Abstract
The strategy of combining a vaccine with immune checkpoint inhibitors has been widely investigated in cancer management, but the complete response rate for this strategy is still unresolved. We describe a genetically engineered cell membrane nanovesicle that integrates antigen self-presentation and immunosuppression reversal (ASPIRE) for cancer immunotherapy. The ASPIRE nanovaccine is derived from recombinant adenovirus-infected dendritic cells in which specific peptide-major histocompatibility complex class I (pMHC-I), anti-PD1 antibody and B7 co-stimulatory molecules are simultaneously anchored by a programmed process. ASPIRE can markedly improve antigen delivery to lymphoid organs and generate broad-spectrum T-cell responses that eliminate established tumours. This work presents a powerful vaccine formula that can directly activate both native T cells and exhausted T cells, and suggests a general strategy for personalized cancer immunotherapy.
Similar content being viewed by others
Main
The successful establishment of antitumour immunity requires antigen-specific lymphocytes to undergo activation, expansion and differentiation1,2. To a large extent, this process is determined by the interaction between T cells and antigen-presenting cells (APCs). Therefore, the control of APC function is a critical step for therapeutic strategies that require T-cell stimulation. The activation of CD8+ cytotoxic T lymphocytes (CTLs) has been proved as a key factor in cancer immunotherapy. This process depends on the tumour-derived peptides that are presented by APCs’ major histocompatibility complex class I (MHC-I) molecules to T cells3,4,5. Conventional vaccines, which include peptide and protein vaccines, rely on random encounters with host APCs, and an inappropriate encounter might lead to the silencing of an immune response6,7,8. The activation of CD8+ T cells is also excessively dependent on an efficient antigen cross-presentation. Both situations could explain some of the shortcomings of current cancer vaccines.
The high manoeuvrability of the activation programme and cell states is the greatest advantage of adoptive cell therapies, such as chimeric antigen receptor T-cell or dendritic cell (DC) vaccine strategies. Especially for DC-based cancer vaccine strategies, it is possible to induce an endogenous antigen-specific CTL response modulated by a typical MHC-I restricted presentation9,10,11,12. However, it is unavoidable that the activated DCs are short-lived in the body after injection, and ultimately only a small portion of the administered DCs migrate to the draining lymph nodes (LNs). This disadvantage greatly limits the activation of CTLs13,14. Despite the progress made in treating haematological malignancies, the use of chimeric antigen receptor T-cell therapy against solid tumours has not been very successful15. Therefore, it remains a daunting task to design a more effective cancer vaccine formula.
Moreover, in the tumour immune microenvironment, immune checkpoints can inhibit antigen-specific CTL responses16. Although immunotherapies based on immune checkpoints have tremendous potential17,18,19,20, only a small fraction of patients experience complete responses, which might be due to the absence of an adequate pre-existing cytotoxic CD8+ T-cell response. Stimulating a strong CD8+ cell response coupled with immune checkpoint therapy would be an optimal solution. The cellular and molecular mechanisms of checkpoint blockade therapy are yet to be fully elucidated. Some recent studies demonstrated a requirement of CD28 co-stimulation for CD8+ T-cell rescue and suggested an important role for the CD28/B7 pathway in PD-1 (programmed cell death protein 1) therapy for cancer patients21,22. Therefore, we hypothesized that a co-delivery system with B7-1 and B7-2 co-stimulatory molecules (which are abundant on the surfaces of mature DCs) and an anti-PD-1 antibody may be a key strategy to overcome the challenges faced in cancer immunotherapy.
Biomimetic synthesis involves the synthesis and display of protein cargos on the cell surface, which is a promising strategy for the efficient production of ligand-oriented nanocarriers23,24,25. Such a biosynthesis process could maintain the native conformation, structure and activity of a functional protein of interest26,27,28. In this study, we report a novel nanovaccine formulation that integrates antigen self-presentation and immunosuppression reversal (ASPIRE) for personalized cancer immunotherapy. The ASPIRE nanovaccine is based on artificial cytomembrane nanovesicles (NVs) derived from DCs that feature the directional presentation of specific antigen epitopes by MHC-I molecules, as well as the co-delivery of anti-PD1 antibody and B7 co-stimulatory molecules via a programmed process. It features nanoscale size, good stability and a homing effect that is mediated by surface adhesion molecules. These features enable the ASPIRE nanovaccine to rapidly enrich in the lymphatic system. Unlike conventional vaccines requiring to be delivered to APCs, the ASPIRE nanovaccine has the ability to present neoantigens to CD8+ T cells directly, which we call ‘antigen self-presentation’, and thus stimulate strong CTL responses. Besides, it also acts as a new enhanced type of checkpoint inhibitor, which strengthens the immunosuppression reversal function of the anti-PD-1 antibody via CD28/B7 co-stimulation and maintains a more sustained CTL response. The ASPIRE nanovaccine shows promise as an efficient strategy to activate strong antitumour immune responses and overcome stubborn immune tolerance.
Engineering and characterization of DC NVs
To induce endogenous presentation, we introduced antigen and stimulated DC differentiation by virus infection. As a proof of principle, we generated a recombinant adenovirus that expresses membrane localization modified green fluorescent protein (rAd-GFP) or ovalbumin (rAd-OVA) (Supplementary Fig. 1a). We also transduced immature DC2.4 cells to express the membrane localization model antigen GFP (Fig. 1a and Supplementary Fig. 1b) and differentiated them into mature DCs (DC-rAd-GFP) (Supplementary Fig. 1c). DCs transduced for 24 hours with rAd-GFP could induce a higher level of MHC-I (Supplementary Fig. 1d,e). This suggests that adenovirus infection promotes the presentation of antigen with MHC-I molecules on DCs. For further verification, we transduced DCs with rAd-OVA, and showed a greater efficacy than those of the other treatments (Supplementary Fig. 2).
a, Generation of DCNVs derived from adenovirus-infected mature dendritic cells. (1) The genes of tumour-specific antigen were genetically engineered into the adenovirus vector. (2) Recombinant adenovirus infected the immature DC2.4 cells to express the modified antigen on the cell surface and stimulate it. (3) Differentiation, maturation and antigen presentation. (4) Harvesting of the induced mature cell membrane and preparation of DCNV-rAd-Ag. b, Schematic illustration of the generation of DCNV-rAd-Ag. c,d, Cryo-electron microscopy (c) and dynamic light scattering analyses (d) showed uniform DCNV-rAd-Ag (approximately 108 nm average diameter, polydispersity index = 0.14) with a vesicle-like morphology. Scale bar, 50 nm. e, The western blot on membrane proteins from DCNV-rAd-GFP demonstrates a similar protein content on the surface compared to that of the parental cells. Panels c–e show representative results of two independent experiments with similar results. f, Comparison of upregulated immune-response-related proteins in NVs and DCs. g, The relative abundance of antigen presentation and migration-related proteins on DCNV-rAd-GFP. r.p.m., revolutions per minute. CCR, CC chemokine receptor; CXCR, C-X-C chemokine receptor; EpCAM, epithelial cellular adhesion molecule; ICAM 1, intercellular adhesion molecule 1; pMHC-I, peptide-major histocompatibility complex class I.
DCNV-rAd-Ag was isolated by multistep density gradient ultracentrifugation (Fig. 1b). From 108 DCs, 37.43 ± 6.68 mg of DCNVs could be obtained. The NVs had a uniform vesicular morphology (Fig. 1c) and the average diameter was approximately 108 nm (Fig. 1d). In fact, such NVs were found to remain stable for many situations (Supplementary Fig. 3a–c). The content of the main functional membrane proteins on the surface of DCNV-rAd-Ag was similar to that of the parental cells, DC-rAd-Ag (Fig. 1e and Supplementary Fig. 3d)29,30. The total proteins of NVs and DCs were then isolated and identified using mass spectrometry. We found that a large number of proteins were upregulated on DCNV-rAd-GFP (Fig. 1f). We found that a variety of co-stimulatory molecules (CD80, CD86, CD40 and so on) were upregulated on DCNV-rAd-GFP. These proteins are known to be involved in the process of antigen presentation and enhance the immune response31 (Fig. 1g). Notably, we also found the overexpression of various chemokines (CCR2, CCR5, CCR7 and so on) on DCNV-rAd-GFP, which can enhance the migration to LNs32.
