Abstract
Global biodiversity loss and mass extinction of species are two of the most critical environmental issues the world is currently facing, resulting in the disruption of various ecosystems central to environmental functions and human health. Microbiome-targeted interventions, such as probiotics and microbiome transplants, are emerging as potential options to reverse deterioration of biodiversity and increase the resilience of wildlife and ecosystems. However, the implementation of these interventions is urgently needed. We summarize the current concepts, bottlenecks and ethical aspects encompassing the careful and responsible management of ecosystem resources using the microbiome (termed microbiome stewardship) to rehabilitate organisms and ecosystem functions. We propose a real-world application framework to guide environmental and wildlife probiotic applications. This framework details steps that must be taken in the upscaling process while weighing risks against the high toll of inaction. In doing so, we draw parallels with other aspects of contemporary science moving swiftly in the face of urgent global challenges.
Similar content being viewed by others
Main
Five of the nine proposed planetary boundaries, which represent the limits within which humans can safely inhabit the Earth1, have now been breached due to human activities: climate change, biosphere integrity loss, land-system change, plastic and chemical pollution, and altered biogeochemical cycles2. We are now firmly entrenched within a sixth mass extinction event3, with loss in corals, bats, bees and amphibians being the most prominent examples of anthropogenically driven biodiversity loss4,5,6 (Supplementary Fig. 1). Currently, widespread extinction events impair the resilience, function and stability of ecosystems, which impacts our existence7,8,9, conceptually referred to as ‘One Health’ (that is, the interconnection between people, animals and the environment)5,6.
In the face of such alarming challenges, recent advances in research have indicated the potential of microbiome-based interventions, such as probiotics to protect wildlife and mitigate environmental impacts, and microbiome transplants to restore ecosystems and improve their resilience10,11,12,13,14,15,16,17,18,19,20,21. We explore synergies between different disciplines, synthesize basic microbiome engineering and symbiosis concepts, and identify critical challenges and safety issues. Importantly, we provide guidelines for designing and implementing microbiome-based strategies to mitigate biodiversity decline using existing examples. We argue that the consequences of biodiversity loss are so serious that the risk of inaction must be weighed against the risk of taking a less-than-perfect action or even the risk of making things worse. This argument implies that regulatory and safety guidelines must be practical, flexible and stipulated using a case-by-case approach, considering the current state of each host and ecosystem as well as scientific insights. In this regard, we discuss crucial ethical considerations, risks, costs and benefits, and opportunities across multiple fields of microbiome management to synthesize an evidence-based framework to accelerate the practical use of emergent approaches. Finally, we draw parallels to other fields of applied science moving swiftly in the face of other current and urgent global challenges.
Biodiversity loss in the Anthropocene
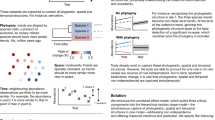
One of the primary hallmarks of the Anthropocene is the progressive elimination of species across a wide taxonomic range—from plants to insects, amphibians, birds and mammals22,23. In contrast, relatively little is known about the decline in microbial diversity and its potential consequences24. Symbiotic relationships between eukaryotic hosts and microbes constituting the holobiont or meta-organism25,26 are key for organism health and support critical ecosystem functions in all habitats on Earth27. In a healthy organism, host–microbe interactions are sufficiently balanced, with certain microbiome members often buffering against biological or environmental disturbances to maintain homoeostasis and function (Fig. 1). Projecting back, microbial symbionts were drivers of plant terrestrialization in early Palaeozoic land ecosystems28, and co-evolution with their hosts resulted in specific and unique host–microbe interactions and microbial assemblages on Earth29. Consequently, the loss of multicellular, eukaryotic hosts documented by massive extinction events in the past and today is in all likelihood accompanied by the currently undocumented loss of microbial genetic and metabolic diversity30. Recent studies have described a microbiome signature of the Anthropocene, which is characterized by a shift towards global homogenization, diversity loss, r‑strategist (that is, fast-growing) microbes and often multiresistant pathogens24,31,32. We posit that such shifts, particularly in host–microbiome associations, are often followed by dysbiosis (that is, the disruption of microbial networks and functions that impact symbiotic relationships within a holobiont)33 and facilitate disease. Dysbiosis is also associated with severe chronic diseases and long-term biotic stress that are well-documented for humans34,35 and crops36,37 but remain understudied in wildlife and natural vegetation.
Anthropogenic impacts can disturb healthy microbiomes, causing dysbiosis characterized by loss of diversity, evenness and homoeostasis, and increased prevalence of r-strategy microbes, hypermutation and antimicrobial resistance. This can result in disturbed ecosystems, the outbreak of pests and pathogens and, thus, increased risk of disease. Microbiome stewardship can exploit the microbiome given that these microbial communities are key members of the holobiont, connect all ecosystem entities, respond rapidly to manipulation with immediate effects and are easier to manipulate than macro-organisms. The use of probiotics or synbiotics is one approach to promote ecosystem functioning and overall one health, avoiding biodiversity loss, pandemics and other impacts while retaining ecosystem services.
