Abstract
Unlike conventional antimicrobials, the study of bacterial resistance to silver nanoparticles (AgNPs) remains in its infancy and the mechanism(s) through which it evolves are limited and inconclusive. The central question remains whether bacterial resistance is driven by the AgNPs, released Ag(I) ions or a combination of these and other factors. Here, we show a specific resistance in an Escherichia coli K-12 MG1655 strain to subinhibitory concentrations of AgNPs, and not Ag(I) ions, as indicated by a statistically significant greater-than-twofold increase in the minimum inhibitory concentration occurring after eight repeated passages that was maintained after the AgNPs were removed and reintroduced. Whole-population genome sequencing identified a cusS mutation associated with the heritable resistance that possibly increased silver ion efflux. Finally, we rule out the effect of particle aggregation on resistance and suggest that the mechanism of resistance may be enhanced or mediated by flagellum-based motility.
Similar content being viewed by others
Main
Nanoenabled antimicrobials comprise a multibillion dollar industry, and silver-based formulations are by far the most widely used1,2,3. Silver nanoparticles (AgNPs) are promising alternatives to conventional antimicrobials (for example, antibiotics and disinfectants) owing to their efficacy against a wide spectrum of bacteria4,5,6. A major challenge associated with use of any antimicrobial is decreased efficacy over time because of the development of resistance in the target microbe. However, unlike conventional antimicrobials, the study of AgNP resistance in bacteria and the mechanism(s) through which it occurs is limited and inconclusive. Resistance to Ag(I) ions has been well-established, whereas the evolution of AgNP resistance in bacteria has only recently been recognized7,8,9,10,11,12,13.
There are many, well-established reports of bacterial resistance to ionic Ag that identify elimination of Ag(I) ions via efflux pumps or reduction to the less toxic Ag(0) oxidation state as resistance mechanisms11,14,15,16,17. Further, Ag-resistant mutants of E. coli have cell membranes that are deficient in outer membrane porins thus decreasing Ag uptake15. On a genetic level, Ag(I) ion resistance is specifically associated with the sil operon in Gram-negative bacteria, which encodes two efflux pumps—a P-type ATPase (SilP) and antiporter (SilCBA)—and two periplasmic silver-binding chaperone proteins (SilE and SilF) that work synergistically to drive Ag(I) ions out of the bacteria16,17.
Over the past 10 years, there have been few studies on bacterial responses to repeated chronic exposure of subinhibitory concentrations of AgNPs. Some studies demonstrate AgNP resistance7,8,9,10,11,12, while other studies postulate that AgNPs pose low risk of bacterial resistance development because they concurrently affect so many aspects of bacterial physiology18,19. AgNP resistance is also shown to be microorganism dependent, with different bacterial taxa exhibiting differential resistance profiles and some not developing any resistance at all7,10,11. Furthermore, comparing resistance evolution in response to Ag(I) ions versus AgNPs reveals another inconsistency with some studies reporting a differential resistance response10 and others observing similar resistance responses between AgNPs and Ag(I) ions8,11,12. Despite these conflicting outcomes, the majority of the scientific community attribute AgNP resistance to the premise that AgNPs serve as reservoirs for Ag(I) ions and elimination of Ag(I) ions occurs via the mechanisms discussed above7,8,9,11. The central question remains: is observed bacterial resistance driven by the AgNPs, Ag(I) ions, or a combination of these and other factors? Other factors include (i) AgNP aggregation, which was recently proposed as a resistance mechanism induced by the bacterial flagellin protein10 and (ii) strain-specific behaviour related to flagellum-mediated motility as a key fitness trait versus simply a protein-binding function. Answering this question is necessary to ensure that AgNPs aid in combatting, rather than contributing to, the global antimicrobial resistance crisis, and to inform the potential to evade resistance through the design of AgNPs.
This study examines the experimental evolution and genetic stability of E. coli resistance in response to repeated long-term exposure to subinhibitory concentrations of Ag(I) ions versus AgNPs over multiple passages in one population, as well as the genetic stability of the imparted resistant phenotype when the pressure is removed. The adopted approach includes a non-selected cell-only control population and antibiotic serial passaging controls, with robust statistical analysis to confirm the resistant phenotypes. Our results further underscore the important role of bacterial motility, a key fitness trait for many bacteria, and other possible strain-specific traits in E. coli AgNP resistance by comparing resistance responses of a hypermotile E. coli K-12 MG1655 (+IS1) strain and a non-motile E. coli K-12 JW1881 strain (ΔflhD::kan) (bacteria strain information in Supplementary Table 1).
E. coli develops resistance to AgNPs but not to Ag(I) ions
Pseudospherical citrate-capped AgNPs were synthesized according to a previously reported procedure20,21,22 and functionalized with poly(ethylene glycol) methyl ether thiol (PEGSH, average molecular weight 1,000 g mol−1) using mass action ligand exchange (Scheme S1 in ref. 20). Poly(ethylene glycol) methyl ether thiol is a widely used biocompatible and charge-neutral ligand20,23 which mitigates electrostatic attractive forces with negatively charged bacteria. After ligand exchange and subsequent purification, the particle size was determined to be 27.2 ± 4.8 nm (Supplementary Fig. 1) by transmission electron microscopy (TEM). PEGSH ligand density of this particle size was determined to be approximately 1.0 PEGSH nm−2 using a combination of proton-nuclear magnetic resonance spectroscopy (1H NMR), TEM and inductively coupled plasma mass spectrometry (ICP-MS) (Supplementary Figs. 3 and 5 in ref. 20). Ultraviolet–visible–near infrared (UV–vis–NIR) spectra of the AgNPs revealed a characteristic localized surface plasmon resonance (LSPR) peak at wavelength λmax ≈ 420 nm over the duration of the bacterial resistance experiments (that is, 5 weeks), indicating the presence of stable, well-dispersed colloidal AgNPs (Supplementary Fig. 2a). Additionally, AgNP size (determined by TEM) did not change significantly over time (Supplementary Fig. 2b). Combined, these evaluations of long-term stability support the conclusion that any fold increase in the minimum inhibitory concentration (MIC) (that is, the lowest concentration of antimicrobial agent that inhibits visible growth of the tested microorganisms) after repeated passaging (vide infra) is not attributable to differences in AgNP morphology and instability over time.
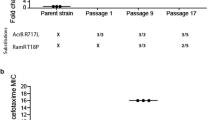
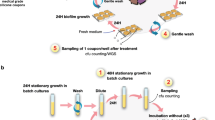
An experimental evolution approach was adopted to study responses to prolonged AgNP and Ag(I) ion exposure, in which the populations were subcultured every 24 h onto Luria–Bertani broth (LB) agar and then into 50 ml fresh LB followed by dosing of fresh AgNPs and Ag(I) ions. This ensures insignificant carry over of Ag species (in the forms of AgNPs or Ag(I) ions) from the previous culture into the subsequent culture and comparable AgNP and Ag(I) ion concentrations at each stage of the prolonged exposure. The incorporation of subculturing periods without Ag exposure in between the prolonged exposure periods also mimics (i) clinical in vivo conditions where bacteria might encounter Ag intermittently through pulse dosing (which would create prolonged dose-free periods allowing Ag exposure levels to diminish until the next dose) and (ii) incidental release of Ag into the environment which may also create similar intermittent or cyclic exposure conditions. Using this approach, we observed an average greater-than-twofold increase (P = 0.0009) in the hypermotile E. coli (+IS1) MIC after eight passages with AgNPs (Fig. 1a; Supplementary Fig. 3 includes the raw MIC data), which was maintained throughout additional passages. An additional replicate lineage was carried out for the AgNPs and nearly all other treatment conditions to confirm the biological response (Supplementary Fig. 4). For each treatment condition, only phenotypic and genotypic data corresponding to the evolution and genetic stability of one of the lineages is reported and discussed below. A non-selected control population (that is, serial passaging of untreated bacteria) was included and maintained to account for potential background effects of repeated passaging on the phenotypic and genetic resistance responses. An aliquot of the non-selected control population was exposed to AgNPs every passage (that is, for the first time) to compare the MIC in sensitive, repeatedly passaged cells against repeatedly treated cells. Thus, we interpret the observed significant fold increase to result from the AgNP selection that was intentionally imposed rather than the bacterial response to repeated passaging in the non-selected control population, as indicated by the significant impact of the treatment group (P < 0.001) in an analysis of variance (ANOVA) and the non-significant change in MIC (that is, less than a twofold MIC increase) in the non-selected control population (P = 1). Detailed comparisons and statistical analyses are summarized in Table 1 and Supplementary Table 2. Shifts in microbial growth kinetics, including the duration of the lag phase and maximum specific growth rate, also occurred at passage 8 and aligns with the increase in MIC as an additional metric indicating evolution of resistance (Supplementary Discussion 1, Supplementary Figs. 5 and 6 and Supplementary Tables 3–5).