Directly activates naive CD8+ T cells
Based on the characteristics described, we next evaluated the antigen presentation of DCNV-rAd-Ag to naive CD8+ T cells (Fig. 2a). CD8+ T cells from the spleens of C57BL/6 mice were co-cultured with various doses of DCNV-rAd-GFP. The DCNV-rAd-GFP group showed significant T-cell activation in comparison with that of other groups. Consistent results were observed in the activation of primary human T cells (Supplementary Fig. 4). We then examined the impact of DCNV-rAd-Ag on T-cell-specific proliferation. The results showed that the DCNV-rAd-OVA promoted a strong OT-1 CD8+ T-cell proliferation and activation. In contrast, 293TNV-rAd-OVA and blank DCNVs failed to induce a T-cell proliferation or activation (Fig. 2e–g and Supplementary Fig. 5). This suggests that DCNV-rAd-Ags are endowed with the complete surface functional proteins of mature DCs, which can directly present antigens to naive T cells in vitro.
a, Schematic describing DCNV-rAd-Ag for specific activation and proliferation of CD8+ T cells. b–d, GFP-specific activation of primary mouse T cells: 2 × 105 CD8+ T cells, selected from the spleen of C57BL/6 mice, per well were co-cultured with different doses of DCNV-rAd-GFP (0.5, 5 and 20 µg ml–1) (b) and enzyme-linked immune absorbent spot (ELISpot) analysis of IFN-γ (c) or TNF-α (d) spot-forming cells among splenocytes after 7 days of incubation. e–g, OVA-specific expansion and activation of primary mouse T cells: 105 OT-I CD8+ T cells per well were co-cultured with different doses of DCNV-rAd-OVA (0.5, 5 and 20 µg m–1l) (e), incubation for 24 h, followed by assessment of the T-cell expansion (f) and activation (g). h–j, DCNV-rAd-Ag for LN targeting. C57BL/6 mice were subcutaneously administered at the tail base with 60 µg of ICG-labelled vesicular vaccine or 2 × 105 complete DC vaccine. Fluorescence signals in the draining inguinal LNs of mice treated with ICG-encased DCNV-rAd-OVA (h) or 293TNV-rAd-OVA (i) were quantified with IVIS after 12 h, or, draining inguinal LNs of mice treated with DCNV-rAd-GFP or DC-rAd-GFP were harvested after 12 h and frozen sections were prepared for confocal microscopy (j). Scale bar, 50 µm. The data are shown as the mean ± s.d. from a representative experiment of 2–3 independent experiments with n = 3 (c,d,f,g) and n = 4 (i) biologically independent samples. The data were analysed by two-way analysis of variance (ANOVA) (c,d,f), one-way ANOVA (g) or two-tailed unpaired Student’s t-test (i) with Bonferroni multiple comparisons post-test. CD8+ T, CD8+ T cell.
Successfully achieving antigen presentation in vivo requires DCNV-rAd-Ags to target LNs and make full contact with T cells. We labelled DCNV-rAd-OVA with indocyanine green (ICG) and quantified the biodistribution after subcutaneous (s.c.) injection at the tail base of mice24. The results showed efficient accumulation of DCNV-rAd-OVA in the peripheral LNs within 12 h, whereas the other organs did not reveal a notable fluorescence signal (Fig. 2h,i, Supplementary Fig. 6). The 293TNV-rAd-OVA had a minimal fluorescence signal in the inguinal draining lymph nodes (dLNs). This may be ascribed to the lymphoid homing molecules derived from the mature DC surface, which induced the LN tropism of DCNV-rAd-Ag29. C57BL/6 mice were injected subcutaneously at the tail base with DC-rAd-GFP and had a minimal GFP signal in inguinal dLNs after 12 h. In contrast, the DCNV-rAd-GFP group exhibited a markedly increased GFP signal in dLNs (Fig. 2j). Benefiting from its nanoscale size and good stability, DCNV-rAd-Ag can more efficiently overcome tissue barriers and enrich the LNs than whole DCs.
Elicitation of robust CD8+ T-cell responses in vivo
We assessed the systemic T-cell activation of DCNV-rAd-OVA. Splenocytes isolated from animals in the DCNV-rAd-OVA treatment group demonstrated the highest concentration of cytokines after re-stimulation ex vivo with a peptide pool (Fig. 3a). There were enlarged spleens and increased cell numbers in the DCNV-rAd-Ag group, which indicated an efficient stimulation (Supplementary Fig. 7). The control vesicle vaccine 293TNV-rAd-OVA induced only 6–7% Ag-specific CTLs after the third immunization. As a benchmark, we also vaccinated animals with 5 μg of Ag emulsified in aluminium(III) hydroxide (AlumOH), which is the only approved adjuvant in the United States33,34. OVA + AlumOH elicited only ∼2% Ag-specific CTLs after priming, yet no remarkable T-cell expansion was observed after immunization three times. This is consistent with recent studies regarding a poor performance of recruiting cytotoxic T cells after immunizations with the alum adjuvant35,36,37. In contrast, the DCNV-rAd-OVA group elicited a peak frequency of ∼26% Ag-specific CD8+ T cells after the third vaccination (Fig. 3b and Supplementary Figs. 8 and 9). When challenged with 2 × 105 Hep1-6-OVA cells, mice immunized with DCNV-rAd-OVA had no detectable tumour masses. In contrast, mice in other groups succumbed to tumours with only marginal survival benefits (Fig. 3c and Supplementary Fig. 10). We observed different degrees of liver metastasis in the other groups at day 18 after Hep1-6-OVA inoculation, whereas tumour metastasis was effectively prevented in the DCNV-rAd-OVA group (Fig. 3d).
a,b, C57BL/6 mice were immunized with the indicated formulations on days 0, 14 and 28. Enzyme-linked immunosorbent assay (ELISA) analysis of the interleukin 2 (IL-2), TNF-α and IFN-γ production of splenocytes after ex vivo re-stimulation with SIINFEKL on day 42 (a), and their representative scatter plots and frequency of SIINFEKL-specific CD8+ T cells in the peripheral blood measured by flow cytometry analysis of tetramer+CD8+ T cells (b). c,d, Tumour challenge experiments. c, On day 42, prevaccinated animals were challenged with a s.c. flank injection of 2 × 105 Hep1-6-OVA cells, and tumour growth was measured over time. d, Representative pictures of the lungs and number of liver metastatic nodules counted on day 16 after the Hep1-6-OVA challenge. e, Tumour growth (± CD4‐ or CD8‐depleting antibody) in the control or DCNV-rAd-Ag-immunized C57BL/6 mice. f, Schematic describing B7-1/2–/– mice. g,h, Tumour challenge in antigen-deficient mice. g, Vaccinated gene knockout animals were challenged after the final immunization by Hep1-6-OVA cells, and tumour growth was measured over time. h, The frequency of peripheral blood CD8+ T cells was measured by flow cytometry analysis. The data are shown as the mean ± s.d. from a representative experiment of 2–3 independent experiments with n = 5 (a–e,g,h) biologically independent samples. The data were analysed by two-way ANOVA (c,e,g) or one-way ANOVA (a,b,d,h) with Bonferroni multiple comparisons post-test. CD8+ T, CD8+ T cell.