Microbiome stewardship
The treatment and management of microbial disruptions have traditionally focused on administering antimicrobials38, with little consideration for the maintenance of the beneficial constituents of the microbiome itself. We adopt the term ‘microbiome stewardship’ to underscore that organismal and ecosystem health can be more effectively managed and maintained by monitoring and suitably manipulating the microbiome. Accordingly, the term encompasses the sum of all approaches, methodologies and technologies to understand and, consequently, manipulate microbiome function, with stewardship emphasizing the advocacy and support for science-based management.
Understanding and manipulating microbiome function is a daunting task, given the metabolic and physiological flexibility of microbes to adapt in response to change39. Additionally, our understanding of how to retain or restore a healthy microbiome is often limited40, in large part by available technologies and the lack of basic understanding of the ecological mechanisms that govern microbiome assembly, growth and evolution39,41.
Microbiome members that are important for holobiont functioning could be harnessed to rescue threatened host species or ecosystems. Competitive microbial interactions and the host immune response are the proposed mechanisms for enriching community members with antagonistic properties against pathogens. Similarly, microbial relief of holobiont stress has been proposed to rescue hosts subjected to environmental effects11,12,13,14,15,16,17,21,42,43,44. Mechanistically, this involves beneficial microbes that can improve the uptake of nutrients, vitamins and minerals, mitigate toxic compounds, control pathogens, and promote growth and fitness, among other favourable roles45. This concept is similar to the pollution-induced tolerance concept introduced by Blanck and Wängberg46 in ecotoxicology or the biological control of pathogens widely explored in agricultural systems47.
Microbe-mediated disease control is becoming more commonly employed, with microbiome-targeted interventions undergoing development. These efforts have focused on humans and plants, with aquaculture species also receiving increasing attention48,49,50. In Box 1, we present a detailed summary of the state-of-the-art microbiome stewardship approaches for some of these hosts and the strategies applied to the tailored design of probiotics across holobionts. This tendency towards defined and tailored treatment has existed for many areas (for example, precision farming and personalized medicine)51,52. While some of these strategies work well53,54, many others have failed or demonstrated inconsistent results55,56. These approaches are not necessarily new or exclusive to a specific host50,57,58,59, and efforts to manipulate the human microbiome date back over a hundred years60. The recent elevated interest in this concept is encouraging and includes a focus on wildlife and natural ecosystems10,11,12,13,14,15,16,17,18,19,20,21.
In general, microbiomes can be managed either by directly applying (1) defined microbes and mixtures/consortia of strains with beneficial properties (probiotics), (2) microbiota transplants, (3) microbiota-active metabolites or (4) mixtures of probiotics and prebiotics (synbiotics), or even indirectly by changing environmental conditions to drive shifts in the microbiome structure and function, turning a dysbiotic holobiont into a healthy one51. Although the definition of a ‘healthy’ microbiome is still being characterized and debated, on the basis of the available data, we propose a ‘microbiome stewardship’ concept that focuses on the use of probiotics. These approaches and the associated outcomes should be measured against the available alternative treatments and potential associated risks (Fig. 1).
Designing probiotics for microbiome stewardship
Designing probiotic products comprises four steps: discovery, screening and evaluation, formulation and application. This process benefits from an ecological and evolutionary understanding of the target ecosystem and the disrupted factors (with a particular focus on function). This strategy has been used to some extent for microbiome stewardship of the human gut microbiome as a treatment for Clostridioides difficile infection61. Disruption of the gut microbiome by antibiotics can lead to C. difficile infection due to reduced microbial competition and habitable niches. Microbiome restoration has been performed with faecal microbiota transplantation (FMT), which can re-establish colonization resistance and thus alleviate disease62. This paradigm of transplanting complex communities from healthy donors with intact communities to health-compromised recipients with disrupted communities could apply to other organisms and ecosystems.
Specific beneficial microbes or defined mixtures/consortia of strains could also be used to promote the health of a host or ecosystem11,13,14. Probiotic strains for such applications can be discovered from various sources, but ecology-based selection strategies that consider the origin and evolutionary history of strains are suggested to ensure adaptation and functionality. For example, microbes that exhibit inverse associations with pathologies are promising candidates for developing probiotics or defined consortia that can be explored for therapies63.
Probiotic candidates must undergo extensive screening to evaluate their efficacy and mode of action (MOA) following enrichment and isolation. The effectiveness of a multifaceted screening approach to obtain the ‘best’ microorganisms from natural bioresources has been demonstrated64. Microbial transplants from resilient wild plants, mosses and lichens were used and their colonization on crop roots and leaves were tracked to re-isolate promising probiotic candidates64. Moreover, a comprehensive set of assays was developed to determine the capability of isolates to protect against different sources of stress and MOA studies were also performed64. The inclusion of multi-omics technologies in the screening process is particularly valuable to assess the full potential of candidate strains57, including their MOA65 and potential off-target or side effects51. Consortia that combine microbes with different MOAs (that is, ‘helper strains’) and metabolic boosters (that is, compounds that can accelerate/trigger the production of specific metabolites) have also been shown to have higher efficiency than single-strain inoculations51.
