Abstract
Cryptococcus neoformans infections cause approximately 15% of AIDS-related deaths owing to a combination of limited antifungal therapies and drug resistance. A collection of clinical and environmental C. neoformans isolates were assayed for increased mutation rates via fluctuation analysis, and we identified two hypermutator C. neoformans clinical isolates with increased mutation rates when exposed to the combination of rapamycin and FK506. Sequencing of drug target genes found that Cnl1 transposon insertions conferred the majority of resistance to rapamycin and FK506 and could also independently cause resistance to 5-fluoroorotic acid and the clinically relevant antifungal 5-flucytosine. Whole-genome sequencing revealed both hypermutator genomes harbour a nonsense mutation in the RNA-interference component ZNF3 and hundreds of Cnl1 elements organized into massive subtelomeric arrays on each of the fourteen chromosomes. Quantitative trait locus mapping in 28 progeny derived from a cross between a hypermutator and wild-type identified a locus associated with hypermutation that included znf3. CRISPR editing of the znf3 nonsense mutation abolished hypermutation and restored small-interfering-RNA production. We conclude that hypermutation and drug resistance in these clinical isolates result from RNA-interference loss and accumulation of Cnl1 elements.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Data generated in this study are available under BioProject PRJNA749953. The BioProject accession numbers for each sample are provided in Supplementary Table 7. The publicly available datasets utilized in this study are: H99 genome (genome assembly ASM301198v1), RepBaseRepeatMaskerEdition-20181026 libraries, RepBase EMBL database (v26.04), H99, and Bt65 Illumina reads were used from published datasets (SRR642222 and SRR647805 for H99; SRR836876, SRR836877, SRR836878, SRR836880, SRR836884, and SRR836885 for Bt65) and the H99 reference genome (FungiDB-46_CneoformansH99_Genome.fasta). Source data are provided with this paper.
Code availability
Genetic variant filtering, QTL mapping, and SNP-effect prediction were conducted in Python (anaconda 3.7.3) via custom scripts available in GitHub (https://github.com/magwenelab/Hypermutator_QTL). All custom Perl scripts reported in Methods for sRNA analysis are also available in GitHub (https://github.com/timdahlmann/smallRNA).
References
Barrick, J. et al. Genome evolution and adaptation in a long-term experiment with Escherichia coli. Nature 461, 1243–1247 (2009).
Harfe, B. D. & Jinks-Robertson, S. Mismatch repair proteins and mitotic genome stability. Mutat. Res. 451, 151–167 (2000).
Billmyre, R. B., Croll, D. & Li, W. Highly recombinant VGII Cryptococcus gattii population develops clonal outbreak clusters through both sexual macroevolution and asexual microevolution. mBio 5, e01494–14 (2014).
Billmyre, R. B., Clancey, S. A. & Heitman, J. Natural mismatch repair mutations mediate phenotypic diversity and drug resistance in Cryptococcus deuterogattii. eLife 6, e28802 (2017).
Healey, K. R. et al. Prevalent mutator genotype identified in fungal pathogen Candida glabrata promotes multi-drug resistance. Nat. Commun. 7, 11128 (2016).
Boyce, K. J. et al. Mismatch repair of DNA replication errors contributes to microevolution in the pathogenic fungus Cryptococcus neoformans. mBio 8, e00595–17 (2017).
Kwon-Chung, K. J. & Chang, Y. C. Aneuploidy and drug resistance in pathogenic fungi. PLoS Pathog. 8, e1003022 (2012).
Janbon, G. et al. Characterizing the role of RNA silencing components in Cryptococcus neoformans. Fungal Genet. Biol. 47, 1070–1080 (2010).
Gusa, A. et al. Transposon mobilization in the human fungal pathogen Cryptococcus is mutagenic during infection and promotes drug resistance in vitro. Proc. Natl Acad. Sci. USA 117, 9973–9980 (2020).
Idnurm, A. et al. Deciphering the model pathogenic fungus Cryptococcus neoformans. Nat. Rev. Microbiol. 3, 753–764 (2005).
Chayakulkeeree, M. & Perfect, J. R. Cryptococcosis. Infect. Dis. Clin. North Am. 20, 507–544 (2006).
Hagen, F. et al. Recognition of seven species in the Cryptococcus gattii/Cryptococcus neoformans species complex. Fungal Genet. Biol. 78, 16–48 (2015).
