC complexes. Nature 375, 148–151 (1995).
Acknowledgements
We thank X. Yu and Q. Zou at Shanghai Jiao Tong University School of Medicine for providing technical assistance. This work is supported by the National Key Research Program (2021YFA0910100 (to L.Q.), 2018YFC1602900 (to L.Q.) and 2019YFA0905800 (to W.T.)), the National Natural Science Foundation of China (NSFC 21922404 (to L.Q.), 22174039 (to L.Q.), 22107027 (to Y.L.) and 21827811 (to W.T.)), the Science and Technology Project of Hunan Province (2021RC4022 (to L.Q.), 2019SK2201 (to W.T.), 2018RS3035 (to L.Q.) and 2017XK2103 (to W.T.)) and Shenzhen excellent science and technology innovation talents training project (RCBS20200714114821377 (to Y.L.)).
Author information
Authors and Affiliations
Contributions
Y.D. and L.Q. conceived and designed the research. Y.D., Y.L., J.L., C.M., Q.Z. and Y.Z. conducted the experiments. Y.D. performed the analysis. Y.D. and Y.L. performed the imaging analysis and reconstruction of cell membrane morphology. Y.D., L.Q. and W.T. drafted and revised the manuscript. All the authors commented on the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Nanotechnology thanks Michael Dustin, Wolfgang Schamel and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–44, Tables 1–4, unprocessed gels for Figs. 2, 15 and 35, Methods, and References.
Source data
Source Data Fig. 1
Statistical source data and unprocessed gels.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Du, Y., Lyu, Y., Lin, J. et al. Membrane-anchored DNA nanojunctions enable closer antigen-presenting cell–T-cell contact in elevated T-cell receptor triggering. Nat. Nanotechnol. 18, 818–827 (2023). https://doi.org/10.1038/s41565-023-01333-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41565-023-01333-2
This article is cited by
-
Assembly of peptide nanostructures with controllable sizes
Nano Research (2024)
-
DNA nanodevices map intracellular ions
Nature Biotechnology (2023)
-
Inhibition of glycolysis-driven immunosuppression with a nano-assembly enhances response to immune checkpoint blockade therapy in triple negative breast cancer
Nature Communications (2023)
-
Precise regulation of intermembrane spacing with DNA nanojunctions for enhanced T-cell receptor triggering
Science China Chemistry (2023)


