Abstract
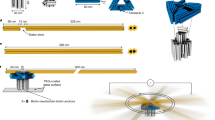
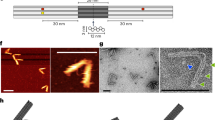
Biological motors are highly complex protein assemblies that generate linear or rotary motion, powered by chemical energy. Synthetic motors based on DNA nanostructures, bio-hybrid designs or synthetic organic chemistry have been assembled. However, unidirectionally rotating biomimetic wheel motors with rotor–stator units that consume chemical energy are elusive. Here, we report a bio-hybrid nanoengine consisting of a catalytic stator that unidirectionally rotates an interlocked DNA wheel, powered by NTP hydrolysis. The engine consists of an engineered T7 RNA polymerase (T7RNAP-ZIF) attached to a dsDNA nanoring that is catenated to a rigid rotating dsDNA wheel. The wheel motor produces long, repetitive RNA transcripts that remain attached to the engine and are used to guide its movement along predefined ssDNA tracks arranged on a DNA nanotube. The simplicity of the design renders this walking nanoengine adaptable to other biological nanoarchitectures, facilitating the construction of complex bio-hybrid structures that achieve NTP-driven locomotion.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Schliwa, M. & Woehlke, G. Molecular motors. Nature 422, 759–765 (2003).
Spetzler, D. et al. Single molecule measurements of F1-ATPase reveal an interdependence between the power stroke and the dwell duration. Biochemistry 48, 7979–7985 (2009).
Macnab, R. M. How bacteria assemble flagella. Annu. Rev. Microbiol. 57, 77–100 (2003).
Krishnan, Y. & Simmel, F. C. Nucleic acid based molecular devices. Angew. Chem. Int. Ed. 50, 3124–3156 (2011).
Zhang, D. Y. & Seelig, G. Dynamic DNA nanotechnology using strand-displacement reactions. Nat. Chem. 3, 103–113 (2011).
Bath, J. & Turberfield, A. J. DNA nanomachines. Nat. Nanotech. 2, 275–284 (2007).
Ketterer, P. Willner, E. M. & Dietz, H. Nanoscale rotary apparatus formed from tight-fitting 3D DNA components. Sci. Adv. 2, e1501209 (2016).
van den Heuvel, M. G. & Dekker, C. Motor proteins at work for nanotechnology. Science 317, 333–336 (2007).
Goel, A. & Vogel, V. Harnessing biological motors to engineer systems for nanoscale transport and assembly. Nat. Nanotech. 3, 465–475 (2008).
Giselbrecht, S., Rapp, B. E. & Niemeyer, C. M. The chemistry of cyborgs–interfacing technical devices with organisms. Angew. Chem. Int. Ed. 52, 13942–13957 (2013).
Feringa, B. L. In control of motion: from molecular switches to molecular motors. Acc. Chem. Res. 34, 504–513 (2001).
von Delius, M. & Leigh, D. A. Walking molecules. Chem. Soc. Rev. 40, 3656–3676 (2011).
Fletcher, S. P., Dumur, F., Pollard, M. M. & Feringa, B. L. A reversible, unidirectional molecular rotary motor driven by chemical energy. Science 310, 80–82 (2005).
Wilson, M. R. et al. An autonomous chemically fuelled small-molecule motor. Nature 534, 235–240 (2016).
Daubendiek, S. L., Ryan, K. & Kool, E. T. Rolling-circle RNA synthesis: circular oligonucleotides as efficient substrates for T7 RNA polymerase. J. Am. Chem. Soc. 117, 7818–7819 (1995).
Elrod-Erickson, M., Rould, M. A., Nekludova, L. & Pabo, C. O. Zif268 protein–DNA complex refined at 1.6 Å: a model system for understanding zinc finger–DNA interactions. Structure 4, 1171–1180 (1996).
Nakata, E. et al. Zinc-finger proteins for site-specific protein positioning on DNA-origami structures. Angew. Chem. Int. Ed. 51, 2421–2424 (2012).
Ngo, T. A., Nakata, E., Saimura, M. & Morii, T. Spatially organized enzymes drive cofactor-coupled cascade reactions. J. Am. Chem. Soc. 138, 3012–3021 (2016).
Lohmann, F., Valero, J. & Famulok, M. A novel family of structurally stable double stranded DNA catenanes. Chem. Commun. 50, 6091–6093 (2014).