Desjardins, C. A. et al. Population genomics and the evolution of virulence in the fungal pathogen Cryptococcus neoformans. Genome Res. 27, 1207–1219 (2017).
Rajasingham, R. et al. Global burden of disease of HIV-associated cryptococcal meningitis: an updated analysis. Lancet Infect. Dis. 17, 873–881 (2017).
Molloy, S. F. et al. Antifungal combinations for treatment of cryptococcal meningitis in Africa. N. Engl. J. Med. 378, 1004–1017 (2018).
Loyse, A., Dromer, F., Day, J., Lortholary, O. & Harrison, T. S. Flucytosine and cryptococcosis: time to urgently address the worldwide accessibility of a 50-year-old antifungal. J. Antimicrob. Chemother. 68, 2435–2444 (2013).
Mourad, A. & Perfect, J. R. The war on cryptococcosis: a review of the antifungal arsenal. Mem. Inst. Oswaldo Cruz 113, e170391 (2018).
Perfect, J. R. & Cox, G. M. Drug resistance in Cryptococcus neoformans. Drug Resist. Updat. 2, 259–269 (1999).
Loftus, B. J. et al. The genome of the basidiomycetous yeast and human pathogen Cryptococcus neoformans. Science 307, 1321–1324 (2005).
Goodwin, T. J. D. & Poulter, R. T. M. The diversity of retrotransposons in the yeast Cryptococcus neoformans. Yeast 18, 865–880 (2001).
Wang, X. et al. Sex-induced silencing defends the genome of Cryptococcus neoformans via RNAi. Genes Dev. 24, 2566–2582 (2010).
Dumesic, P. A. et al. Stalled spliceosomes are a signal for RNAi-mediated genome defense. Cell 152, 957–968 (2013).
Feretzaki, M., Billmyre, R. B., Clancey, S. A., Wang, X. & Heitman, J. Gene network polymorphism illuminates loss and retention of novel RNAi silencing components in the Cryptococcus pathogenic species complex. PLoS Genet. 12, e1005868 (2016).
Feretzaki, M. & Heitman, J. Genetic circuits that govern bisexual and unisexual reproduction in Cryptococcus neoformans. PLoS Genet. 9, e1003688 (2013).
Wang, X., Darwiche, S. & Heitman, J. Sex-induced silencing operates during opposite-sex and unisexual reproduction in Cryptococcus neoformans. Genetics 193, 1163–1174 (2013).
Breuder, T., Hemenway, C. S., Movva, N. R., Cardenas, M. E. & Heitman, J. Calcineurin is essential in cyclosporin A- and FK506-sensitive yeast strains. Proc. Natl Acad. Sci. USA 91, 5372–5376 (1994).
Heitman, J., Movva, N. R. & Hall, M. N. Targets for cell cycle arrest by the immunosuppressant rapamycin in yeast. Science 253, 905–909 (1991).
Billmyre, R. B., Clancey, S. A., Li, L. X., Doering, T. L. & Heitman, J. 5-Fluorocytosine resistance is associated with hypermutation and alterations in capsule biosynthesis in Cryptococcus. Nat. Commun. 11, 127 (2020).
Song, M. H. et al. A flucytosine-responsive Mbp1/Swi4-like protein, Mbs1, plays pleiotropic roles in antifungal drug resistance, stress response, and virulence of Cryptococcus neoformans. Eukaryot. Cell 11, 53–67 (2012).
Calo, S. et al. Antifungal drug resistance evoked via RNAi-dependent epimutations. Nature 513, 555–558 (2014).
Litvintseva, A. P., Thakur, R., Vilgalys, R. & Mitchell, T. G. Multilocus sequence typing reveals three genetic subpopulations of Cryptococcus neoformans var. grubii (serotype A), including a unique population in Botswana. Genetics 172, 2223–2238 (2006).
Liu, O. W. et al. Systematic genetic analysis of virulence in the human fungal pathogen Cryptococcus neoformans. Cell 135, 174–188 (2008).
Priest, S. J. et al. Factors enforcing the species boundary between the human pathogens Cryptococcus neoformans and Cryptococcus deneoformans. PLoS Genet. 17, e1008871 (2021).
Kwon-Chung, K. J., Varma, A., Edman, J. C. & Bennett, J. Selection of ura5 and ura3 mutants from the two varieties of Cryptococcus neoformans on 5-fluoroorotic acid medium. J. Med. Vet. Mycol. 30, 61–69 (1992).
