Abstract
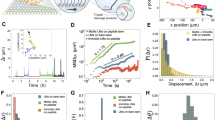
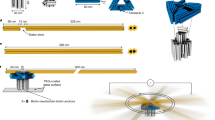
Engineering biomolecular motors can provide direct tests of structure–function relationships and customized components for controlling molecular transport in artificial systems1 or in living cells2. Previously, synthetic nucleic acid motors3,4,5 and modified natural protein motors6,7,8,9,10 have been developed in separate complementary strategies to achieve tunable and controllable motor function. Integrating protein and nucleic-acid components to form engineered nucleoprotein motors may enable additional sophisticated functionalities. However, this potential has only begun to be explored in pioneering work harnessing DNA scaffolds to dictate the spacing, number and composition of tethered protein motors11,12,13,14,15. Here, we describe myosin motors that incorporate RNA lever arms, forming hybrid assemblies in which conformational changes in the protein motor domain are amplified and redirected by nucleic acid structures. The RNA lever arm geometry determines the speed and direction of motor transport and can be dynamically controlled using programmed transitions in the lever arm structure7,9. We have characterized the hybrid motors using in vitro motility assays, single-molecule tracking, cryo-electron microscopy and structural probing16. Our designs include nucleoprotein motors that reversibly change direction in response to oligonucleotides that drive strand-displacement17 reactions. In multimeric assemblies, the controllable motors walk processively along actin filaments at speeds of 10–20 nm s−1. Finally, to illustrate the potential for multiplexed addressable control, we demonstrate sequence-specific responses of RNA variants to oligonucleotide signals.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
02 March 2018
An incorrect Supplementary Information file was originally published. The file has been replaced with the correct one.
References
Korten, T., Månsson, A. & Diez, S. Towards the application of cytoskeletal motor proteins in molecular detection and diagnostic devices. Curr. Opin. Biotechnol. 21, 477–488 (2010).
Goodman, B. S., Derr, N. D. & Reck-Peterson, S. L. Engineered, harnessed, and hijacked: synthetic uses for cytoskeletal systems. Trends Cell Biol. 22, 644–652 (2012).
Lund, K. et al. Molecular robots guided by prescriptive landscapes. Nature 465, 206–210 (2011).
Wickham, S. F. J. et al. A DNA-based molecular motor that can navigate a network of tracks. Nat. Nanotech. 7, 169–173 (2012).
Pan, J., Li, F., Cha, T.-G., Chen, H. & Choi, J. H. Recent progress on DNA based walkers. Curr. Opin. Biotechnol. 34, 56–64 (2015).
Manstein, D. J. Molecular engineering of myosin. Philos. Trans. R. Soc. Lond. B 359, 1907–1912 (2004).
Chen, L., Nakamura, M., Schindler, T. D., Parker, D. & Bryant, Z. Engineering controllable bidirectional molecular motors based on myosin. Nat. Nanotech. 7, 252–256 (2012).
Schindler, T. D., Chen, L., Lebel, P., Nakamura, M. & Bryant, Z. Engineering myosins for long-range transport on actin filaments. Nat. Nanotech. 9, 33–38 (2014).
Nakamura, M. et al. Remote control of myosin and kinesin motors using light-activated gearshifting. Nat. Nanotech. 9, 693–697 (2014).
Furuta, A. et al. Creating biomolecular motors based on dynein and actin-binding proteins. Nat. Nanotech. 12, 233–237 (2017).
Miyazono, Y., Hayashi, M., Karagiannis, P., Harada, Y. & Tadakuma, H. Strain through the neck linker ensures processive runs: a DNA–kinesin hybrid nanomachine study. EMBO J. 29, 93–106 (2010).
Derr, N. D. et al. Tug-of-war in motor protein ensembles revealed with a programmable DNA origami scaffold. Science 338, 662–665 (2012).
Furuta, K. et al. Measuring collective transport by defined numbers of processive and nonprocessive kinesin motors. Proc. Natl Acad. Sci. USA 110, 501–506 (2013).
Wollman, A. J. M., Sanchez-Cano, C., Carstairs, H. M. J., Cross, R. A. & Turberfield, A. J. Transport and self-organization across different length scales powered by motor proteins and programmed by DNA. Nat. Nanotech. 9, 44–47 (2013).
Hariadi, R. F. Mechanical coordination in motor ensembles revealed using engineered artificial myosin filaments. Nat. Nanotech. 10, 696–700 (2015).
Cheng, C. Y. et al. Consistent global structures of complex RNA states through multidimensional chemical mapping. eLife 4, e07600 (2015).
Yurke, B., Turberfield, A. J., Mills, A. P., Simmel, F. C. & Neumann, J. L. A DNA-fuelled molecular machine made of DNA. Nature 406, 605–608 (2000).
Spudich, J. A. The myosin swinging cross-bridge model. Nat. Rev. Mol. Cell Biol. 2, 387–392 (2001).
Liao, J.-C., Elting, M. W., Delp, S. L., Spudich, J. A. & Bryant, Z. Engineered myosin VI motors reveal minimal structural determinants of directionality and processivity. J. Mol. Biol. 392, 862–867 (2009).
Li, H. et al. RNA as a stable polymer to build controllable and defined nanostructures for material and biomedical applications. Nano Today 10, 631–655 (2015).
Chen, Y.-J., Groves, B., Muscat, R. A. & Seelig, G. DNA nanotechnology from the test tube to the cell. Nat. Nanotech. 10, 748–760 (2015).