a–c, Resistance profiles in response to AgNPs (black triangles, a), Ag(I) ions (blue triangles, b) and ampicillin (positive control) (purple triangles, c) plotted as fold increase in MIC versus passage number. The non-selected passage control for each condition is included as red circles and pertains to serial passaging of untreated bacteria, with an aliquot being exposed to the antimicrobial of interest each passage (that is, for the first time). The tan box represents the absence of the antimicrobial pressure. Mean ± s.d. of n = 2–3 replicates. The asterisk signifies a greater-than-twofold average increase in the MIC (passages 8–20) that is statistically significant at P < 0.001. The hypermotile E. coli (+IS1) strain developed resistance to AgNPs at a comparable rate to the onset of resistance to ampicillin but not in the case of Ag(I) ions.
The greater-than-twofold MIC increase occurring throughout passages 8–20 suggests resistance evolved to AgNPs24,25 and aligns with other AgNP resistance studies that cover a wide range of sizes and surface chemistries7,8,9,10,11,12. For example, multiple strains of E. coli and Pseudomonas aeruginosa (P. aeruginosa) experienced a fourfold MIC increase upon exposure to AgNPs within 6–13 passages10, while Bacillus subtilis adapted to concentrations of AgNPs and Ag2O-NPs that were at a 1.5-fold MIC increase after 13 passages9. The fold increase observed in our study indicates that E. coli K-12 resistance to AgNPs develops more slowly compared with conventional antibiotics reported in the literature (for example, gentamicin, ampicillin, tetracycline and chloramphenicol), which impart a higher fold increase in the MIC in a shorter time period26,27,28,29. For example, E. coli (ATCC 11775) demonstrates resistance to gentamicin after just one passage and a tenfold MIC increase after ten passages28. The slower evolution of resistance to AgNPs may indicate that the cell–AgNP interaction leading to adaptation is more complex than the simple direct interaction of antibiotics with the cell and is probably due to the range of AgNP modes of action on cellular processes, whereas antibiotics target a single specific target28. Alternatively, the intermittent periods of subculturing on agar may have diminished the effective population size, power of natural selection and mutation supply, rendering the development of AgNP resistance a slower process.
Ampicillin is an antibiotic known to induce resistance in several bacterial species, including E. coli26,29,30 and was evaluated as a positive control treatment in our study. Here, we saw a greater-than-twofold statistically significant increase in the MIC (Fig. 1c) after 8–20 passages with ampicillin (P < 0.001), yet no change in MIC was observed in the non-selected control population (P = 1). Analysis of variance testing further revealed a significant impact between treatment group (ampicillin exposure versus non-selected passage control, P < 0.001) (Table 1 and Supplementary Table 2). Therefore, the observed evolution of ampicillin resistance, using fold change in the MIC as an indicator, upon prolonged exposure to ampicillin validates our methodology and confirms that the increase in fold change after repeated passaging with AgNPs was also due to evolution of resistance. The magnitude of ampicillin resistance observed in this study is however significantly less than has been reported in other ampicillin and antibiotics studies26,29, possibly owing to the different strains of E. coli K-12 used between studies, fitness epistasis (that is, genetic variation within populations) and the experimental evolution approach applied.
No significant change in the MIC (that is, less than a twofold increase) was observed over 20 passages with Ag(I) ions in the hypermotile E. coli (+IS1) (P = 1), or in the non-selected control population (P = 1), indicating that no resistance to Ag(I) ions evolved (Fig. 1b and Table 1). The absence of bacterial resistance to Ag(I) ions was also observed by Panacek et al.10, yet is contrary to other studies that observe resistance to Ag(I) ions8,11,12,14,15, albeit some of the resistance to Ag(I) ions was found to evolve more slowly than for AgNPs12. These differences could be attributed to the use of different bacterial taxa, different methods used to select for resistance and different bacteria growth media containing various halide ion concentrations12,14,15,31. For example, Haefeli et al. and Li et al. found that when there was no sodium chloride present in the system, the MIC of Ag decreased significantly for E. coli and Pseudomonas stutzeri14,15. Additionally, the mixed observations of resistance to Ag(I) ions within the literature could be due to fitness epistasis, that is, genetic variation within populations, coupled with differences in the external environment, both of which can dictate specific evolutionary paths and mutation effects in evolution experiments32,33. In our study, the decreased susceptibility of the hypermotile E. coli (+IS1) to AgNPs compared with Ag(I) ions (controlling for total Ag and Ag(I) content, see Supplementary Discussion 2) suggests a nanoparticle-specific bacterial response similar to the findings of Panacek et al.10. Bacteria probably employ different resistance mechanisms in response to the challenges imposed by different forms of Ag to adapt and mitigate their antimicrobial activity, which we expect given the demonstrated unique antimicrobial mechanisms of AgNPs versus Ag(I) ions34.
Permanent AgNP resistance is conferred by cusS mutation
The heritability and stability of the resistant phenotype was investigated to classify the observed AgNP resistance as reversible or permanent and thus provide some insight into the mechanism conferring resistance. After the AgNP pressure was removed and reintroduced following a few passages of non-selective bacteria growth in broth (Fig. 1a), the strain maintained resistance, suggesting the change was heritable. Thus, the type of resistance conferred can be characterized as permanent and non-reversible35,36, which is corroborated by other studies that found Ag-resistance genes to be constitutively expressed (that is, always turned on) even in the absence of Ag and regardless of form (nanoparticulate or ionic Ag)9,11,12,31,37. This is contrary to the proposal that expression of Ag-resistance genes or a Ag-resistant phenotype should be lost in the absence of Ag owing to fitness costs2,18,38,39. Other studies investigating AgNP and Ag(I) ion adaptations have shown bacterial resistance but have not demonstrated a stable resistant phenotype in the absence of Ag7,40, which could be attributed to the use of different bacterial species and different resistance selection methods, particularly duration and concentration of exposure, of which phenotypic stability is highly dependent25,35.