Nielsen, K. et al. Sexual cycle of Cryptococcus neoformans var. grubii and virulence of congenic a and α isolates. Infect. Immun. 71, 4831–4841 (2003).
Janbon, G. et al. Analysis of the genome and transcriptome of Cryptococcus neoformans var. grubii reveals complex RNA expression and microevolution leading to virulence attenuation. PLoS Genet. 10, e1004261 (2014).
Yadav, V. et al. RNAi is a critical determinant of centromere evolution in closely related fungi. Proc. Natl Acad. Sci. USA 115, 3108–3113 (2018).
Vézina, C., Kudelski, A. & Sehgal, S. N. Rapamycin (AY-22,989), a new antifungal antibiotic. J. Antibiot. 28, 721–726 (1975).
Kino, T. et al. Structure of FK506: a novel immunosuppressant isolated from a Streptomyces. J. Am. Chem. Soc. 40, 1249–1255 (1987).
So, Y.-S., Lee, D.-G., Idnurm, A., Ianiri, G. & Bahn, Y.-S. The TOR pathway plays pleiotropic roles in growth and stress responses of the fungal pathogen Cryptococcus neoformans. Genetics 212, 1241–1258 (2019).
Juvvadi, P. R., Lamoth, F. & Steinbach, W. J. Calcineurin as a multifunctional regulator: unraveling novel functions in fungal stress responses, hyphal growth, drug resistance, and pathogenesis. Fungal Biol. Rev. 28, 56–59 (2014).
Drinnenberg, I. A. et al. RNAi in budding yeast. Science 326, 544–550 (2009).
Rahnama, M. et al. Transposon-mediated telomere destabilization: a driver of genome evolution in the blast fungus. Nucleic Acids Res. 48, 7197–7217 (2020).
Pardue, M.-L. & Debaryshe, P. G. Retrotransposons provide an evolutionarily robust non-telomerase mechanism to maintain telomeres. Annu. Rev. Genet. 37, 485–511 (2003).
Weiberg, A. et al. Fungal small RNAs suppress plant immunity by hijacking host RNA interference pathways. Science 342, 118–123 (2013).
McClintock, B. The origin and behavior of mutable loci in maize. Proc. Natl Acad. Sci. USA 36, 344–355 (1950).
Berman, J. & Krysan, D. J. Drug resistance and tolerance in fungi. Nat. Rev. Microbiol. 18, 319–331 (2020).
Vitte, C. & Panaud, O. LTR retrotransposons and flowering plant genome size: emergence of the increase/decrease model. Cytogenet. Genome Res. 110, 91–107 (2005).
Drinnenberg, I. A., Fink, G. R. & Bartel, D. P. Compatibility with killer explains the rise of RNAi-deficient fungi. Science 333, 1592 (2011).
Barton, E. S. et al. Herpesvirus latency confers symbiotic protection from bacterial infection. Nature 447, 326–329 (2007).
Clancey, S. A., Ruchti, F., Leibundgut-Landmann, S., Heitman, J. & Ianiri, G. A novel mycovirus evokes transcriptional rewiring in the fungus Malassezia and stimulates beta interferon production in macrophages. mBio 11, e01534-20 (2020).
Radchenko, E. A., McGinty, R. J., Aksenova, A. Y., Neil, A. J. & Mirkin, S. M. Quantitative analysis of the rates for repeat-mediated genome instability in a yeast experimental system. Methods Mol. Biol. 1672, 421–438 (2018).
Heitman, J., Movva, N. R., Hiestand, P. C. & Hall, M. N. FK 506-binding protein proline rotamase is a target for the immunosuppressive agent FK 506 in Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 88, 1948–1952 (1991).
Madeira, F. et al. The EMBL–EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res. 47, W636–W641 (2019).
Pitkin, J. W., Panaccione, D. G. & Walton, J. D. A putative cyclic peptide efflux pump encoded by the TOXA gene of the plant-pathogenic fungus Cochliobolus carbonurn. Microbiology 142, 1557–1565 (1996).
Sun, S., Priest, S. J. & Heitman, J. Cryptococcus neoformans mating and genetic crosses. Curr. Protoc. Microbiol. 53, e75 (2019).
Yadav, V., Sun, S., Coelho, M. A. & Heitman, J. Centromere scission drives chromosome shuffling and reproductive isolation. Proc. Natl Acad. Sci. USA 117, 7917–7928 (2020).