Ackermann, D., Jester, S. S. & Famulok, M. Design strategy for DNA rotaxanes with a mechanically reinforced PX100 axle. Angew. Chem. Int. Ed. 51, 6771–6775 (2012).
Ackermann, D. et al. A double-stranded DNA rotaxane. Nat. Nanotech. 5, 436–442 (2010).
Weigandt, J., Chung, C.-L., Jester, S.-S. & Famulok, M. A daisy chain rotaxane interlocked DNA nanostructure. Angew. Chem. Int. Ed. 55, 5512–5516 (2016).
Lionberger, T. A. & Meyhofer, E. Bending the rules of transcriptional repression: tightly looped DNA directly represses T7 RNA polymerase. Biophys. J. 99, 1139–1148 (2010).
Lee, W., von Hippel, P. H. & Marcus, A. H. Internally labeled Cy3/Cy5 DNA constructs show greatly enhanced photo-stability in single-molecule FRET experiments. Nucleic Acids Res 42, 5967–5977 (2014).
Yang, W. P., Wu, H. & Barbas, C. F. 3rd Surface plasmon resonance based kinetic studies of zinc finger–DNA interactions. J. Immunol. Methods 183, 175–182 (1995).
Ko, S. H., Gallatin, G. M. & Liddle, J. A. Nanomanufacturing with DNA origami: factors affecting the kinetics and yield of quantum dot binding. Adv. Funct. Mater. 22, 1015–1023 (2012).
Abendroth, J. M., Bushuyev, O. S., Weiss, P. S. & Barrett, C. J. Controlling motion at the nanoscale: rise of the molecular machines. ACS Nano 9, 7746–7768 (2015).
Pan, J. et al. Visible/near-infrared subdiffraction imaging reveals the stochastic nature of DNA walkers. Sci. Adv. 3, e1601600 (2017).
Pan, J., Li, F., Cha, T. G., Chen, H. & Choi, J. H. Recent progress on DNA based walkers. Curr. Opin. Biotechnol. 34, 56–64 (2015).
Yehl, K. et al. High-speed DNA-based rolling motors powered by RNase H. Nat. Nanotech. 11, 184–190 (2016).
Tomov, T. E. et al. DNA bipedal motor achieves a large number of steps due to operation using microfluidics-based interface. ACS Nano 11, 4002–4008 (2017).
Zhang, D. Y. & Winfree, E. Control of DNA strand displacement kinetics using toehold exchange. J. Am. Chem. Soc. 131, 17303–17314 (2009).
Jung, C., Allen, P. B. & Ellington, A. D. A stochastic DNA walker that traverses a microparticle surface. Nat. Nanotech. 11, 157–163 (2016).
Omabegho, T., Sha, R. & Seeman, N. C. A bipedal DNA Brownian motor with coordinated legs. Science 324, 67–71 (2009).
Berna, J. et al. Macroscopic transport by synthetic molecular machines. Nat. Mater. 4, 704–710 (2005).
Kudernac, T. et al. Electrically driven directional motion of a four-wheeled molecule on a metal surface. Nature 479, 208–211 (2011).
Lu, C. H., Cecconello, A., Elbaz, J., Credi, A. & Willner, I. A three-station DNA catenane rotary motor with controlled directionality. Nano Lett. 13, 2303–2308 (2013).
Venkataraman, S., Dirks, R. M., Rothemund, P. W., Winfree, E. & Pierce, N. A. An autonomous polymerization motor powered by DNA hybridization. Nat. Nanotech. 2, 490–494 (2007).
Turberfield, A. J. et al. DNA fuel for free-running nanomachines. Phys. Rev. Lett. 90, 118102 (2003).
Bath, J., Green, S. J. & Turberfield, A. J. A free-running DNA motor powered by a nicking enzyme. Angew. Chem. Int. Ed. 44, 4358–4361 (2005).
Lund, K. et al. Molecular robots guided by prescriptive landscapes. Nature 465, 206–210 (2010).
Eelkema, R. et al. Molecular machines: nanomotor rotates microscale objects. Nature 440, 163–163 (2006).
Liu, M. H. et al. Biomimetic autonomous enzymatic nanowalker of high fuel efficiency. ACS Nano 10, 5882–5890 (2016).
Cheng, J. et al. Bipedal nanowalker by pure physical mechanisms. Phys. Rev. Lett. 109, 238104 (2012).
Loh, I. Y., Cheng, J., Tee, S. R., Efremov, A. & Wang, Z. From bistate molecular switches to self-directed track-walking nanomotors. ACS Nano 8, 10293–10304 (2014).