To validate our conclusion of a permanent genetic mutation imparting the observed stable resistant phenotype, we carried out a detailed genomic analysis using whole-population genome sequencing. Whole-population genome sequencing of the population exposed to repeated AgNP treatment revealed several mutations which arose and remained stable after 25 passages. Specifically, a missense mutation (R292L) in the cusS gene arose at passage 8 and was maintained at all passages thereafter at 100% frequency (Table 2). The presence of other mutations in the ancestor and control populations are explained in the Supplementary Information (see Supplementary Discussion 3). CusS encodes a sensory histidine kinase that is part of a two-component regulatory system with CusR, which activates expression of the cusCFBA periplasmic efflux pump in response to copper or silver exposure41,42. At the amino acid level, this mutation changes the positively charged arginine residue (residue 292) to a non-polar hydrophobic leucine residue (Supplementary Fig. 7). The exact functional effect of this change is unknown and more research is needed to elucidate the structure of the cytoplasmic domain and its functional roles. We speculate that AgNP resistance is imparted through increased silver ion efflux, which may result from autophosphorylation of the CusS cytoplasmic domain caused by this mutation. Residue 292 resides within the active site of the dimerization and the histidine phosphotransfer (DHp) domain, which works with the catalytic and ATP-binding domain to catalyse the transfer of a phosphate group from ATP to the response regulator CusR41,42. Since the cusS mutation rose to fixation over several passages in the treatment line and was absent in the non-selected control line, this genotype is a likely response to AgNP selection and could not have risen to high frequency by genetic drift over this time scale43.
Although this particular cusS mutation does not seem to have been documented, increased AgNP resistance has been observed and attributed to the autophosphorylation or phosphoacceptor residues of CusS, which are essential for endogenous metal resistance in E. coli42. Other work has also identified mutations in cusS as a resistance response to AgNPs and Ag(I) ions, albeit that several of these have been located in different domains (Supplementary Fig. 8)8,44,45.
AgNP resistance depends on bacterial motility
The observed increase in the MIC for the hypermotile E. coli (+IS1) strain upon repeated passaging with AgNPs and not Ag(I) ions suggests a specificity of resistance evolution to AgNPs. To determine whether resistance evolves from interaction with the particle or as a consequence of the experimental system, we pursued evaluation of (i) strain-dependent resistance by investigating a non-motile E. coli strain (ΔflhD::kan) and (ii) particle aggregation under experimental conditions. Motility, in particular, is a key fitness trait for many bacteria and confers the selective advantage of enabling cells to escape from poor conditions and search for more favourable environments (for example, reducing exposure to an antimicrobial, increasing nutrient availability and relocating to unpopulated areas)46,47.
Unlike the hypermotile strain, the average MIC fold increase for the non-motile E. coli strain (ΔflhD::kan) was not significantly greater than a twofold increase (P = 0.88) during repeated passaging with AgNPs (Fig. 2a and Supplementary Fig. 3) nor for the non-selected control population (P = 0.99), indicating no resistance had evolved. Bacterial resistance also did not evolve towards ampicillin, as there was no significant MIC fold increase (P = 1) (that is, less than twofold increase) upon repeated passaging with ampicillin (Fig. 2b). Detailed comparisons and statistical analyses are summarized in Table 1 and Supplementary Table 2. Additionally, in both cases, the duration of the lag phase and maximum specific growth rate did not show any change (Supplementary Figs. 5 and 6 and Supplementary Tables 4 and 5). The lack of observed resistance in both treatments is expected because AgNP resistance typically influences bacteria features and functions that are also responsive to common antibiotics (for example, cell wall synthesis, protein synthesis and nucleic acid synthesis)7. The difference in resistance response between E. coli strains (hypermotile and non-motile) to AgNPs suggests that AgNP resistance is a strain-specific response and that differences in motility may have facilitated different interactions between the bacteria and AgNPs. However, the non-motile strain contains a kanamycin selection marker that was inserted to knock out the regulatory region of flhD (Supplementary Table 1), which inactivates the entire flhDC operon that encodes the master regulator of flagellum biosynthesis48,49. This marker confers resistance to kanamycin and could have also shaped the differential resistance response to ampicillin and AgNPs. We also recognize that the insertion of a IS1 element in the putative diguanylate cyclase (Table 2) in the hypermotile strain may not only contribute to phenotypes related to flagellum-mediated motility but can be pleiotropic and affect things such as cell adhesion, biofilm formation, growth rate and population size50. This would suggest the possibility that, in addition to or instead of motility, other strain-specific differences could have played a role in the evolution of resistance and would require further investigation of phenotypic traits imparting resistance. Still, this observation is in line with other studies that have noted enhanced resistance to antibiotics in organisms that swarm, surf or have increased motility51,52,53. Other researchers have also identified plasmid-mediated mechanisms of resistance, with the plasmid being associated with motility and biofilm formation54. Unlike other studies which have simply used a motility and/or flagellin inhibitor such as pomegranate rind extract to demonstrate resistance suppression10,55, our study is unique in that we used a knockout E. coli K-12 strain displaying a non-motile phenotype to demonstrate lack of resistance onset.
a,b, Resistance profiles in response to AgNPs (black triangles, a) and ampicillin (positive control) (purple triangles, b) plotted as fold increase in MIC versus passage number. The passage control is included as red circles. Average ± s.d.of n = 2–3 replicates. The non-motile E. coli (ΔflhD::kan) strain did not develop resistance to AgNPs nor to ampicillin. (Note that these experiments were halted after passage 13 because of COVID-19 laboratory closures. Resistance evolved in the hypermotile strain, Fig. 1, around passage 8.)
Particle aggregation will influence particle–bacteria interactions and has been shown to result from experimental media34 and proteins produced by the bacteria (for example, flagellin)10. Therefore, we evaluated particle aggregation as a function of bacteria strain and susceptibility, total AgNP concentration and time in the experimental media. To provide insight into their potential influence on resistance evolution, UV–vis–NIR and dynamic light scattering (DLS) data of AgNPs were collected after incubation in (1) growth media alone and (2) growth media with each E. coli strain (that is, hyper- and non-motile), including both susceptible (20 untreated passages) and resistant (20 treated passages) bacteria of the hypermotile strain. Samples were taken at two time points over the duration of one MIC assay (that is, a mid-point and end-point at 14 and 21 h, respectively) and three AgNP concentrations (6.3, 12.5 and 25 µg ml−1 AgNP) to see if time and concentration elicited differential aggregation. Generally, an increase in particle size will shift the LSPR peak obtained with UV–vis–NIR towards longer wavelengths (red-shift), result in peak broadening and decrease the intensity of the peak56. These spectroscopic effects are mimicked when smaller particles aggregate. Additionally, changes in hydrodynamic diameter (Dh) observed with DLS can indicate changes in the colloid size and suggest particle aggregation57,58. Raw aliquots of each post-incubation condition were studied, as well as samples that were filtered with a 0.45 μm polyvinylidene fluoride (PVDF) membrane filter to isolate the AgNPs (that is, remove bacterial cells and eliminate their potential influence on changes in Dh). The filtrate was then analysed by DLS and UV–vis–NIR spectroscopy, with the presence of the LSPR peak suggesting that AgNPs remained in the filtrate (Fig. 3). However, any aggregates greater than 450 nm, if formed, would also have been removed (AgNP loss is reported in Supplementary Fig. 9). Both samples before and after filtration are included in Table 3 to show the change in particle size by DLS upon removal of the bacteria. Only data corresponding to the 21-h time point and AgNP concentration of 12.5 µg ml−1 are reported and discussed below (Fig. 3 and Table 3) as the other conditions showed similar aggregation behaviour.
Aggregation of AgNPs (12.5 µg ml−1 [AgNP]) after 21 h incubation in broth and in the presence of E. coli before and after filtering with a 0.45-μm PVDF membrane filter to remove bacterial cells. a,c, The increase in particle size (hydrodynamic diameter) is revealed by DLS after incubation with E. coli. b,d, The UV–vis–NIR absorption spectra of the AgNPs in arbitrary units (a.u.) reveals broadening of the characteristic LSPR at λmax ≈ 420 nm after incubation with E. coli to indicate that aggregation is occurring. a,b, AgNPs in the presence of non-motile E. coli (AgNP-sensitive only). c,d, AgNPs in the presence of AgNP-sensitive and AgNP-resistant strains (after passage 20) of hypermotile E. coli, which all exhibit similar aggregation behaviour. Plots show the average of three replicates (for DLS, 20 scans per replicate) for bacteria-containing samples or two replicates for AgNP-only controls (in broth). Note: aggregation of AgNPs at 14 h (and at 6.3 and 25 µg ml−1 [AgNP]) was similar to the data presented here and thus was not included.