Bao, W., Kojima, K. K. & Kohany, O. Repbase update, a database of repetitive elements in eukaryotic genomes. Mob. DNA 6, 11 (2015).
Storer, J., Hubley, R., Rosen, J., Wheeler, T. J. & Smit, A. F. The Dfam community resource of transposable element families, sequence models, and genome annotations. Mob. DNA 12, 2 (2021).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Garrison, E. & Marth, G. Haplotype-based variant detection from short-read sequencing. Preprint at https://doi.org/10.48550/arXiv.1207.3907 (2012).
Roth, C., Sun, S., Billmyre, R. B., Heitman, J. & Magwene, P. M. A high-resolution map of meiotic recombination in Cryptococcus deneoformans demonstrates decreased recombination in unisexual reproduction. Genetics 209, 567–578 (2018).
Xu, S. Genetic mapping and genomic selection using recombination breakpoint data. Genetics 195, 1103–1115 (2013).
Churchill, G. A. & Doerge, R. W. Empirical threshold values for quantitative trait mapping. Genetics 138, 963–971 (1994).
Visscher, P. M., Thompson, R. & Haley, C. S. Confidence intervals in QTL mapping by bootstrapping. Genetics 143, 1013–1020 (1996).
Fan, Y. & Lin, X. Multiple applications of a transient CRISPR–Cas9 coupled with electroporation (TRACE) system in the Cryptococcus neoformans species complex. Genetics 208, 1357–1372 (2018).
Arras, S. D. M., Chitty, J. L., Blake, K. L., Schulz, B. L. & Fraser, J. A. A genomic safe haven for mutant complementation in Cryptococcus neoformans. PLoS ONE 10, e0122916 (2015).
Idnurm, A., Reedy, J. L., Nussbaum, J. C. & Heitman, J. Cryptococcus neoformans virulence gene discovery through insertional mutagenesis. Eukaryot. Cell 3, 420–429 (2004).
Fang, Y., Cui, L., Gu, B., Arredondo, F. & Tyler, B. M. Efficient genome editing in the oomycete Phytophthora sojae using CRISPR/Cas9. Curr. Protoc. Microbiol. 44, 21A.1.1–21A.1.26 (2017).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Dahlmann, T. A. & Kück, U. Dicer-dependent biogenesis of small RNAs and evidence for microRNA-like RNAs in the penicillin producing fungus Penicillium chrysogenum. PLoS ONE 10, e0125989 (2015).
Ianiri, G. et al. Mating-type-specific ribosomal proteins control aspects of sexual reproduction in Cryptococcus neoformans. Genetics 214, 635–649 (2020).
Acknowledgements
We thank B. Billmyre for initial project guidance, S. Clancey for instruction in conducting fluctuation assays and dsRNA enrichment protocols, J. Granek for preliminary analyses of hypermutator genomes, Z. Chang for assistance with the sRNA isolation, K. Sylvester for assistance with the screening of C. neoformans isolates, and the laboratory of C. Holley at Duke University for the use of their Nanodrop and BioAnalyzer equipment for preliminary sRNA analyses. We thank M. Farman and M. Rahnama for stimulating discussion on the impacts of transposons on telomere dynamics. We thank K. Zhu for assistance with the generation of figures. We also thank S. Sun, B. Billmyre, A. Alspaugh, S. Jinks-Robertson, A. Gusa, and K. Sylvester for critical reading and comments on the manuscript. This work was funded by NIH/NIAID F31 Fellowship 1F31AI143136-02A1 awarded to S.J.P., NIH/NIAID R37 MERIT award AI39115-23 and R01 grant AI50113-16 awarded to J.H., and R01 grant AI33654-04 awarded to P.M.M. and J.H. These studies were supported by a Visiting Professor travel grant awarded by Ruhr-Universität Bochum, Germany to J.H. J.H. is co-director and Fellow of the CIFAR program Fungal Kingdom: Threats and Opportunities. We also thank the Madhani laboratory and NIH grant R01 AI100272 for the KN99α msh2Δ deletion strain. T.A.D. and U.K. are funded by the German Research Foundation (DFG; Bonn Bad Godesberg, Germany; grant no. KU 517/15-1).