Bath, J., Green, S. J., Allen, K. E. & Turberfield, A. J. Mechanism for a directional, processive, and reversible DNA motor. Small 5, 1513–1516 (2009).
Tian, Y., He, Y., Chen, Y., Yin, P. & Mao, C. A DNAzyme that walks progressively and autonomously along a one-dimensional track. Angew. Chem. Int. Ed. 44, 4355–4358 (2005).
Yang, Y. et al. A photoregulated DNA-based rotary system and direct observation of its rotational movement. Chemistry 23, 3979–3985 (2017).
Michelotti, N., de Silva, C., Johnson-Buck, A. E., Manzo, A. J. & Walter, N. G. A bird’s eye view tracking slow nanometer-scale movements of single molecular nano-assemblies. Methods Enzymol. 475, 121–148 (2010).
Aitken, C. E., Marshall, R. A. & Puglisi, J. D. An oxygen scavenging system for improvement of dye stability in single-molecule fluorescence experiments. Biophys. J. 94, 1826–1835 (2008).
Fu, J. et al. Multi-enzyme complexes on DNA scaffolds capable of substrate channelling with an artificial swinging arm. Nat. Nanotech. 9, 531–536 (2014).
Suddala, K. C. & Walter, N. G. Riboswitch structure and dynamics by smFRET microscopy. Methods Enzymol. 549, 343–373 (2014).
Suddala, K. C., Wang, J., Hou, Q. & Walter, N. G. Mg2+ shifts ligand-mediated folding of a riboswitch from induced-fit to conformational selection. J. Am. Chem. Soc. 137, 14075–14083 (2015).
Fu, J. et al. Assembly of multienzyme complexes on DNA nanostructures. Nat. Protoc. 11, 2243–2273 (2016).
Rashid, F. et al. Single-molecule FRET unveils induced-fit mechanism for substrate selectivity in flap endonuclease 1. Elife 6, e21884 (2017).
Rueda, D. et al. Single-molecule enzymology of RNA: essential functional groups impact catalysis from a distance. Proc. Natl Acad. Sci. USA 101, 10066–10071 (2004).
Douglas, S. M. et al. Rapid prototyping of 3D DNA-origami shapes with caDNAno. Nucleic Acids Res. 37, 5001–5006 (2009).
Stahl, E., Martin, T. G., Praetorius, F. & Dietz, H. Facile and scalable preparation of pure and dense DNA origami solutions. Angew. Chem. Int. Ed. 53, 12735–12740 (2014).
Acknowledgements
The authors thank K. Rotscheidt, V. Vieberg and D. Keppner for technical assistance, and D. Ackermann, A. Kristofferson and A. Lange for performing preliminary studies. This work was supported by the Alexander von Humboldt Foundation and the European Research Council (ERC Advanced Grant 267173), the Max-Planck Society and the University of Bonn. N.G.W. acknowledges partial funding by Department of Defense grant W911NF-12-1-0420 and NSF grant DMR-1607854. M.F. thanks H. Famulok (1932–2017) for his genuine and encouraging interest in this work.
Authors contributions
M.F. and J.V. developed the concepts of interlocked bio-hybrid nanoengines and the walking principle. J.V. performed and designed, with M.F., most of the included studies. M.F. supervised the research project. N.P., S.D. and N.G.W. planned and performed the single-molecule fluorescence experiments. All authors discussed the experimental results and contributed to writing the manuscript (J.V. and M.F. performed the bulk of the writing).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Text, Supplementary Figures 1–34, Supplementary Tables 1–5 and Supplementary References
Rights and permissions
About this article
Cite this article
Valero, J., Pal, N., Dhakal, S. et al. A bio-hybrid DNA rotor–stator nanoengine that moves along predefined tracks. Nature Nanotech 13, 496–503 (2018). https://doi.org/10.1038/s41565-018-0109-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41565-018-0109-z
This article is cited by
-
A rhythmically pulsing leaf-spring DNA-origami nanoengine that drives a passive follower
Nature Nanotechnology (2024)
-
A synthetic tubular molecular transport system
Nature Communications (2021)
-
Building machines with DNA molecules
Nature Reviews Genetics (2020)
-
Programmable Assembly of DNA-protein Hybrid Structures
Chemical Research in Chinese Universities (2020)
-
Design, assembly, characterization, and operation of double-stranded interlocked DNA nanostructures
Nature Protocols (2019)