After incubation in growth media alone, the AgNPs had a Dh of approximately 60 nm (Table 3), which is larger than the inorganic core size determined by TEM (d = 27.2 ± 4.8 nm) and is expected given that the PEGSH ligand and the solvent layer surrounding the particle also contribute to the measured Dh (ref. 59). As a control, the AgNPs were filtered from growth media alone to ensure that the process of filtering did not alter the size distribution of the AgNPs. The Dh and full-width at half-maximum (FWHM) of the DLS and LSPR peaks remained consistent before and after filtering, indicating that there was no change in particle size due to the filtering process (Table 3 and Fig. 3).
Comparing the size of AgNPs in broth alone to AgNPs in the presence of E. coli after filtration indicates that particle aggregation occurred to a similar extent under all exposure conditions, including for both the hypermotile and non-motile strains as well as in the case of both susceptible and resistant phenotypes (Table 3). E. coli were also filtered from growth media alone to ensure that the 0.45 μm pore size was sufficient to remove all bacteria (length of 1–2 μm, diameter of 0.25 μm) (Supplementary Fig. 10). Particle Dh increased from 60 nm to a range of 110–160 nm for filtered samples (Table 3 and Fig. 3a,c), which is corroborated by the LSPR peak broadening (increase in FWHM) and red-shifting, indicating a shift to larger particle diameters (Table 3 and Fig. 3b,d). Since all strains induce similar aggregation behaviour (that is, aggregation is independent of strain susceptibility and bacterial motility) aggregation alone cannot be responsible for the differences in resistance emerging in the hyper- and non-motile E. coli. Currently unrealized, alternative mechanisms of interaction around bacterial motility (for example, physical distancing or relocating away from the AgNPs to reduce exposure or increase nutrient availability) may be at play in inducing these differential AgNP resistance profiles. Further, the non-motile strain has the regulatory region of flhD knocked out, which inactivates the entire flhDC operon that encodes the master regulator of flagellum biosynthesis48,49, meaning there should be no flagellin produced (that is, the major protein hypothesized as being responsible for AgNP aggregation10). Since aggregation occurs in the absence of flagellin (that is, in the non-motile strain experimental conditions), there may be other proteins (for example, pili, outer membrane proteins, ribosomal proteins and enzymes involved in cellular metabolism released upon cell death10) present in bacteria that cause aggregation.
Conclusion
Findings from this study present different resistance responses for hypermotile and non-motile E. coli strains after repeated exposure to subinhibitory concentrations of Ag(I) ions and AgNPs. AgNP resistance (that is, a statistically significant greater-than-twofold increase in the MIC) was found in hypermotile E. coli and remained stable, with no reversion back to AgNP sensitivity, while non-motile E. coli demonstrated no resistance. A permanent mutation in cusS was found in the hypermotile E.coli strain and suggests a direct mechanism of resistance through increased silver ion efflux that may be co-mediated or enhanced by a motile phenotype. These findings provide critical insight into bacterial development of resistance to AgNPs, underscoring the potential role of bacterial motility. Identifying that bacteria motility may play an independent role from particle aggregation is an exciting result because it directs future research to uncover detailed mechanism(s) of resistance. In addition, the findings from this study are promising for reducing the burden of pathogenic diseases and infection because there are many non-motile, clinically relevant, drug-resistant bacterial strains that are potential targets for AgNP treatments. The Center for Disease Control and Prevention lists Clostridioides difficile, P. aeruginosa, Staphylococcus aureus and various types of Acinetobacter and Streptococci as urgent and serious threats60, all of which are non-motile, or in the case of P. aeruginosa, may emerge as non-motile during late and chronic infections61. Our data suggest that AgNPs could be effective and remain effective in non-motile strains, with no resistance or, potentially, delayed resistance emerging.
It should be noted that even in the absence of observed resistance, it is possible that resistance could evolve over longer time scales (for example, more than 20 passages) or when multiple independent replicate populations are used. Uncovering these possibilities will require additional investigation. We recognize that the stochastic nature of evolutionary adaptation may not impart the same phenotype or involve the same genes, and thus may present multiple mechanisms and levels of resistance, such that replication and understanding the impact of different mutations arising in different populations is important. Possible additional mechanisms of resistance may be uncovered in future work that incorporates several replicate populations. Additionally, predicting how frequently the cusS mutation or other mutations occur to impart resistance can also be achieved by studying multiple replicate populations. Different exposure concentrations, periods of exposure and the presence of other stressors will also influence evolution of resistance. Furthermore, our study examined one type of AgNP (that is, one size, shape and surface ligand) within one growth medium; additional research is needed to establish global conclusions that account for the wide variability of nanoparticle-dependent biological interactions and underlying genetic responses, as well as media components and concentrations that influence particle behaviour in suspension. The observed absence of bacterial resistance to ionic silver in motile strains and the evolution of resistance upon exposure to AgNPs indicate a mechanism specific to nanoparticulate silver and provide an opportunity to evade resistance through particle design. The particle parameters—size, shape and surface chemistry—can be used to modulate differentially the identified AgNP properties that govern their efficacy and minimal resistance in different bacteria. Additionally, and particularly for motile strains, the focus of future work could be on designing the particle to maximize efficacy (for example, high, sustained ion release) under given use conditions or designing for a high biological fitness cost.
Finally, our findings caution against the ubiquitous use of AgNPs in consumer products (for example, washing machines, textiles and paints) that result in potentially high human and environmental exposures. To limit and not contribute to the growing global resistance challenge, AgNP use should be reserved for those applications necessitating antimicrobial function, such as in the health care sector and for medical devices where resistant bacteria cause life-threatening situations.
Methods
Chemicals
Silver nitrate (AgNO3, 99.9999%), tannic acid (purissimum grade), sodium citrate tribasic dehydrate (citrate, ≥99%), ampicillin sodium salt, nitric acid (HNO3, >99.999%, trace metal basis) and isopropyl alcohol (IPA, 99.90%) were purchased from Sigma-Aldrich. Hydrochloric acid (HCl, >99.999%, trace metal basis), acetonitrile (ACN, 99.8%), sodium chloride (NaCl, 99.6%), potassium phosphate monobasic (KH2PO4), Luria–Bertani broth (LB, Miller), BD Bacto dehydrated agar and glycerol were purchased from Fisher Scientific. Poly(ethylene glycol) methyl ether thiol (PEGSH, average molecular weight 1,000 g mol−1) was obtained from Laysan Bio, Inc. Deuterium oxide (D2O, 99.9%) was obtained from Cambridge Isotope Laboratories. All reagents were used as received unless otherwise indicated. All chemicals were dissolved in deionized (DI) water obtained from a Milli-Q ultrapure water purification system. NANOpure (Thermo Scientific, >18.2 MΩ cm) water was used in the preparation of all solutions associated with AgNP synthesis. Before use, all glassware and Teflon-coated stir bars were washed with aqua regia (3/1 ratio of concentrated HCl and HNO3 by volume) and rinsed thoroughly with water. Caution: aqua regia is highly toxic and corrosive and requires proper personal protective equipment. Aqua regia should be handled in a fume hood only.