Author information
Authors and Affiliations
Contributions
S.J.P., V.Y., C.R., T.A.D., U.K., P.M.M., and J.H. designed experiments, interpreted data, and wrote the paper. S.J.P. performed experiments and analysed the fluctuation assay and Sanger sequencing data. V.Y. conducted the Nanopore sequencing and analysed all of the resulting data. C.R. and P.M.M. analysed the sequencing data from Bt65 × H99 crg1Δ F1 progeny and conducted QTL mapping and analysis. T.A.D. and U.K. analysed the sRNA sequencing data. S.J.P., U.K., P.M.M., and J.H. provided resources.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Bt65 and Bt81 do not display a hypermutator phenotype on 5-FC or 5-FOA.
Mutation rates of closely related VNBII strains and controls on (A) YNB + 5-FC and (B) YNB + 5-FOA media. Heights of bars represent the mutation rate and error bars represent 95% confidence intervals; mutation rates represent the number of mutations per cell per generation. Fluctuation analysis was conducted once for all strains in each experiment, with n = 10 biologically independent cultures per strain. Schematic depicts the phylogenetic relationships of all strains included in fluctuation analyses based on Desjardins et al. 201713.
Extended Data Fig. 2 Growth at elevated temperature does not result in increased mutation rates in C. neoformans strains.
Fluctuation assays were used to quantify the mutation rates of strains grown overnight at 30 °C or 37 °C and plated on YPD medium with R + F. Heights of bars indicate mean mutation rate and error bars indicate 95% confidence intervals. Mutation rates represent the number of mutations per cell per generation. Fluctuation analysis was conducted once for all strains, with n = 10 biologically independent cultures per strain.
Extended Data Fig. 3 Gel electrophoresis of FRR1, URA5, and FUR1 PCR products from resistant colonies.
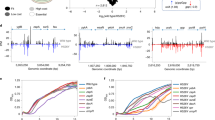
Gel electrophoresis of FRR1 PCR products from (A) all H99 R + FR colonies and a subset of (B) Bt65 and Bt81 R + FR colonies sequenced in Fig. 1D. PCR amplification of wild-type FRR1 in C. neoformans produces a 1,165 bp electrophoretic species (primers ZC7/8). Gel electrophoresis of a subset of (C) URA5 PCR products from H99, Bt65, and Bt81 5-FOAR colonies and (D) FUR1 and (E) UXS1 PCR products from 5-FCR colonies of Bt65 and Bt81.
Extended Data Fig. 4 Protein length differences of genes within QTL.
In the upper panels, points mark the strength of association (y-axis) between bi-allelic SNP sites and hypermutation for Chr3 and Chr11 (top left and right, respectively). Grey dashed lines depict the 95% confidence intervals (CI) of the two QTL. For the bi-allelic SNPs within the two QTL 95% CIs, P = 1.46868 × 10−5 (Kruskal–Wallis H-test). Lower panels show the predicted differences in lengths of proteins (y-axis) encoded by annotated genes in Bt65 compared to H99 within each 95% CI of the QTL (x-axis) on Chr3 and Chr11 (bottom left and right, respectively). The name of each gene with a predicted nonsense mutation is annotated. Blue and red colours denote the gene orientation.
Extended Data Fig. 5 QTL associated with the hypermutator phenotype span a chromosomal translocation.
(A) Nanopore whole-genome sequencing followed by synteny analysis was used to identify all indicated genomic rearrangements with respect to the reference strain H99. There is a chromosomal translocation between Chr3 and Chr11 unique to H99, and a translocation between H99 Chr1 and Chr13 unique to Bt65 and Bt81. Phylogenetic relationships of these strains are depicted in the top schematic, telomeric repeat sequences accurately identified in genomic assemblies are indicated by black half circles, and centromeres are indicated by white circles. (B) Haplotype maps of Bt65 x H99 crg1Δ F1 progeny utilized for QTL mapping. For QTLs on Chr3 and Chr11, the haplotypes (x-axis) are inferred by SNP data per segregant (y-axis) and coloured blue or orange if inherited from H99 crg1Δ or Bt65, respectively. Segregants are sorted along the y-axis by their mutation rate; largest to smallest, top to bottom. Vertical red lines display boundaries of the QTL(s). Vertical black lines depict approximate location of the translocation between H99 and Bt65. Boundaries of the QTG, ZNF3, are depicted by vertical green lines. Vertical white spaces indicate approximate locations of centromeres.
Extended Data Fig. 6 Mutation rates of Bt81 x H99 crg1Δ F1 progeny.