We depleted CD4+ or CD8+ T cells from mice. The results showed that the immunotherapeutic enhancement facilitated by DCNV-rAd-OVA was lost after CD8+ T-cell depletion, but not after CD4+ T-cell depletion (Fig. 3e). The greater therapeutic efficacy also resulted in an improved survival (Supplementary Fig. 11). Thus, it can be concluded that the antitumour effect of DCNV-rAd-Ag is mainly due to the activation of CD8+ T cells. We assessed whether DCNV-rAd-OVA directly present the antigen to CD8+ T cells proactively or deliver the antigen to CD8+ T cells after they are passively taken up by the APCs. This was studied in antigen-presentation-disabled mice in which B7-1/2 on the APCs were knocked out (Fig. 3f and Supplementary Fig. 12). DCNV-rAd-OVA also showed a strong resistance to tumours in B7-1/2–/– mice, but the other groups were unable to inhibit tumour growth in the mice (Fig. 3g). Impressively, the DCNV-rAd-OVA treatment strategy yielded a 100% complete response. Additionally, animals treated with DCNV-rAd-OVA had more peripheral blood CD8+ T cells than those that received other vaccine treatments (Fig. 3h). These data imply that DCNV-rAd-OVA can directly present antigens to CD8+ T cells in vivo and activate a strong and effective antigen-specific CTL response.
We evaluated the DCNV platform using a B16F10 murine melanoma model to demonstrate its utility for vaccination against neoantigens. This type of tumour is highly aggressive and poorly immunogenic, which thus makes it hard to treat with conventional cancer vaccines38. To prevent a tumour immune escape by the loss of a single mutant allele, we sought to elicit broad-spectrum T-cell responses by employing multiple antigens (multiAgs), which included the recently reported B16F10 mutated neoepitopes M27 and M30, as well as tyrosinase-related protein 2 (TRP2), which is a melanoma-associated Ag. These antigens were all engineered on the DCNVs. Compared with the traditional multiAgs–AlumOH vaccine, therapeutic vaccination with DCNV-rAd-MultiAgs induced polyfunctional interferon-γ+ (IFN-γ+) and IFN-γ+ tumour necrosis factor-α+ (TNF-α+) multiAgs-specific CD8+ T cells and substantially delayed B16F10 tumour growth. However, no tumour rejection was observed in either vaccine group (Supplementary Figs. 13 and 14). This could potentially be due to the immunosuppressive tumour microenvironment, which features high expression levels of PD-1 and its ligand PD-L1 among dLN CD8+ T cells and tumour cells, respectively (Supplementary Fig. 15).
Strong antitumour effect of ASPIRE
ASPIRE was further engineered based on the DCNV platform to break the immunosuppressive PD1/PD-L1 pathway while activating the immune response. The anti-PD1 antibody was pre-expressed on DC surfaces by transmembrane modification18, and αPD1 DCs were obtained. The αPD1 DCs were then induced to translate into αPD1-DC-rAd-MultiAgs by rAd-MultiAgs. After further nanosizing treatment, as mentioned above, ASPIRE was finally prepared (Fig. 4a and Supplementary Fig. 16). ASPIRE treatment led to complete tumour regression in all the mice. In contrast, a tumour regression rate of ∼40% occurred when using general combination immunotherapy with DC-rAd-MultiAgs and the same dose of free anti-PD-1 (Fig. 4b,c). Notably, 100% of the surviving mice rejected a subsequent rechallenge with B16F10 cells intravenously injected on day 80 (Fig. 4d). This result indicates that there is a strong immunological memory against tumour recurrence. We speculate that the enhanced antitumour effect may be related to B7-1/2, important co-stimulatory molecules for anti-PD1 therapy21,22, on the surface of ASPIRE. Importantly, throughout our studies, we did not observe any signs of toxicity, autoimmunity or immune responses directed against the DCNVs in animals that were immunized multiple times with ASPIRE (Supplementary Fig. 17).
a, Process for preparing ASPIRE from DCs. b,c, C57BL/6 mice were inoculated subcutaneously with 105 B16F10 tumour cells and vaccinated with the indicated formulations (60 µg NVs or 10 nmol of each antigen peptide combined with 10.3 µg of anti-PD-1) on days 10, 17 and 24. Average (b) and individual (c) B16F10 tumour growth curves are shown with the fraction of complete tumour regression (CR). d, ASPIRE vaccinated and surviving mice were intravenously rechallenged with 5 × 104 B16-Luc cells two months after the third vaccination. The representative photos show progression of the lung metastases after the B16-Luc challenge. The data are given as the mean ± s.d. from a representative experiment of 2–3 independent experiments with n = 5 (b) biologically independent samples. The data were analysed by two-way ANOVA (b), or log rank (Mantel–Cox) test (b) with Bonferroni multiple comparisons post-test.
B7 co-stimulation enhances anti-PD1-based therapy
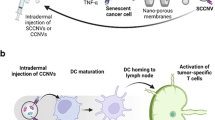
We explored whether the co-delivery of B7-1/2 and anti-PD1 promotes immunosuppression reversal and reactivates exhausted CD8+ T cells to restart the CTL response (Fig. 5a). To this end, αPD1-DCNVs were prepared after the αPD1-DCs were induced to maturity by cytokines. We validated the effect using a Lewis lung carcinoma (LLC) model, which is sensitive to anti-PD1 therapy39,40. To eliminate interference from T-helper cells, we predepleted CD4+ T cells in mice (Fig. 5b). The results showed that CD8+ T cells isolated from tumours in the αPD1-DCNV treatment group demonstrated the highest percentage of granzyme B (Grz B) and IFN-γ-secreting CD8+ T cells (Fig. 5c). There was rapid tumour growth in all the untreated mice and the mice that received 10 μg of free anti-PD1, whereas αPD1-DCNVs elicited tumour regression in eight of ten animals. In contrast, all the mice that received αPD1-DCNVB7–/– (αPD1-DCNVs with B7-1/2 knockout) showed tumour progression.
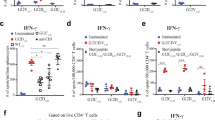
a, Schematic describing ASPIRE for the co-delivery of B7-CD28 and anti-PD1 to promote immunosuppression reversal. b, Mice were depleted of CD4+ T cells for the duration of the experiment. LLC tumour-bearing mice were enrolled into different treatment groups as indicated. c, The data shown are frequencies of Grz B or IFN-γ that produced tumour-infiltrating CD8+ T cells harvested on day 23 from mice that received different treatment regimens, quantified by flow cytometry analysis with intracellular cytokine staining. d, Individual tumour growth, represented by tumour volume. The data show one representative experiment out of three independent experiments. e, Survival curves from the data in d. The data show one representative experiment (ten mice per group) out of three independent experiments. f, Percentage of mice unable to control tumour growth. The data show a summary of three independent experiments with n = 10 biologically independent samples. For the box plots in c, the centre of the box shows the median, the bounds show the interquartile range, the lower whisker extends from the lowest value (minimum) to the 25th percentile and the upper whisker extends from the 76th percentile to the highest value (maximum). The data are shown as the mean ± s.d. from a representative experiment out of three independent experiments with n = 10 (c,e) biologically independent samples. The data were analysed by one-way ANOVA (c,f) with Bonferroni multiple comparisons post-test, or log rank (Mantel–Cox) test (e). Exhausted T, Exhausted T cell.
The effectiveness of PD1 therapy to suppress LLC tumour growth resulted in a significant improvement in the overall survival of mice treated with αPD1-DCNV in comparison with those of the untreated mice, those treated with free anti-PD1 and those that received αPD1-DCNVB7–/–. Furthermore, there was no substantial improvement in the survival of mice treated with αPD1-DCNVB7–/– in comparison with those treated with free anti-PD1 (Fig. 5d,e). A summary of three independent experiments is shown in Fig. 5f. As expected, αPD1-DCNVB7–/– failed to control the growth of the LLC tumour in mice with a deficiency of B7-1/2 co-stimulatory molecules. Our results show that B7-1/2 co-stimulation is important to enhance anti-PD1 therapy by αPD1-DCNVs.