Bacterial strains and cultivation
The bacterial strains used in this work were the wild-type E. coli K-12 MG1655(Seq) strain (CGSC no. 7740) and the E. coli K-12 JW1881 ΔflhD::kan strain from the Keio Collection (CGSC no. 11772), both obtained from the Coli Genetic Stock Center at Yale University (Supplementary Table 1). Since E. coli MG1655 has IS1 elements in the flhDC promoter and a putative diguanylate cyclase dgcJ, it is hypermotile with hyper-flhDC expression48,49 and will be referred to as the hypermotile E. coli (+IS1) strain. E. coli JW1881 ΔflhD::kan has a deletion in the regulatory region of flhD. This deletion inactivates the entire flhDC operon with no flhDC expression48,49 and thus the strain will be referred to as the non-motile E. coli (ΔflhD::kan) strain. Microorganism stock solutions were prepared in LB broth supplemented with 25% glycerol and stored at −80 °C. Microorganisms were cultured overnight in LB broth at 37 °C and 150 r.p.m. Overnight cultures were re-inoculated in fresh medium, grown to exponential phase and diluted 1000-fold in LB to obtain approximately 106 colony-forming units (CFU) ml−1.
Synthesis and ligand exchange of AgNPs
Citrate-capped AgNPs (diameter, d = 27.2 ± 4.8 nm) were synthesized according to a previously reported procedure20,21,22. Briefly, a 100 ml aqueous solution of citrate (5.00 mM) and tannic acid (0.40 mM) was brought to reflux in a 250-ml, three-neck round-bottomed flask while stirring at a rate of 340 r.p.m. The molar ratio of tannic acid to citrate was 0.08/1 to achieve ~25 nm AgNPs22. Once 100 °C was reached and a rapid drip rate was achieved (drip rate ~1 s−1), 1.00 ml AgNO3 (25.00 mM) was quickly injected. The reaction mixture changed from a transparent, pale yellow colour (because of the tannic acid) to a dark, golden brown within 1 min. The resulting AgNP solution was allowed to reflux for 5 min before it was removed from heat and cooled to room temperature by sitting on the benchtop. The citrate-capped AgNPs were concentrated and washed to remove excess citrate and tannic acid by centrifuging 1.50-ml aliquots in 1.50-ml centrifuge tubes at ~20,000 relative centrifugal force for 6 min (Eppendorf 5424 centrifuge). The supernatants were removed and the pellet was resuspended in another 1.50-ml aliquot of AgNP solution from the round-bottomed flask and centrifuged again. After the second removal of the supernatants, the particles were resuspended in 1.00 ml H2O. The prepared citrate-capped AgNPs, as 1.00 ml H2O suspensions, were functionalized with PEGSH (average molecular weight 1,000 g mol−1) using mass action ligand exchange (Scheme S1 in ref. 20). Here, 12.90 mM PEGSH was added to each tube containing the particles in 1.00 ml H2O. These mixtures were vortexed and then placed on a temperature-controlled mixer (Eppendorf R Thermomixer) in the dark at 400 r.p.m. at 25 °C for 24 h. Immediately after mixing, the PEGSH-coated AgNPs in each tube were centrifuged, resuspended in 1.00 ml H2O and transferred to a clean glass media bottle wrapped in aluminium foil for refrigerated storage (~4 °C). This synthesis and ligand exchange process was repeated to obtain approximately ten batches of AgNPs over a span of 3 days and pooled into the same glass media bottle for storage. This was completed at the start of the bacterial resistance experiments (see below).
At the time of each bacterial passage (vide infra), the AgNPs were further concentrated and purified with a series of washing steps to remove excess PEGSH and displaced citrate; 1.50-ml aliquots (eight tubes in total) were centrifuged and their supernatants removed. Two of the pellets were combined with 1.00 ml H2O so that a total of four total tubes remained. This process of combining pellets after centrifuging was repeated a further two times, keeping the same 1.00 ml total volume, until one concentrated tube remained. The particles were then washed by centrifuging once more, the supernatant was removed and the particles were resuspended in 850 µl H2O to provide sufficient volume of concentrated AgNPs for the experiments. Of this solution, 10.0 µl were diluted in 500 µl H2O to (i) obtain a concentration that would fall under the upper limit of quantification to measure reliably the initial AgNP concentration by UV–vis–NIR and (ii) provide good resolution for characterizing particle size by TEM. The remaining solution was used in the bacterial resistance experiments and diluted to obtain 400 µg ml−1 AgNPs.
For initial ligand density quantification, the same purification process was repeated on four additional tubes of 1.50-ml aliquots until one tube remained. After the last wash cycle, the supernatant was removed to yield the concentrated pellet of PEGSH-AgNPs which was saved for NMR spectroscopy, TEM and ICP-MS analyses.
AgNP characterization
AgNP solutions at each bacterial passage were characterized by UV–vis–NIR absorption spectroscopy using a Cary 5000 spectrophotometer (Agilent, Inc.). The spectrum baseline was corrected with respect to the H2O spectrum, or in the case of the aggregation experiments, to the spectrum of LB broth. To evaluate the stability of the refrigerated AgNP stock solution over time, NP size distributions were determined by TEM. An aliquot was obtained weekly and diluted with H2O prior to being drop cast onto a Formvar-coated copper TEM grid (Ted Pella, Inc.). Samples were allowed to air dry slowly and then were dried under vacuum overnight before characterization with a Hitachi H-9500 environmental TEM at 300 kV (NanoScale Fabrication and Characterization Facility, Petersen Institute of NanoScience and Engineering, Pittsburgh, PA). NP size distributions were determined from measuring at least 300 NPs (ImageJ v.1.52a, National Institutes of Health, USA) in multiple images collected from various areas of the grid. For the aggregation experiments, AgNP solutions were placed in a disposable plastic cuvette (Fisherbrand). Dynamic light scattering was used to measure the hydrodynamic diameter of AgNPs (Anton Paar Litesizer 500). Spectra were averaged over 20 scans. All NMR measurements were performed on a Bruker 400 Ultrashield magnet with AVANCE III 400 Console (Bruker Bio-spin) at 298 K (Supplementary Figs. 4 and 5 in ref. 20). ICP-MS analysis was performed using an argon flow with a NexION spectrometer (Perkin Elmer, Inc.) (Supplementary Fig. 3 in ref. 20).
Quantifying resistance in response to Ag(I) ions and AgNPs
Initial antimicrobial activity of Ag(I) ions (as AgNO3) and AgNPs, as well as ampicillin as a positive control, were evaluated using standard planktonic MIC measurements. These experiments were carried out in 96-well flat-bottomed microplates (Corning Costar) and done in triplicate. A dispersion of AgNPs or a solution of AgNO3 were serially diluted in H2O and inoculated with 50 μl E. coli (5 × 105 CFU ml−1) so that the total volume of each well was 100 μl. The final tested silver concentrations were 200, 100, 50, 25, 12.5, 6.3, 3.1, 1.6, 0.8 and 0.4 µg ml−1 for AgNPs and 100, 50, 25, 12.5, 6.3, 3.1, 1.6, 0.8, 0.4 and 0.2 µg ml−1 for AgNO3. The final tested ampicillin concentrations were 25, 12.5, 6.3, 3.1, 1.6, 0.8, 0.4, 0.2, 0.1 and 0.05 µg ml−1. Controls, including broth (50 μl) and water (50 μl) alone, broth (50 μl) and water with the antimicrobial agent (50 μl) and water (50 μl) and untreated bacteria (50 μl), were run simultaneously. Cells were incubated at 37 °C for 24 h with medium linear shaking (~500 cycles per minute) in a Microplate Reader (Synergy HTX Multi-Mode, BioTek). Cell growth was monitored by measuring absorbance at 600 nm every 30 min. The mean absorbance and s.d. for at least three biological replicates is reported.