Fluctuation analysis was used to quantify the mutation rates of the indicated strains on YPD + rapamycin + FK506 medium (y-axis) – sorted smallest to largest, left to right – for F1 progeny and the parental strains, H99 crg1Δα and Bt81 (x-axis). Heights of bars indicate the mean mutation rate and error bars represent 95% confidence intervals. Mutation rates represent the number of mutations per cell per generation. Inheritance of the Bt81 znf3 allele or H99 crg1Δ ZNF3 allele in the F1 progeny is indicated above mutation rates. Fluctuation analysis was conducted once for all strains, with n = 10 biologically independent cultures per strain.
Extended Data Fig. 7 Subtelomeric and centromeric retrotransposons in Bt89 and Bt133.
Distributions of the Tcn1–Tcn6 LTR retrotransposons and the Cnl1 non-LTR retrotransposon in the genomes of (A) Bt89 and (B) Bt133. 50 kb of subtelomeric regions as well as centromeric regions are displayed for both strains. Shading corresponds to the lengths of the Cnl1 elements, and gene arrowheads indicate the direction of transcription for all retrotransposons.
Extended Data Fig. 8 Centromere lengths do not significantly differ among H99, Bt65, Bt81, Bt89, and Bt133.
The length of each centromere (y-axis) is plotted for each strain (x-axis). The thin horizontal black line indicates average centromere length and the thicker black error bars indicate the standard error of the mean. No significant difference was found between the average centromere length of each strain (one-way analysis of variance, P = 0.153).
Extended Data Fig. 9 Distribution of Cnl1 among Bt65 x H99 crg1Δ F1 progeny and parental strains.
The Cnl1 non-LTR elements identified in the Nanopore-based whole-genome assemblies are depicted for H99, Bt65, three hypermutator F1 progeny (P02, P08, and P34, all on the left), and three non-hypermutator F1 progeny (P14, P18, and P20, all on the right). Blue and orange bars under the subtelomeric region of each chromosome indicate which parental strain the region was inherited from (orange for Bt65, blue for H99 crg1Δ). Red asterisks indicate invasion of Cnl1 into an H99 crg1Δ subtelomeric region that previously had zero Cnl1 copies/fragments. Accurate assembly of telomeric repeat sequences at the end of each chromosome is indicated by a black half circle. Cnl1 length is also indicated by the shade of black for each element.
Extended Data Fig. 10 Enrichment for dsRNA does not identify any fragments likely to be dsRNA mycoviruses.
Pictured on the left are RNA samples following LiCl enrichment for dsRNA run on a 1% agarose gel. Total RNA prior to dsRNA enrichment is pictured on the right on a 1% agarose gel. Ms + is a Malassezia sympodialis strain that harbours a dsRNA virus, and Ms- is a congenic virus-cleared strain69. Two biological replicates for all samples are shown and labelled (1) and (2). The TriDye 1 kb DNA ladder (NEB) was used to estimate RNA fragment sizes.
Supplementary information
Supplementary Information
Supplementary Tables 1–7, Supplementary Figs. 1–10 and Source Data Supplementary Figs. 9,10.
Source data
Source Data Fig. 1
Raw colony counts and mutation frequencies from fluctuation analysis and Sanger sequencing results.
Source Data Fig. 2
Raw colony counts and mutation frequencies from fluctuation analysis.
Source Data Fig. 5
Raw colony counts and mutation frequencies from fluctuation analysis, differential expression analysis and quantification of sense and antisense sRNA reads corresponding to transposable elements.
Source Data Extended Data Fig. 1
Raw colony counts and mutation frequencies from fluctuation analysis.
Source Data Extended Data Fig. 2
Raw colony counts and mutation frequencies from fluctuation analysis.
Source Data Extended Data Fig. 3
Original gel images.
Source Data Extended Data Fig. 4
Genetic variants and predicted changes in genes within the QTL between H99 and Bt65.
Source Data Extended Data Fig. 6
Raw colony counts and mutation frequencies from fluctuation analysis.
Source Data Extended Data Fig. 10
Original gel images.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Priest, S.J., Yadav, V., Roth, C. et al. Uncontrolled transposition following RNAi loss causes hypermutation and antifungal drug resistance in clinical isolates of Cryptococcus neoformans. Nat Microbiol 7, 1239–1251 (2022). https://doi.org/10.1038/s41564-022-01183-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-022-01183-z
This article is cited by
-
Telomere transposon takeover in Cryptococcus
Nature Microbiology (2022)