Moore, T., Zhang, Y., Fenley, M. O. & Li, H. Molecular basis of box C/D RNA–protein interactions; cocrystal structure of archaeal L7Ae and a box C/D RNA. Structure 12, 807–818 (2004).
Ménétrey, J. et al. The structure of the myosin VI motor reveals the mechanism of directionality reversal. Nature 435, 779–785 (2005).
Ménétrey, J. et al. Processive steps in the reverse direction require uncoupling of the lead head lever arm of myosin VI. Mol. Cell 48, 75–86 (2012).
Saito, H. et al. Synthetic translational regulation by an L7Ae-kink-turn RNP switch. Nat. Chem. Biol. 6, 71–78 (2010).
Ohno, H. et al. Synthetic RNA–protein complex shaped like an equilateral triangle. Nat. Nanotech. 6, 116–120 (2011).
Stapleton, J. A. et al. Feedback control of protein expression in mammalian cells by tunable synthetic translational inhibition. ACS Synth. Biol. 1, 83–88 (2012).
Turner, B., Melcher, S. E., Wilson, T. J., Norman, D. G. & Lilley, D. M. J. Induced fit of RNA on binding the L7Ae protein to the kink-turn motif. RNA 11, 1192–1200 (2005).
Wells, A. L. et al. Myosin VI is an actin-based motor that moves backwards. Nature 401, 505–508 (1999).
Lister, I. et al. A monomeric myosin VI with a large working stroke. EMBO J. 23, 1729–1738 (2004).
Tsiavaliaris, G., Fujita-Becker, S. & Manstein, D. J. Molecular engineering of a backwards-moving myosin motor. Nature 427, 558–561 (2004).
Geary, C. C., Chworos, A. A. & Jaeger, L. L. Promoting RNA helical stacking via A-minor junctions. Nucleic Acids Res. 39, 1066–1080 (2011).
Ohno, H. & Saito, H. RNA and RNP as building blocks for nanotechnology and synthetic biology. Prog. Mol. Biol. Transl. Sci. 139, 165–185 (2016).
Geary, C., Rothemund, P. W. K. & Andersen, E. S. RNA nanostructures. A single-stranded architecture for cotranscriptional folding of RNA nanostructures. Science 345, 799–804 (2014).
Yang, Y. R., Liu, Y. & Yan, H. DNA nanostructures as programmable biomolecular scaffolds. Bioconjug. Chem. 26, 1381–1395 (2015).
Leontis, N. B. N. B. & Westhof, E. E. Geometric nomenclature and classification of RNA base pairs. RNA 7, 499–512 (2001).
Elting, M. W., Bryant, Z., Liao, J.-C. & Spudich, J. A. Detailed tuning of structure and intramolecular communication are dispensable for processive motion of myosin VI. Biophys. J. 100, 430–439 (2011).
Gibson, D. G., Smith, H. O., Hutchison, C. A., Venter, J. C. & Merryman, C. Chemical synthesis of the mouse mitochondrial genome. Nat. Methods 7, 901–903 (2010).
Lebel, P., Basu, A., Oberstrass, F. C., Tretter, E. M. & Bryant, Z. Gold rotor bead tracking for high-speed measurements of DNA twist, torque and extension. Nat. Methods 11, 456–462 (2014).
Acknowledgements
The authors thank M. Nakamura, T. Schindler, H. Ennomani and other members of the Bryant laboratory for discussions and assistance. This work was supported by a National Institutes of Health (NIH) Fellowship F32GM09442 to T.O., the Division of Intramural Research of the National Heart, Lung, and Blood Institute (G.A.), a Women & Science Fellowship from the Rockefeller University (to P.G.), a Human Frontiers Science Program Long-Term Fellowship (to P.V.R.), NIH High-Risk Research Grants 1DP2 OD004690 (to Z.B.) and 7DP5OD17885 (to G.A.), NIH R01 Grants GM100953 and GM102519 (to R.D.) and a grant from the W.M. Keck Foundation to Manu Prakash and Z.B.
Author information
Authors and Affiliations
Contributions
T.O. and Z.B. conceived the project. T.O. designed molecules, performed research and analysed data. P.S.G. and L.Y.K. performed cryo-EM research. P.S.G. analysed cryo-EM data. C.Y.C. performed MOHCA-seq research and analysed data. P.V.R. performed research and contributed methods. Z.B., G.M.A. and R.D. supervised research. T.O., Z.B., P.S.G., G.M.A. and C.Y.C. wrote the manuscript. All authors provided expertise, discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing financial interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
A correction to this article is available online at https://doi.org/10.1038/s41565-017-0028-4.
Electronic supplementary material
Supplementary information
Supplementary information
Rights and permissions
About this article
Cite this article
Omabegho, T., Gurel, P.S., Cheng, C.Y. et al. Controllable molecular motors engineered from myosin and RNA. Nature Nanotech 13, 34–40 (2018). https://doi.org/10.1038/s41565-017-0005-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41565-017-0005-y
This article is cited by
-
Optical control of fast and processive engineered myosins in vitro and in living cells
Nature Chemical Biology (2021)
-
Synthetic biology approaches to dissecting linear motor protein function: towards the design and synthesis of artificial autonomous protein walkers
Biophysical Reviews (2020)
-
Switchable DNA-origami nanostructures that respond to their environment and their applications
Biophysical Reviews (2018)