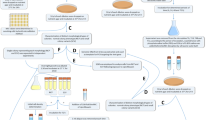
To evaluate evolution of resistance, bacteria were repeatedly exposed to subinhibitory concentrations of Ag(i) ions and AgNPs, as well as ampicillin, in 20 successive passages (note: the experiment for the non-motile strain was interrupted by COVID-19-related shut down of the laboratory, so only 13 passages were completed). We prepared a parent culture that was used to inoculate a serially diluted series of AgNPs ranging from 0.4 to 200 µg ml−1 AgNPs (vide supra). The culture was incubated and after 24 h the MIC concentration was determined to be the lowest concentration of antimicrobial agent that inhibited visible growth of the tested microorganisms. Sub-MIC cultures with surviving bacteria (100 µl taken from each of the first three wells below the MIC) were mixed and then subcultured on LB agar at 37 °C for 16–18 h. A single colony was used for inoculum preparation at a density of 106 CFU ml−1 in the next passage where they were exposed to the second serially diluted series of AgNPs and incubated overnight. The entire procedure described above, from the initial inoculation to the preparation of the new inoculum, was considered to constitute one passage for the development of bacterial resistance. The next day the MIC (which remained the same or increased) was determined and the sub-MIC cultures from the latest passage were used to inoculate a new series of diluted AgNPs. The process continued for up to 20 passages, a standard timeframe for observing resistance development in evolutionary experiments10,29,35,63,64 and over time bacteria began surviving at higher concentrations. Dispersions of AgNPs and solutions of ampicillin having higher concentrations (for example, 400–800 µg ml−1) were used when the MIC increased past the initial concentration range (that is, when resistance evolved). Non-selected control populations, that is, untreated bacteria successively cultured in the absence of any antimicrobial agents and exposed to the antimicrobial agent for the first time at each passage, were also included. In all cases where resistance developed, the stability of the resistant phenotype was investigated by removing the selective pressure (that is, AgNPs or ampicillin) and re-introducing it after a few passages of non-selective bacteria growth in broth (that is, without the antimicrobial agent) to see if the MIC remained elevated (that is, displayed a stable resistant phenotype) or decreased back to pre-exposure levels (that is, returned to its susceptible phenotype).
Fold increase in the MIC (equations (1) and (2)) was calculated by dividing the average MIC for the passage of interest X by the average MIC of the initially susceptible bacteria:
where ‘experimental’ denotes that each passage had an antimicrobial exposure (for example, passage 10 = 10 passages of antimicrobial exposure), whereas ‘control’ is the non-selected repeated passaging control. At each new passage, the non-selected bacteria population was exposed to the antimicrobial for the first time (for example, passage 10 = 9 passages untreated, 1 passage of antimicrobial exposure on the 10th passage). The ‘initial’ MICs corresponding to the experimental and control were always set equal. Those values were obtained from the first passage, except for two of the conditions (that is, AgNPs and Ag(I) ions in the case of the hypermotile strain) where those values were obtained as the average of the first five passages because of the high variability in the initial MIC data. Additionally, multiple skip wells (that is, wells having no bacteria growth in between wells at a higher and lower antimicrobial concentration having bacteria growth65,66) appeared several times throughout the MIC assays following AgNP exposure and not in the Ag(I) ions or ampicillin treatments. The observation of skip wells in the case of AgNPs is probably due to the nature of the colloidal solution and aggregation behaviour when bacteria and growth media are added. It is possible that differential aggregation occurs between wells, particularly at higher AgNP concentrations67. However, an alternative explanation for the presence of skip wells could be attributed to heteroresistance, which is the formation of resistant subpopulations, where subpopulations exhibit a range of susceptibilities to antimicrobial agents65,66. Replicates that contained more than one skip well were excluded from the data analysis as they are considered uninterpretable65,68,69. Replicates containing just one skip well were included by considering the well above the skip well (that is, the well at the higher antimicrobial concentration) as the lowest dilution having no bacteria growth and therefore the MIC65,68,69.
Genomic analysis
Whole-population genome sequencing was performed to identify genomic variants associated with the resistance observed in the AgNP treatment populations compared to the ancestor and non-selected control population. DNA was extracted from the ancestral E. coli K-12 MG1655 stock and both control and treatment populations at multiple time points, using samples preserved following each MIC assay. These included: (i) passage 8, which represents the earliest observation of the resistant phenotype in the treated populations, (ii) passage 20, which represents the end of the repeated exposure and persistent resistant phenotype in the treated populations, (iii) passage 24, which represents the absence of AgNP pressure and (iv) passage 25, which represents the re-introduction of AgNP pressure in the treated populations. Genomic DNA was extracted from overnight cultures using a Qiagen DNeasy Tissue Kit according to manufacturer’s instructions (Qiagen). Illumina library construction and sequencing were conducted using the Illumina Nextera DNA Sample Prep Kit with 150-bp paired-end reads and libraries were sequenced on the NextSeq 550 sequencing platform (Illumina, San Diego, CA). Whole-population genome sequencing was performed at the Microbial Genome Sequencing Centre. Variant calling and the identification of mutations (for example, single nucleotide polymorphisms, insertion-deletion polymorphisms) was achieved by aligning AgNP variant sequences to the reference E. coli K-12 MG1655 genome70 using the breseq v.0.35.4 pipeline62.
Monitoring AgNP aggregation
AgNPs at three different concentrations (6.3, 12.5 and 25 µg ml−1) were incubated in LB broth and in the presence of both E. coli strains at 37 °C for 14 and 21 h in 96-well microplates. Twelve wells were pooled together to comprise one sample, which was analysed by DLS and UV–vis–NIR spectroscopy. The samples were then filtered using a 33-mm Millex-HV Durapore syringe filter with a 0.45 µm pore size and hydrophilic PVDF membrane (Millipore–Sigma). The filtrate was then analysed by DLS and UV–vis–NIR spectroscopy. This experiment was carried out in triplicate and was also repeated in the presence of AgNP-resistant bacteria.
Statistics
All statistical analyses were carried out in R (v.4.0.0 (Arbor Day) released 2020, University of Auckland, New Zealand). A two-way ANOVA was used to analyse the difference in fold increases in the MIC between treatment group and passage number under different exposure conditions and bacteria strains to determine if either treatment group or passage number impact the fold increase in the MIC. Experimental and passage control data were pooled together to increase the power of the statistical analyses. The level of this significance was then probed for individual groups using a paired Wilcoxon test with at least a sample size of n = 8. Specifically, the fold increase in the MIC at the end of the experiment between treatment groups and versus the initial fold increase in the MIC was compared. Similar analyses were carried out for maximum specific growth rate and duration of the lag phase. A backwards power analysis revealed that with our sample size we could determine statistical significance with an alpha = 0.05 and a power of 80%. This translates to being able to detect significant differences in MIC for the hypermotile strain and passage control of at least 0.55 (that is, n = 52, s.d. = 1.39) and 0.63 (that is, n = 40, s.d. = 1.39), respectively. When comparing only the last five passages (that is, n = 10, s.d. = 1.68), we can detect significance at a difference of size 1.7. Alternatively, using the same power and significance for the non-motile data set requires at least a difference of 0.96 to be present before a significant difference can be detected.
Data availability
Figures 1–3, Tables 1–3 and all compiled supplementary figures and tables were composed from raw data that are available from the corresponding author. Whole-population genome sequencing data is available through NCBI SRA BioProject ID PRJNA729974.
References
Mijnendonckx, K., Leys, N., Mahillon, J., Silver, S. & Van Houdt, R. Antimicrobial silver: uses, toxicity and potential for resistance. BioMetals 26, 609–621 (2013).
Molling, J. W., Seezink, J. W., Teunissen, B. E., Muijrers-Chen, I. & Borm, P. J. Comparative performance of a panel of commercially available antimicrobial nanocoatings in Europe. Nanotechnol. Sci. Appl. 7, 97–104 (2014).