ASPIRE for immunosuppression reversal
We explored the impact of the ASPIRE integration strategy on the interaction between antigen self-presentation and immunosuppression reversal (Fig. 6a). This was accomplished by using a murine MC-38 colon carcinoma model that harbours an MHC I-restricted neoepitope (ASMTNMELM)41. PD-1+CD38hiCD8+ T cells have been described as a population of dysfunctional cells that fail to respond to antigenic stimulation and do not elicit effector functions42,43. When ASPIRE (αPD1-DCNV-rAd-neo) was given, there was a significant decrease in the frequency of PD-1+CD38hi in tumour-infiltrating CD8+ T cells and a significant increase in the frequency of tumour-infiltrating antigen-specific CD8+ T cells. Moreover, ∼80% of the tumour-infiltrating CD8+ T cells were IFN-γ+ and Grz B+ T cells. In contrast, 60% IFN-γ+ and 40% Grz B+ T cells were obtained when using the non-integrated combination strategy of ASPIRE groups with αPD1-DCNV+DCNV-rAd-neo (Fig. 6b,c and Supplementary Fig. 18). ASPIRE treatment led to an impressive rate of MC-38 tumour rejection with 100% of the mice being free of tumours. The general combination therapies of αPD1-DCNV+DCNV-rAd-neo and anti-PD1+DCNV-rAd-neo led to tumour regression in ∼60% and 20% of the animals, respectively (Fig. 6d).
a, Schematic describing ASPIRE for antigen self-presentation and immunosuppression blockade. b, C57BL/6 mice were inoculated subcutaneously with 3 × 105 MC-38 tumour cells and vaccinated with the indicated formulations (60 µg of NVs or combined with 100 µl of anti-PD1 antibody) on days 10, 17 and 24. c, The frequency of CD38hi or antigen-specific T cells in tumour-infiltrating CD8+ T cells, and percentages of Grz B or IFN-γ that produced tumour-infiltrating CD8+ T cells harvested on day 20 from mice that received different treatment regimens quantified by flow cytometry analysis with intracellular cytokine staining. d, Individual MC-38 tumour growth curves with the fraction of complete tumour regression and survival monitored over time. e, Schedule of mouse treatments. Tumour-infiltrating T cells were collected 7 days after treatment. f, Frequency of PD-1+CD38hi cells in the total CD8+ T cells. g, Schedule of mouse treatments. Tumour-infiltrating central and effector memory T cells were collected 17 days after treatment. h, Frequency of CD62L+CD44+ and CD62L-CD44+ cells in the total CD8+ T cells after various treatments. i, C57BL/6 mice were inoculated subcutaneously with 3 × 105 heterogeneous tumour cells (2.4 × 105 MC-38-OVA cells mixed with 6 × 104 MC-38 cells) and vaccinated with the indicated formulations (60 µg of NVs, 2 × 105 DCs or 0.2 nmol OVA) on days 10, 17 and 24. j, The proportions of OVA-negative tumour cells in the tumour tissue of the phosphate buffered solution (PBS) group detected on days 10 and 20. k, Shown are the percentages of mature DCs (CD11c+CD80+MHCII+) in tumour-infiltrating DCs on day 20. l, Survival curves. m, Individual tumour growth curves are shown with the fraction of complete tumour regression (CR). For box plots in c, f and h, the centre of the box shows the median, the bounds show the interquartile range, the lower whisker extends from the lowest value (minimum) to the 25th percentile and the upper whisker extends from the 76th percentile to the highest value (maximum). The data are shown as the mean ± s.d. from a representative experiment of 2–3 independent experiments with n = 10 biologically independent samples. The data were analysed by one-way ANOVA (c,k) with Bonferroni multiple comparisons post-test, or log rank (Mantel–Cox) test (d,l). NS, not significant.
To understand the reason behind the antitumour effect of ASPIRE, we profiled the T-cell infiltrates in the tumour microenvironment using schedules based on time and sequence difference (Fig. 6e). Vaccination using αPD1-DCNV+DCNV-rAd-neo resulted in a significant decrease in PD-1+CD38hiCD8+ T cells in the MC-38 model, which was further decreased when ASPIRE was given. On the other hand, when αPD1-DCNV was given before or after the DCNV-rAd-neo vaccine, the level of total PD-1+CD38hiCD8+ T cells was slightly higher or did not change compared to the αPD1-DCNV+DCNV-rAd-neo vaccine group and was significantly higher than that of the ASPIRE group (Fig. 6f). We found that mice treated with ASPIRE had significantly higher levels of central (CD62L+CD44+) and effector (CD62L−CD44+) memory T cells than the other groups (Fig. 6g,h and Supplementary Fig. 19). The concomitant administration of αPD1-DCNV and DCNV-rAd-neo or the administration of αPD1-DCNV three days after the DCNV-rAd-neo vaccination could enhance the memory T cell infiltration. However, administering αPD1-DCNV before the DCNV-rAd-neo vaccine obviously attenuated such an enhancement in the tumour microenvironment. These results are consistent with recent reports that PD-1 blockade in unprimed or suboptimally primed CD8+ cells induces resistance through the induction of PD-1+CD38hiCD8+ cells43, and such resistance is reversed by ASPIRE.
Therefore, we consider ASPIRE to be an optimal vaccine strategy for both immune activation and immunosuppression reversal. This bioinspired nanosystem takes advantage of genetically engineered cell membranes as a natural medium for most biological reactions and as an integrated platform to express functional proteins. Specifically, this system shows a promising spatiotemporal effectiveness for multiple co-stimulatory signal transduction.
The remarkable antitumour effect of ASPIRE benefits from various functional protein molecules on its surface, of which the two most important ones are B7-1 and B7-2. The B7-CD28 pathway provides co-stimulatory signals not only for antigen presentation, but also for anti-PD1-based therapy. CTLs were transferred into MC-38 tumour-bearing B7-1/2–/– mice and combined with different forms of PD1 therapy (Supplementary Fig. 20), and the results suggest that the B7-CD28 signalling pathway is necessary for co-stimulation in anti-PD1-based therapy, which is consistent with the latest findings21,22.
The pressure of tumour vaccine therapy may lead to the selection of target antigen negative cancer cell clones, which is an important mechanism of tumour immune escape. The antitumour efficiency of ASPIRE (αPD1-DCNV-rAd-OVA) was also verified in heterogeneous tumour models with various MC-38 to MC-38-OVA cell ratios, although tumours with 50% or more MC-38 cells escaped immune control (Fig. 6i–m and Supplementary Figs. 21 and 22). Notably, all antigen-presentation-deficient (DC incompetence) mice that received ASPIRE showed tumour progression and the survival of mice with heterogeneous tumours did not substantially improve (Supplementary Fig. 23).
Conclusions
By taking advantage of established manufacturing procedures and excellent safety profiles, the ASPIRE vaccine platform shows notable promise to dramatically improve the delivery of antigens to LNs (Fig. 2). It could also break the general routine of vaccine development, increase the efficiency of antigen presentation through antigen self-presentation and drive CD8+ T-cell-dominated immunity against tumours with strong CTL immune responses (Fig. 3). Conventional vaccines often fail to eliminate tumours due to the immunosuppressive PD-L1/PD-1 pathway. ASPIRE integrates antigen self-presentation and immunosuppression reversal, maximizes the cytotoxic potential of T cells, achieves a potently amplified therapeutic effectiveness, effectively suppresses tumour growth and provides long-term immunoprotection (Fig. 4). We also confirmed the important role of the B7-CD28 co-stimulation signal in anti-PD1-based therapy (Fig. 5). B7 co-delivery was applied to anti-PD1-based therapy in a demonstration of antitumour efficacy with a personalized vaccine formula that has the power to directly activate both native T cells and exhausted T cells (Fig. 6).
The present work provides a framework for future clinical translation. The ASPIRE system is designed to induce neoantigen-specific CTL immunity, which requires the identification and screening of tumour neoantigens. This can be achieved with the rapid development of sequencing technology. The recent biomedical breakthroughs in whole-exome sequencing, immunogenomics and computational immunology provided technologies for the comprehensive exploitation of neoantigens for clinical applications44,45,46,47. It is also worth mentioning that, although the production of the ASPIRE formulation is somewhat more complicated than that of a DC vaccine, the chemistry, manufacturing and control of ASPIRE are not a major concern as we have established a standard operating procedure for a scaled synthesis with a proper quality control of the membrane vesicles. Compared with conventional DC vaccines, cell-free ASPIRE has unique advantages in storage and long-distance transportation, which greatly reduces the production cost and labour intensiveness (Supplementary Fig. 24).