Silver nanoparticles market by application (electronics & electrical, healthcare, food & beverages, textiles) and segment forecasts to 2022. GrandView Research (2015); https://www.grandviewresearch.com/industry-analysis/silver-nanoparticles-market
Panacek, A. et al. Silver colloid nanoparticles: synthesis, characterization, and their antibacterial activity. J. Phys. Chem. B 110, 16248–16253 (2006).
Franci, G. et al. Silver nanoparticles as potential antibacterial agents. Molecules 20, 8856–8874 (2015).
Duran, N. et al. Silver nanoparticles: a new view on mechanistic aspects on antimicrobial activity. Nanomedicine 12, 789–799 (2016).
Lara, H. H., Ayala-Nunez, N. V., Turrent, L. & Padilla, C. R. Bactericidal effect of silver nanoparticles against multidrug-resistant bacteria. World J. Microbiol. Biotechnol. 26, 615–621 (2010).
Graves, J. L. et al. Rapid evolution of silver nanoparticle resistance in Escherichia coli. Front. Genet. 6, 42 (2015).
Gunawan, C., Teoh, W. Y., Marquis, C. P. & Amal, R. Induced adaptation of Bacillus sp. to antimicrobial nanosilver. Small 9, 3554–3560 (2013).
Panacek, A. et al. Bacterial resistance to silver nanoparticles and how to overcome it. Nat. Nano. 13, 65–71 (2018).
Losasso, C. et al. Antibacterial activity of silver nanoparticles: sensitivity of different Salmonella serovars. Front. Microbiol. 5, 227 (2014).
Valentin, E. et al. Heritable nanosilver resistance in priority pathogen: a unique genetic adaptation and comparison with ionic silver and antibiotics. Nanoscale 12, 2384–2392 (2020).
Gunawan, C. et al. Widespread and indiscriminate nanosilver use: genuine potential for microbial resistance. ACS Nano 11, 3438–3445 (2017).
Haefeli, C., Franklin, C. & Hardy, K. Plasmid-determined silver resistance in Pseudomonas stutzeri isolated from a silver mine. J. Bacteriol. 158, 389–392 (1984).
Li, X. Z., Nikaido, H. & Williams, K. E. Silver-resistant mutants of Escherichia coli display active efflux of Ag+ and are deficient in porins. J. Bacteriol. 179, 6127–6132 (1997).
Gupta, A., Matsui, K., Lo, J. F. & Silver, S. Molecular basis for resistance to silver cations in Salmonella. Nat. Med. 5, 183–188 (1999).
Silver, S. Bacterial silver resistance: molecular biology and uses and misuses of silver compounds. FEMS Microbiol. Rev. 27, 341–353 (2003).
Pelgrift, R. Y. & Friedman, A. J. Nanotechnology as a therapeutic tool to combat microbial resistance. Adv. Drug Deliv. Rev. 65, 1803–1815 (2013).
Wang, L., Hu, C. & Shao, L. The antimicrobial activity of nanoparticles: present situation and prospects for the future. Int J. Nanomed. 12, 1227–1249 (2017).
Johnston, K. A. et al. Impacts of broth chemistry on silver ion release, surface chemistry composition, and bacterial cytotoxicity of silver nanoparticles. ES: Nano 5, 304–312 (2018).
Johnston, K. A., Smith, A. M., Marbella, L. E. & Millstone, J. E. Impact of as-synthesized ligands and low-oxygen conditions on silver nanoparticle surface functionalized. Langmuir 32, 3820–3826 (2016).
Bastus, N. G., Merkoci, F., Piella, J. & Puntes, V. Synthesis of highly monodisperse citrate-stabilized silver nanoparticles of up to 200 nm: kinetic control and catalytic properties. Chem. Mater. 26, 2836–2846 (2014).
Ivanova, E. P., Bazaka, K. & Crawford, R. J. in New Functional Biomaterials for Medicine and Healthcare Ch. 3, 71–99 (Woodhead, 2014).
Olofsson, S. K. & Cars, O. Optimizing drug exposure to minimize selection of antibiotic resistance. Clin. Infect. Dis. 45, S129–S136 (2007).
Gullberg, E. et al. Selection of resistant bacteria at very low antibiotic concentrations. PLoS Pathog. 7, e1002158 (2011).
Adam, M., Murali, B., Glenn, N. O. & Potter, S. S. Epigenetic inheritance based evolution of antibiotic resistance in bacteria. BMC Evol. Biol. 8, 52 (2008).
George, A. M. & Levy, S. B. Amplifiable resistance to tetracycline, chloramphenicol, and other antibiotics in Escherichia coli: involvement of a non-plasmid-determined efflux of tetracycline. J. Bacteriol. 155, 531–540 (1983).
Zhao, Y. et al. Small molecule-capped gold nanoparticles as potent antibacterial agents that target Gram-negative bacteria. J. Am. Chem. Soc. 132, 12349–12356 (2010).
Ji, F. et al. Tetrabromobisphenol A (TBBPA) exhibits specific antimicrobial activity against Gram-positive bacteria without detectable resistance. Chem. Commun. 53, 3512–3515 (2017).
Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Susceptibility Testing: Twentieth Informational Supplement Vol. 30(1) (Approved Standard M100-S20, Clinical and Laboratory Standards Institute, 2010).
Silver, S. in Molecular Biology, Pathogenicity, and Ecology of Bacterial Plasmids (eds Levy, S. B. et al.) 179–189 (Plenum, 1981).
Flynn, K. M., Cooper, T. F., Moore, F. B. G. & Cooper, V. S. The environment affects epistatic interactions to alter the topology of an empirical fitness landscape. PLoS Genet. 9, e1003426 (2013).
Hall, A. E. et al. Environment changes epistasis to alter trade-offs along alternative evolutionary paths. Evolution 73, 2094–2105 (2019).
Stabryla, L. M., Johnston, K. A., Millstone, J. E. & Gilbertson, L. M. Emerging investigator series: it’s not all about the ion: support for particle-specific contributions to silver nanoparticle antimicrobial activity. ES: Nano 5, 2047–2068 (2018).
Sandoval-Motta, S. & Aldana, M. Adaptive resistance to antibiotics in bacteria: a systems biology perspective. WIREs Syst. Biol. Med 8, 253–267 (2016).
Normark, B. H. & Normark, S. Evolution and spread of antibiotic resistance. J. Intern. Med. 252, 91–106 (2002).
Lok, C.-N. et al. Proteomic identification of the cus system as a major determinant of constitutive Escherichia coli silver resistance of chromosomal origin. J. Proteome Res. 7, 2351–2356 (2008).
Knetsch, M. L. & Koole, L. H. New strategies in the development of antimicrobial coatings: the example of increasing usage of silver and silver nanoparticles. Polymers 3, 340–366 (2011).
Chopra, I. The increasing use of silver-based products as antimicrobial agents: a useful development or a cause for concern? J. Antimicrob. Chemother. 59, 587–590 (2007).
Bridges, K., Kidson, A., Lowbury, E. J. L. & Wilkins, M. D. Gentamicin- and silver-resistant Pseudomonas in a burns unit. Brit. Med. J. 1, 446–449 (1979).
Gudipaty, S. A. & McEvoy, M. M. The histidine kinase CusS senses silver ions through direct binding by its sensor domain. Chem. Biochem. 1844, 1656–1661 (2014).
Affandi, T. & McEvoy, M. M. Mechanism of metal ion-induced activation of a two-component sensor kinase. Biochem. J. 476, 115–135 (2019).
Cooper, V. S. Experimental evolution as a high-throughput screen for genetic adaptations. mSphere 3, e00121–18 (2018).
Tajkarimi, M. et al. Selection for ionic- confers silver nanoparticle resistance in Escherichia coli. JSM Nanotechnol. Nanomed. 5, 1047 (2017).
Randall, C. P., Gupta, A., Jackson, N., Busse, D. & O’Neill, A. J. Silver resistance in Gram-negative bacteria: a dissection of endogenous and exogenous mechanisms. J. Antimicrob. Chemother. 70, 1037–1046 (2015).