In conclusion, we have developed a novel nanovaccine platform with the ability to activate the immune response and break immune tolerance. We have demonstrated its ability to stimulate powerful CTL responses and enhance the immune checkpoint blockade with a remarkable therapeutic efficacy. Owing to the tolerance of ASPIRE to a large protein insertion in a natural form, our approach provides a powerful and facile way to produce personalized cancer vaccines. Furthermore, this platform technology may be generally applicable to treat other diseases as well, such as chronic viral infection, in which T-cell exhaustion often occurs during infection and prevents the optimal viral control48,49.
Methods
Mice and cell lines
All the animals were cared for and treated according to the instructions and approval of the Institutional Animal Care and Use Committee of Xiamen University. Male and female C57Bl/6 mice (6–8 weeks old) were purchased from SLAC (Shanghai). B7-1/2-deficient mice with a C57BL/6 genetic background were generated by the Xiamen University Laboratory Animal Center. OT-1 mice were provided by the Chinese Academy of Medical Sciences. The mice were maintained under specific pathogen-free conditions in the animal facility at Xiamen University. 293T HEK, DC2.4, Hep1-6-OVA, B16F10, B16F10-Luc, MC-38, MC-38-OVA and LLC cells were acquired from the National Institute of Diagnostics and Vaccine Development in Infectious Diseases (Xiamen University). For the isolation of T cells from spleen or tumour tissue, the Pan T-cell Isolation Kit (Miltenyi Biotec) was used. For the isolation of CD8+ T cells, peripheral blood monocytes (PBMCs) from volunteers were isolated, or spleens from C57BL/6 or OT-1 mice were digested by Liberase (Roche Diagnostics) and DNase I (SIGMA-ALDRICH) to generate single-cell suspensions, which were then stained with R-phycoerythrin (PE)-conjugated anti-CD8 antibody and applied to anti-PE microbeads (Miltenyi Biotec) for isolation. Human PBMCs were freshly isolated from Chinese healthy adult volunteers with informed consent. All the experiments that used human PBMCs were approved by the Medical Ethics Committee of the School of Public Health of Xiamen University. Splenic lymphocytes were isolated by a splenic lymphocyte kit (Dakewe Biotech). Unless otherwise specified, DC2.4 is referred to as DC in the article. DC2.4 cells were cultured in a medium with rmGM-CSF (recombinant granulocyte-macrophage colony-stimulating factor)and rmIL-4 (all purchased from ThermoFisher). The culture medium used for DCs and T cells was RPMI-1640 (Gibco), supplemented with 10% fetal bovine serum (Gibco), 100 U ml–1 penicillin (Invitrogen) and 100 µg ml–1 streptomycin (Invitrogen). The culture medium used for 293T HEK, Hep1-6-OVA, B16F10, B16F10-Luc, MC-38, MC-38-OVA and LLC cells was Dulbecco′s Modified Eagle Medium (Gibco) supplemented with 10% fetal bovine serum, 100 U ml–1 penicillin and 100 µg ml–1 streptomycin. Throughout the studies, all the cells were used as received and tested negative for mycoplasma contamination and rodent pathogens.
Recombinant adenovirus construction
A signal sequence led by a Kozak consensus sequence was fused in a frame to the N terminus of an antigen protein/epitope (GFP/OVA, or M27-M30-TRP2/ASMTNMELM with ten copies) gene-constant region, a PDGFR transmembrane domain was fused to the C terminus and a FLAG Tag fused after the transmembrane domain to analyse the epitope expression26,27. The frame was cloned to shuttle plasmid vector pDC316. Finally, the recombinant adenovirus (rAd-GFP/OVA, rAd-MultiAgs and rAd-neo) was produced using the simple system of AdMax (Ad5) (Supplementary Fig. 1). The sequences were designed and analysed with Snapgene (version 2.3.2).
Preparation of DC-rAd-Ag
Immature DC2.4 cells were plated at 1 × 106 cells per well in 12-well plates. After 24 h, the DCs were infected with a recombinant adenovirus titre of 100 multiplicity of infection (MOI), or cells in the control groups were incubated with 20 µg of pDC316-GFP or 50 µg of antigen proteins in various formulations (that is, PBS/Ag/Liposome@Ag/Ag@IONPs), in complete media with the addition of a maturation factor combination (1,000 U ml–1 rmGM-CSF, 1,000 U ml–1 rmIL-4 and 200 U ml–1 rmTNF-α) for different lengths of time (2, 6, 24 and 48 h) at 37 °C with 5% CO2. DCs were harvested, washed with FACS buffer (1% fetal bovine serum in PBS), incubated with anti-CD16/32 (Biolegend) at room temperature and then stained on ice with anti-B7-1-APC/anti-B7-2-FITC/anti-MHC-I(H-2Kb)-PE antibodies or PE-conjugated antimouse SIINFEKL/H-2Kb monoclonal antibody 25-D1.16 (all purchased from eBioscience). Cells were analysed by flow cytometry (Cyan 5, Beckman Coulter). DCs cultured in medium with 1,000 U ml–1 rmGM-CSF and 1,000 U ml–1 rmIL-4 were stained as the background to quantify the MHC-I expression level.
Preparation and characterization of DCNV-rAd-Ag
Immature DC2.4 cells were infected with a recombinant adenovirus titre of 100 MOI, and cultured in the medium with the addition of a maturation factor combination for 24 h.
According to our previous work26,27,28, the cells were collected and washed in cold PBS mixed with protease inhibitor twice to remove cellular debris and culture medium. Next, the cells were suspended in saline and sonicated in a sterile 1.5 ml EP tube under a low power (22.5 W, 1 min) on ice. Then the NVs were isolated by multistep density gradient ultracentrifugation before being resuspended in saline. The product was then introduced to a Mini-Extruder (Avanti Polar Lipids) equilibrated in saline (200 nm pore-sized membrane filters) to further purify uniform NVs (DCNV-rAd-Ag).
DCNV-rAd-MultiAgs wash, DCNV-Ad was prepared from DC2.4 cells that had been infected with a blank adenovirus titre of 100 MOI for 24 h, DCNV was prepared from DC2.4 cells without any treatment, 293TNV-rAd-Ag was prepared from 293T HEK cells that had been infected with a rAd-Ag titre of 100 MOI for 24 h. All the NVs were prepared as mentioned above.
hDCNV-rAd-Ag/hDCNV-Ad/hDCNV was prepared from DCs of human PBMCs. Briefly, human PBMCs were isolated from healthy blood donors using Ficoll density gradient centrifugation (Biocoll, Biochrom AG). CD14-positive monocytes were isolated by MACS selection (Miltenyi), and cultured for 5 days in complete media with the addition of a stimulating factor combination (1,000 U ml–1 rhGM-CSF and 1,000 U ml–1 rhIL-4), and then immature DCs were obtained. Immature DCs were infected with a recombinant adenovirus titre of 100 MOI, and cultured in the medium with the addition of a maturation factor combination (1,000 U ml–1 rhGM-CSF, 1000 U ml–1 rhIL-4 and 200 U ml–1 rhTNF-α) for 24 h, and the corresponding NVs were prepared according to the DCNV-rAd-Ag preparation procedure.
The presence of NVs was verified through the morphological examination by cryo-electron microscopy, transmission electron microscopy and size measurement using dynamic light scattering (Zetasizer software 7.13). To detect the presence of major functional membrane proteins on the DCNV-rAd-Ag, a western blot assay was performed. Briefly, membrane proteins were extracted from 0.5 ml (1 mg ml–1 total protein) of DCNV-rAd-Ag by a ProteoExtract Transmembrane Protein Extraction Kit (Merck). Samples were probed with antibodies against GFP/OVA, MHC-I, B7-1, B7-2, ICAM-I or CCR7 (all purchased from Invitrogen). The major functional membrane proteins were visualized with an HRP-antimouse/rabbit immunoglobulin G antibody (Invitrogen) and ECL substrate (Thermo). To determine the content of Ag in the total amount of protein, a GFP/OVA ELISA kit or FLAG ELISA kit (Abcam) was used to directly or indirectly quantify the amount of Ag. Membrane proteins on DCs and NVs were extracted and then identified by liquid chromatography/tandem mass spectroscopy analysis. Raw data files were processed using Proteome Discoverer (PD) version 1.4 (Thermo Scientific).