Koskella, B., Taylor, T. B., Bates, J. & Buckling, A. Using experimental evolution to explore natural patterns between bacterial motility and resistance to bacteriophages. ISME J. 5, 1809–1817 (2011).
Samad, T. et al. Swimming bacteria promote dispersal of non-motile Staphylococcal species. ISME J. 11, 1933–1937 (2017).
Gauger, E. J. et al. Role of motility and the flhDC operon in Escherichia coli MG1655 colonization of the mouse intestine. Infect. Immun. 75, 3315–3324 (2007).
Barker, C. S., Prub, B. M. & Matsumura, P. Increased motility of Escherichia coli by insertion sequence element integration into the regulatory region of the flhD operon. J. Bacteriol. 186, 7529–7537 (2004).
Sanchez-Torres, V., Hu, H. & Wood, T. K. GGDEF proteins YeaI, YedQ, and YfiN reduce early biofilm formation and swimming motility in Escherichia coli. Appl. Microbiol. Biotechnol. 90, 651–658 (2011).
Butler, M. T., Wang, Q. & Harshey, R. M. Cell density and mobility protect swarming bacteria against antibiotics. Proc. Natl Acad. Sci. USA 107, 3776–3781 (2010).
Lai, S., Tremblay, J. & Deziel, E. Swarming motility: a multicellular behaviour conferring antimicrobial resistance. Environ. Microbiol. 11, 126–136 (2009).
Sun, E. et al. Broad-spectrum adaptive antibiotic resistance associated with Pseudomonas aeruginosa mucin-dependent surfing motility. Antimicrob. Agents Chemother. 62, e00848–18 (2018).
Zhang, H. et al. Stress resistance, motility and biofilm formation mediated by a 25kb plasmid pLMSZ08 in Listeria monocytogenes. Food Control 94, 345–352 (2018).
Asadishad, B., Hidalgo, G. & Tufenkji, N. Pomegranate materials inhibit flagellin gene expression and flagellar-propelled motility of uropathogenic Escherichia coli strain CFT073. FEMS Microbiol. Lett. 334, 87–94 (2012).
Paramelle, D. et al. A rapid method to estimate the concentration of citrate capped silver nanoparticles from UV-visible light spectra. Analyst 139, 4855–4861 (2014).
Zhang, W., Crittenden, J., Li, K. & Chen, Y. Attachment efficiency of nanoparticle aggregation in aqueous dispersions: modeling and experimental validation. Environ. Sci. Technol. 46, 7054–7062 (2012).
Li, X., Lenhart, J. J. & Walker, H. W. Dissolution-accompanied aggregation kinetics of silver nanoparticles. Langmuir 26, 16690–16698 (2010).
Ma, R. et al. Size-controlled dissolution of organic-coated silver nanoparticles. Environ. Sci. Technol. 46, 752–759 (2012).
Biggest Threats and Data (Centers for Disease Control and Prevention, 2020); https://www.cdc.gov/drugresistance/biggest-threats.html
Demirdjian, S. et al. Phosphatidylinositol-(3,4,5)-trisphosphate induces phagocytosis of nonmotile Pseudomonas aeruginosa. Infect. Immun. 86, 215–218 (2018).
Deatherage, D. E. & Barrick, J. E. Identification of mutations in laboratory-evolved microbes from next-generation sequencing data using breseq. Methods Mol. Biol. 1151, 165–188 (2014).
Narayanaswamy, V. P. et al. In vitro activity of novel glycopolymer against clinical isolates of multidrug-resistant Staphylococcus aureus. PLoS One 13, e0191522 (2018).
Li, X. et al. Functional gold nanoparticles as potent antimicrobial agents against multi-drug-resistant bacteria. ACS Nano 8, 10682–10686 (2014).
Landman, D., Salamera, J. & Quale, J. Irreproducible and uninterpretable polymyxin B MICs for Enterobacter cloacae and Enterobacter aerogenes. J. Clin. Microbiol. 51, 4106–4111 (2013).
El-Halfawy, O. M. & Valvano, M. A. Antimicrobial heteroresistance: an emerging field in need of clarity. Clin. Microbiol. Rev. 28, 191–207 (2015).
Baalousha, M. et al. The concentration-dependent behavior of nanoparticles. Environ. Chem. 13, 1–3 (2015).
Lecture 11: Antimicrobial Susceptibility Testing—Broth Dilution. Online video clip. Technical Univ. Denmark (n.d); https://www.coursera.org/lecture/antimicrobial-resistance/lecture-11-antimicrobial-susceptibility-testing-broth-dilution-VeNw0
Clinical and Laboratory Standards Institute. Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria that Grow Aerobically 9th edn, Vol. 32(2) (Approved Standard M7-A9, Clinical and Laboratory Standards Institute, 2012).
Riley, M. et al. Escherichia coli K-12: a cooperatively developed annotation snapshot—2005. Nucleic Acids Res. 34, 1–9 (2006).
Acknowledgements
L.M.S. and L.M.G. acknowledge the generous support from the Department of Defense (DoD) through the National Defense Science and Engineering Graduate Fellowship (NDSEG) Program, the 3M Non-Tenured Faculty Award and the Department of Civil and Environmental Engineering at the University of Pittsburgh. Additionally, K.A.J., N.A.D. and J.E.M. thank the Research Corporation for Science Advancement and the Department of Chemistry at the University of Pittsburgh for their support in carrying out this research. We thank M. J. Fritz for her assistance with the DNA extractions prior to whole-population genome sequencing.
Author information
Authors and Affiliations
Contributions
L.M.S. and L.M.G. conceived the project and designed the evolutionary experiments dealing with bacteria resistance to AgNPs and aggregation. L.M.S. carried out the microbiological and evolutionary experiments, the synthesis and long-term stability study of AgNPs, the aggregation experiments, the microbial growth kinetics analyses and statistical analyses. K.A.J. and N.A.D. were responsible for the characterizations of the AgNPs (that is, ICP-MS for Ag atom concentration and NMR for ligand density and TEM for particle size, respectively). V.S.C. conducted the genomic variant analysis. S.-J.H. contributed guidance on the microbiological and evolutionary experiments, the analysis of microbial growth kinetics and the statistical analyses. J.E.M. aided in discussion and data interpretation of the AgNP characterization and stability experiments as well as making edits to the manuscript. L.M.S. and L.M.G. wrote the paper with input from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Nanotechnology thanks Joseph Graves, Cindy Gunawan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10, Discussions 1–3 and Tables 1–5.
Rights and permissions
About this article
Cite this article
Stabryla, L.M., Johnston, K.A., Diemler, N.A. et al. Role of bacterial motility in differential resistance mechanisms of silver nanoparticles and silver ions. Nat. Nanotechnol. 16, 996–1003 (2021). https://doi.org/10.1038/s41565-021-00929-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41565-021-00929-w
This article is cited by
-
Surface Morphology, Porosity and Antibiofilm Activity of Orthosiphon aristatus-Phytosynthesized–Silver Nanoparticles Supported Zeolite A
Arabian Journal for Science and Engineering (2024)
-
Ag nanocomposite hydrogels with immune and regenerative microenvironment regulation promote scarless healing of infected wounds
Journal of Nanobiotechnology (2023)
-
Antibacterial micro/nanomotors: advancing biofilm research to support medical applications
Journal of Nanobiotechnology (2023)
-
Visible light-induced synergic antimicrobial activity of silver nanoparticles/methylene blue nanohybrids encapsulated in chitosan and pluronic polymers
Chemical Papers (2023)
-
Antibacterial Alternatives to a Dying Antibiotic Pipeline
Current Treatment Options in Infectious Diseases (2023)