Preparation and characterization of ASPIRE
A signal sequence was fused in the N-terminus of the anti-PD1 scFv gene region, and a HIS-Tag was fused before the scFv gene region. To facilitate the analyses of epitope expression, a flexible peptide linker (GGGGS)3 and a PDGFR transmembrane domain were fused to the C terminus. The gene encoding anti-PD1 ScFv was finally cloned into plasmid vector pcDNA3.1(+) to express membrane-localized anti-PD1 antibodies in DCs (αPD1-DC) (Supplementary Fig. 16). Immature DCs transfected with pcDNA3.1(+)-αPD1 (105 cells µg–1) were cultured in the culture medium with the addition of growth factor combination (that is, 1,000 U ml–1 rmGM-CSF and 1,000 U ml–1 rmIL-4). After 48 h of culture, the cells were infected with the recombinant adenovirus mentioned previously (rAd-GFP/OVA, rAd-MultiAgs, rAd-neo), and cultured in the culture medium with the addition of maturation factor combination (1,000 U ml–1 rmGM-CSF, 1,000 U ml–1 rmIL-4 and 200 U ml–1 rmTNF-α) for another 12 h. The cells were collected and NVs were extracted in a similar procedure to that in Preparation and characterization of DCNV-rAd-Ag.
αPD1-DCNV/αPD1-DCNVB7–/– were prepared after αPD1-DC/αPD1-DCB7–/– were induced by maturation factor combination with 12 h of incubation. The cells were collected and NVs were extracted in a similar procedure to that in Preparation and characterization of DCNV-rAd-Ag.
To detect the orientation of the αPD-1 antibody on ASPIRE or αPD1-DCNVs, an immunoprecipitation assay was performed. Briefly, 0.5 ml (1 mg ml–1 total protein) of ASPIRE or αPD1-DCNVs were incubated with beads (Santa Cruz Biotechnology) conjugated to protein A/G for 1 h at room temperature after the addition of 2 μg of anti-His antibody specific for the fusion protein HIS-αPD1. The beads were washed gently thrice with PBS. Sample mixtures were resolved and subjected to 10% SDS–PAGE and analysed by western immunoblot assay.
Antigen self-presentation to activate naive CD8+ T cells
To assess the ability of DCNV-rAd-Ag for antigen self-presentation and to directly activate naive T cells, CD8+ T cells (2 × 105 per well) selected from the spleen of C57BL/6 mice were co-cultured with different doses of DCNV-rAd-GFP/293TNV-rAd-GFP/DCNV-Ad/DCNV formulations (0.5, 5 and 20 µg ml–1 NVs) in RPMI 1640 culture medium. After 7 days of incubation, the cells were transferred to ELISPOT plates (Merck), which were pretreated with 95% ethanol and washed with sterile water and PBS before coating with 15 µg ml–1 anti-IFN-γ or anti-TNF-α (Mabtech) overnight at 4 °C. Unbound antibodies were removed by washing with sterile filtered PBS. After 24 h of incubation, the cells were washed away and the plates were incubated with 1 µg ml–1 biotinylated anti-IFN-γ or anti-TNF-α antibody (Mabtech) for 2 h at room temperature. Thereafter, the plates were washed with PBS and incubated for 90 min at room temperature with streptavidin–alkaline phosphatase (Mabtech) (1/1,000 dilution) followed by washing with PBS. The plates were finally incubated with a BCIP/NBT substrate solution (CALBIOCHEM) at room temperature until spots emerged, which took approximately 1 h. The colour development was stopped by repeated washings with tap water. After drying, the spots were counted with an ELISPOT reader using AID ELISPOT software. To detect the activation of primary human T cells, human PBMCs were isolated from healthy blood donors using Ficoll density gradient centrifugation. CD8-positive T cells were isolated by MACS selection (Miltenyi). CD8+ T cells (2 × 105 per well) were co-cultured with different doses of hDCNV-rAd-GFP/293TNV-rAd-GFP/hDCNV-Ad/hDCNV formulations prepared in PBMCs isolated from the same volunteer (0.5, 5 and 20 µg ml–1 NVs) in RPMI 1640 culture medium. After 7 days of incubation, the cells were transferred to the ELISPOT plates (Merck), and T-cell activation was detected by ELISPOT assay as mentioned above.
To further verify that DCNV-rAd-Ag directly activates T cells and stimulates T-cell specific proliferation, 105 OT-I CD8+ T cells per well were co-cultured with different doses of DCNV-rAd-OVA and 293TNV-rAd-OVA formulations (0.5, 5 and 20 µg ml–1 NVs) in RPMI 1640 culture medium. After 24 h of incubation, the media were aspirated and 150 µl of CPRG/lysis buffer was added and incubated for 90 min, measured at 570 nm using a microplate reader. The IL-2 and TNF-α in the culture supernatant were determined by a Mouse IL-2 ELISA kit or a TNF-α ELISA kit (both purchased from Dakewe Biotech).
For the detection of T-cell proliferation dilution, an OT-I CD8+ T-cell suspension was stained with a carboxyfluorescein succinimidyl ester solution, and co-cultured with 20 µg ml–1 NVs in RPMI 1640 culture medium for 48 h, followed by flow cytometric analysis.
In vivo imaging of DCNVs
For LN draining studies, C57BL/6 mice were treated with DCNV-rAd-OVA@ICG or 293TNV-rAd-OVA@ICG by s.c. injection at the tail base24,26. Under isoflurane anaesthesia, in vivo near-infrared fluorescence imaging was performed using an IVIS Lumina II imaging system at 12 h postinjection. C57BL/6 mice were injected with DCNV-rAd-GFP or DCNV-rAd-GFP. After 12 h, inguinal LNs were harvested, and after a 10% formaldehyde solution fixation, paraffin embedding and freezing section, the GFP fluorescence signal was measured.
In vivo immunization and cancer immunotherapy studies
All the mice used for immunological studies were 6–8-week-old females with a C57Bl/6 background. All the animals were cared for and treated according to the instructions and approval of the Institutional Animal Care and Use Committee of Xiamen University (no. XMULAC20190146), and it was defined that the tumour load of mice must not exceed 1.7 cm (diameter). The tumour volume throughout this study was calculated by the equation: tumour volume = length × width2 × 0.5. Animals were euthanized when the tumour masses reached 1.5 cm in diameter or when the animals became moribund with severe weight loss or ulceration. C57BL/6 mice were immunized with different formulations: NVs (60 µg per mouse), recombinant adenovirus (2 × 107 vector particles per mouse) or complete dendritic cell vaccine (2 × 105 cells per mouse) in 100 µl by s.c. injection at the tail base at the indicated time points. In some studies, antigen emulsified in AlumOH served as a positive control. Briefly, antigen protein (2 nmol) in 0.5 ml of PBS was thoroughly emulsified in 0.5 ml of AlumOH until the mixture was homogeneous, and then administered subcutaneously in a 100-µl-injection volume.
For prophylactic tumour challenge studies, vaccinated animals were challenged on day 14 after the final immunization by the s.c. injection of 2 × 105 Hep1-6-OVA cells per mouse on the right flank, and tumour growth was monitored every other day. Livers were excised on day 18, followed by enumeration of the Hep1-6-OVA lung tumour nodules. For the CD4+/CD8+ cell depletion, 250 µg of GK1.5 antibody or 53-6.7 antibody (BioXcell) were administered to the vaccinated tumour-bearing mice by intraperitoneal injection every 2 weeks. For B7-1/2–/– mice, vaccinated gene knockout animals were challenged after the final immunization by Hep1-6-OVA cells.
For the therapeutic tumour vaccination studies, C57BL/6 mice were inoculated with 1 × 105 B16F10 or MC-38 cells per mouse on the right flank by s.c. injection on day 0 and vaccinated on the indicated days. For the combinatorial therapy groups, the antimouse PD-1 antibody (10.3 µg per mouse; clone, J43 (BioXcell)) was administered intraperitoneally after each vaccination. For the lung metastasis model, the surviving mice after ASPIRE treatment were rechallenged by the intravenous injection of 5 × 104 B16F10-Luc cells per mouse, and a group of untreated mice were inoculated as a control. Under isoflurane anaesthesia, the fluorescence signal of lungs metastasis was measured every day using an IVIS Lumina II imaging system.
For the co-stimulatory study of the anti-PD1 antibody and B7-1/2, C57BL/6 mice were subcutaneously inoculated with 2 × 105 LLC cells and CD4 depletion was maintained for the duration of the experiment by repeated injections of 250 µg of GK1.5 antibody every 7 days. αPD1-DCNVs were administered by s.c. injection at the tail base every 3 days. For the co-stimulatory study in KO mice, B7-1/2–/– mice were inoculated with 1 × 105 MC-38 per mouse by s.c. injection on day 0, and MC-38-specific CTLs were intraperitoneally administered after each vaccination on the indicated days.
For the therapeutic tumour vaccination studies, C57BL/6 mice were inoculated with 1 × 105 MC-38 cells per mouse on the right flank by s.c. injection on day 0, and vaccinated on days 10 and 13 with αPD1-DCNV or DCNV-rAd-neo separately in the order indicated, or vaccinated on day 13 with αPD1-DCNV, DCNV-rAd-neo or ASPIRE (60 µg per dose).
For the antitumour study of ASPIRE in heterogeneous tumours, C57BL/6 mice and antigen-presentation-deficient mice were inoculated subcutaneously with 3 × 105 heterogeneous tumour cells (MC-38-OVA cells mixed with MC-38 cells in various proportions) and vaccinated with the indicated formulations (60 µg of NVs, 2 × 105 DCs or 0.2 nmol OVA) on days 10, 17 and 24. The mature DCs (CD11c+CD80+MHCII+) in tumour-infiltrating DCs were detected on day 20. Intracellular protein staining and flow cytometric analysis were used for the analysis of OVA in heterogeneous tumour cells at the indicated time points.
Phenotypic and functional assessment of T cells
For the analysis of the activation of peripheral blood lymphocytes, at 2 weeks after the final immunization peripheral blood was harvested from the animals in all the treatment groups. The peripheral blood lymphocytes were prepared and incubated in RPMI 1640 media with 1 μM of the SIINFEKL peptides added. After 24 h, the IL-2/TNF-α/IFN-γ production was measured with the ELISA assay system (purchased from Dakewe Biotech) according to the manufacturer’s instructions.
Immunized mice were analysed for the percentages of tumour antigen-specific CD8+ T cells using the tetramer staining assay with a peptide–MHC tetramer tagged with PE (H-2Kb-restricted SIINFEKL, MBL Beijing Biotech). Briefly, the peripheral blood lymphocytes were resuspended in a mouse CD16/32 antibody (0.025 mg ml–1) solution to block non-specific and FcR-mediated antibody binding. The suspension was incubate for 10 min at room temperature and washed five times with FACS buffer. Then H-2Kb OVA Tetramer-SIINFEKL-PE solution was added to each sample and incubated for 30 min on ice. Anti-CD8-APC was added to each experimental sample and incubated for 20 min on ice. The samples were washed twice with FACS buffer, fixed and the cells resuspended for flow cytometry analysis. To assess the functionality of the primed CD8+ T cells, PBMCs were stimulated ex vivo with the peptide pool (1 μM of each antigen peptide, M27, M30 and TRP2) for 6 h, fixed, permeabilized, stained with anti-IFN-γ-eFluor 660/TNF-α-FITC and anti-CD8-APC, and analysed by flow cytometry.
For the analysis of tumour cells or tumour-infiltrating T cells, tumour tissues were excised at the indicated time points, cut into small pieces of 2–4 mm and then placed in a dissociation buffer (1 mg ml–1 collagenase type IV and 0.1 mg ml–1 DNase I in RPMI) for 30 min at 37 °C with gentle shaking. The cell suspension was passed through a 70 µm strainer, washed with FACS buffer and stained with the indicated antibodies or its isotype control, followed by flow cytometric analysis. The intracellular cytokine staining assays on tumour-infiltrating T cells were performed with anti-IFN-γ-eFluor 660 and anti-TNF-α-FITC.
For the analysis of CD8+ T cells in the draining LNs, the draining LNs were excised at the indicated time points. Cell suspensions were prepared and stained with anti-CD8-APC, followed by flow cytometric analysis.
Immunized MC-38 mice were analysed for the percentages of tumour-infiltrating antigen-specific CD8+ T cells using a tetramer staining assay with the peptide–MHC tetramer tagged with PE (H-2Kb-restricted ASMTNMELM, MBL Beijing Biotech), functional cell subtypes were analysed by staining with FITC-anti- PD-1/eFluor450-anti-CD38/PerCP-eFluor710-anti-Granzyme B/FITC-anti-CD44/PE-antimouse CD62L for flow cytometry analysis.
Statistical analysis
All the animal studies were performed after randomization. Experiments were not performed in a blinded fashion. Data were analysed by one- or two-way ANOVA, followed by Bonferroni post hoc test for comparison of multiple groups with Prism (v5.0 and v7.0) (GraphPad Software). Data were normally distributed and the variance between groups was similar. P values less than 0.05 were considered statistically significant. All the values were reported as mean ± s.d. with the indicated sample size. No sample in any representative experiment was excluded from the analysis.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The authors declare that data supporting the findings of this study are available within the article and its Supplementary Information. All the relevant data can be provided by the authors upon reasonable request
References
Couzin-Frankel, J. Cancer immunotherapy. Science 342, 1432–1433 (2013).
Chen, D. S. & Mellman, I. Oncology meets immunology: the cancer-immunity cycle. Immunity 39, 1–10 (2013).
Huppa, J. B. & Davis, M. M. T-cell-antigen recognition and the immunological synapse. Nat. Rev. Immunol. 3, 973–983 (2003).
Fuertes, M. B. et al. Host type I IFN signals are required for antitumor CD8+ T cell responses through CD8α+ dendritic cells. J. Exp. Med. 208, 2005–2016 (2011).
Steinman, R. M. Decisions about dendritic cells: past, present, and future. Annu. Rev. Immunol. 30, 1–22 (2012).
Heath, W. R. & Carbone, F. R. Cross-presentation, dendritic cells, tolerance and immunity. Annu. Rev. Immunol. 19, 47–64 (2001).
Heath, W. R. & Carbone, F. R. Cross-presentation in viral immunity and self-tolerance. Nat. Rev. Immunol. 1, 126–134 (2001).
Albert, M. L. & Bhardwaj, N. Resurrecting the dead: DCs cross-present antigen derived from apoptotic cells on MHC I. Immunologist 6, 194–198 (1998).
Palucka, A. K. et al. Dendritic cells loaded with killed allogeneic melanoma cells can induce objective clinical responses and MART-1 specific CD8+ T-cell immunity. J. Immunother. 29, 545–557 (2006).
Steinman, R. M. & Banchereau, J. Taking dendritic cells into medicine. Nature 449, 419–426 (2007).
Palucka, K. & Banchereau, J. Cancer immunotherapy via dendritic cells. Nat. Rev. Cancer 12, 265–277 (2012).
Banchereau, J. & Steinman, R. M. Dendritic cells and the control of immunity. Nature 392, 245–252 (1998).
Nopora, A. & Brocker, T. Bcl-2 controls dendritic cell longevity in vivo. J. Immunol. 169, 3006–3014 (2002).
Park, D., Lapteva, N., Seethammagari, M., Slawin, K. M. & Spencer, D. M. An essential role for Akt1 in dendritic cell function and tumor immunotherapy. Nat. Biotechnol. 24, 1581–1590 (2006).
Shah, N. N. & Fry, T. J. Mechanisms of resistance to CAR T cell therapy. Nat. Rev. Clin. Oncol. 16, 372–385 (2019).
Ribas, A. & Wolchok, J. D. Cancer immunotherapy using checkpoint blockade. Science 359, 1350–1355 (2018).
Garon, E. B. et al. Pembrolizumab for the treatment of non-small-cell lung cancer. N. Engl. J. Med. 372, 2018–2028 (2015).
Wolchok, J. D. et al. Nivolumab plus ipilimumab in advanced melanoma. N. Engl. J. Med. 369, 122–133 (2013).
Hamid, O. et al. Safety and tumor responses with lambrolizumab (anti-PD-1) in melanoma. N. Engl. J. Med. 369, 134–144 (2013).
Brahmer, J. R. et al. Safety and activity of anti-PD-L1 antibody in patients with advanced cancer. N. Engl. J. Med. 366, 2455–2465 (2012).
Hui, E. F. et al. T cell costimulatory receptor CD28 is a primary target for PD-1-mediated inhibition. Science 355, 1428–1433 (2017).
Kamphorst, A. O. et al. Rescue of exhausted CD8 T cells by PD-1-targeted therapies is CD28-dependent. Science 355, 1423–1427 (2017).
Chen, Z. W., Wang, Z. J. & Gu, Z. Bioinspired and biomimetic nanomedicines. Acc. Chem. Res. 52, 1255–1264 (2019).
Zhang, P. F. et al. Genetically engineered liposome-like nanovesicles as active targeted transport platform. Adv. Mater. 30, 1705350 (2018).
Hu, C. M. J. et al. Nanoparticle biointerfacing by platelet membrane cloaking. Nature 526, 118–121 (2015).
Liu, X. et al. Vesicular antibodies: a bioactive multifunctional combination platform for targeted therapeutic delivery and cancer immunotherapy. Adv. Mater. 31, 1808294 (2019).
Zhang, P. F. et al. Virus-mimetic nanovesicles as a versatile antigen-delivery system. Proc. Natl Acad. Sci. USA 112, E6129–E6138 (2015).
Liu, X. et al. Bioinspired artificial nanodecoys for hepatitis B virus. Angew. Chem. Int. Ed. 57, 12499–12503 (2018).
Sallusto, F., Lenig, D., Forster, R., Lipp, M. & Lanzavecchia, A. Two subsets of memory T lymphocytes with distinct homing potentials and effector functions. Nature 401, 708–712 (1999).
Munro, J. M. Endothelial-leukocyte adhesive interactions in inflammatory diseases. Eur. Heart J. 14, 72–77 (1993).
Chen, L. & Flies, D. B. Molecular mechanisms of T cell co-stimulation and co-inhibition. Nat. Rev. Immunol. 13, 227–242 (2013).
Worbs, T., Hammerschmidt, S. I. & Forster, R. Dendritic cell migration in health and disease. Nat. Rev. Immunol. 17, 30–48 (2017).
Li, H. Y., Li, Y. H., Jiao, J. & Hu, H. M. Alpha-alumina nanoparticles induce efficient autophagy-dependent cross-presentation and potent antitumour response. Nat. Nanotechnol. 6, 645–650 (2011).
Flach, T. L. et al. Alum interaction with dendritic cell membrane lipids is essential for its adjuvanticity. Nat. Med. 17, 479–487 (2011).
Wheeler, C. M. et al. Efficacy, safety, and immunogenicity of the human papillomavirus 16/18 AS04-adjuvanted vaccine in women older than 25 years: 7-year follow-up of the phase 3, double-blind, randomised controlled VIVIANE study. Lancet Infect. Dis. 16, 1154–1168 (2016).
Zhu, F. C. et al. Efficacy and safety of a recombinant hepatitis E vaccine in healthy adults: a large-scale, randomised, double-blind placebo-controlled, phase 3 trial. Lancet 376, 895–902 (2010).
Leslie, M. Solution to vaccine mystery starts to crystallize. Science 341, 26–27 (2013).
Verdegaal, E. M. E. et al. Neoantigen landscape dynamics during human melanoma–T cell interactions. Nature 536, 91–95 (2016).
Bertrand, F. et al. TNF alpha blockade overcomes resistance to anti-PD-1 in experimental melanoma. Nat. Commun. 8, 2256 (2017).
Li, H. Y. et al. The tumor microenvironment regulates sensitivity of murine lung tumors to PD-1/PD-L1 antibody blockade. Cancer Immunol. Res. 5, 767–777 (2017).
Yadav, M. et al. Predicting immunogenic tumour mutations by combining mass spectrometry and exome sequencing. Nature 515, 572–576 (2014).
Chen, L. M. et al. CD38-mediated immunosuppression as a mechanism of tumor cell escape from PD-1/PD-L1 blockade. Cancer Discov. 8, 1156–1175 (2018).
Verma, V. et al. PD-1 blockade in subprimed CD8 cells induces dysfunctional PD-1+CD38hi cells and anti-PD-1 resistance. Nat. Immunol. 20, 1555–1555 (2019).
Bulik-Sullivan, B. et al. Deep learning using tumor HLA peptide mass spectrometry datasets improves neoantigen identification. Nat. Biotechnol. 37, 55–63 (2019).
Robbins, P. F. et al. Mining exomic sequencing data to identify mutated antigens recognized by adoptively transferred tumor-reactive T cells. Nat. Med. 19, 747–752 (2013).
Liu, X. S. & Mardis, E. R. Applications of immunogenomics to cancer. Cell 168, 600–612 (2017).
Schumacher, T. N. & Schreiber, R. D. Neoantigens in cancer immunotherapy. Science 348, 69–74 (2015).
Im, S. J. et al. Defining CD8+ T cells that provide the proliferative burst after PD-1 therapy. Nature 537, 417–421 (2016).
Blackburn, S. D. et al. Coregulation of CD8+ T cell exhaustion by multiple inhibitory receptors during chronic viral infection. Nat. Immunol. 10, 29–37 (2009).
Acknowledgements
This work was supported by the Major State Basic Research Development Program of China (2017YFA0205201) (G.L.), the National Natural Science Foundation of China (81925019, 81422023 and U1705281) (G.L.), the National University of Singapore Startup Fund (NUHSRO/2020/133/Startup/08) (X.C.), the Nanomedicine Translational Research Programme (NUHSRO/2021/034/TRP/09/Nanomedicine) (X.C.), the Fundamental Research Funds for the Central Universities (20720190088 and 20720200019) (G.L.), the China Postdoctoral Science Foundation (2021M702738) (C.L.) and (2021M700115) (Xue Liu) and the Program for New Century Excellent Talents in University, China (NCET-13-0502) (G.L.).
Author information
Authors and Affiliations
Contributions
C.L. and G.L. conceived and designed the experiments. C.L., Xue Liu, X.X., X.P., S.C., L.Z. and X.W. performed the experiments. C.L., Xue Liu, Xuan Liu, Y.Z., E.R., P.L., X.C. and G.L. analysed the data. C.L., Xue Liu, X.C. and G.L. wrote the manuscript. G.L. supervised the entire project. All the authors discussed the results and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
G.L. and C.L. are inventors on a patent (No. ZL202010070724.9) related to this study. The other authors declare no competing interests.
Peer review
Peer review information
Nature Nanotechnology thanks Michele De Palma, Lana Kandalaft and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–24.
Rights and permissions
About this article
Cite this article
Liu, C., Liu, X., Xiang, X. et al. A nanovaccine for antigen self-presentation and immunosuppression reversal as a personalized cancer immunotherapy strategy. Nat. Nanotechnol. 17, 531–540 (2022). https://doi.org/10.1038/s41565-022-01098-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41565-022-01098-0
This article is cited by
-
Harnessing adenovirus in cancer immunotherapy: evoking cellular immunity and targeting delivery in cell-specific manner
Biomarker Research (2024)
-
Roles of exosomes in immunotherapy for solid cancers
Cell Death & Disease (2024)
-
Insights into vaccines for elderly individuals: from the impacts of immunosenescence to delivery strategies
npj Vaccines (2024)
-
Modification of immune cell-derived exosomes for enhanced cancer immunotherapy: current advances and therapeutic applications
Experimental & Molecular Medicine (2024)
-
SRSF7 is a promising prognostic biomarker in hepatocellular carcinoma and is associated with immune infiltration
Genes & Genomics (2024)