Once data on their activity, genetic profile and interactions with other microbes or the host has been assembled, promising candidates or consortia can be chosen for a formulation process. In some cases, stabilizing additives are used to provide long shelf life. Subsequent large-scale production and registration are the remaining important hurdles for probiotic development. While the focus of the probiotic design is often the efficacy of the strains, other important aspects, such as safety and risk assessment, knowledge translation and ethics, must be considered during the early stages of development.
Risk and safety considerations of microbiome stewardship
Risk assessment is an important requirement that is currently among the greatest bottlenecks when registering a novel probiotic: they are costly, time-consuming and often inefficient in considering the specific features of each bioproduct66.
The regulation of bioproducts for microbial management varies between countries and areas of application, and are not compatible with each other. For example, in the United States, the Food and Drug Administration developed a safety evaluation framework that may assign microbial products the ‘Generally Recognized as Safe’ status before being marketed (https://www.fda.gov/home). However, the European Union Food Safety Authority uses a different framework based on a list of microbial species that may be considered safe depending on their taxonomic affiliation, existing scientific knowledge, pathogenicity and virulence, and safety for the environment. However, several microbial species still require full safety assessment and the number of included species is limited. Thus, most environmental probiotic candidates require a full safety assessment as they are not included in this list. This is further complicated by the fact that microbial products rely on living organisms that themselves have complex and ever-changing physiologies. Although multi-omics technologies often allow us to obtain detailed insight into the potential safety of probiotics for environmental applications, the current lack of a consensus and globally recognized framework for safety assessment still constitutes a major bottleneck in their development.
We believe that an extensive network of collaboration between research, industry and regulators is required to resolve these issues. Integrating microbiome-based concepts, as well as improving and adapting assays to evaluate the effects of new candidate probiotics on living systems offer novel possibilities for efficient risk assessment51,66. A more flexible, science-based system is needed to evaluate risks associated with the use of specific probiotics, where each situation will be evaluated through a case-by-case assessment, as detailed in Fig. 2. Whole-genome sequencing can contribute to understanding a candidate’s mode of (inter)action in detail and detecting genes encoding bioactive metabolites, virulence and antimicrobial resistance. Detecting genes of interest is especially critical if present on mobile genetic elements. However, basing the risk assessment only on genomic traits or inferred functions is probably insufficient due to other crucial factors (for example, epigenetic patterns). Therefore, assessments should be supplemented with additional investigations and methods.
The science-based flexible framework includes clear ethical and environmental safety management considerations for developing and implementing probiotics for applications to the environment in a case-by-case approach. The proposed steps could be followed sequentially or in combination and include the specific aspects that need to be considered for the selection and application of probiotics for wildlife. We highlight that risk assessment is the first step, that it should also weigh the risk of inaction against the need for rapid action and that the selection of probiotics should follow a science-informed exclusion of potential pathogens. The framework also suggests an overall survey on the application regime and potential side effects, as well as parallel research to improve our knowledge on the mechanisms driving host–microbiome interactions.
Combining ecological, genomic, transcriptomic and physiological data is of scientific value and should be included as much as possible for research on probiotic candidates. However, stipulating clear criteria for which features require investigation for a given probiotic application is challenging, and consideration must be given to the primary outcome. We suggest that knowledge-based assessments on a case-by-case basis using experimental data on microbe behaviour are necessary for environmental microbiome stewardship to counterbalance all aspects discussed above. Notably, such broadly defined individual assessments could represent an even longer process that is difficult to regulate. By comparison, a clear and universal framework that includes a fair, flexible and straightforward roadmap to follow would be more beneficial. This flexible framework should include experimental evaluations of the risks and benefits of using probiotics. In addition, the inherent risks and costs associated with inaction must be considered, as they create the opportunity to select, test and validate potentially innovative and efficient probiotics for environmental applications that current legislations would otherwise prohibit.
Lessons learned from global challenges
We should assess when rapid action is critical and would justify potential and calculated risks, as described in Fig. 2. The knowledge-based assessments of risk outlined above could benefit from an integrative analysis of the use of probiotics for different hosts, delivery methods and quick bench-to-host pathways. The rapid development of multiple vaccines against SARS-CoV-2 is an example of science moving swiftly in response to a medical emergency. Global efforts bridging stakeholders, scientists, and the public and private sectors greatly accelerated these developments, as well as adaptations in legislation, licensing and authorization systems (rapid emergency use authorization), and risk assessment systems, moving from long-term passive/active to real-time safety surveillance. In this emergent framework, rare adverse effects from vaccines were deemed acceptable considering the risk–benefit considerations to immunize most of the human adult population67,68.
As another example, while FMTs have been described for many years, they have not yet been proven safe beyond doubt and adverse effects, although rare, have been reported. Despite this, they are now regularly applied to treat human disease69,70. For C. difficile infections, the gain of curing71 most patients (up to 90%) facing life-threatening diseases must be carefully weighed against the risk of rare but serious adverse outcomes, as reported when one patient died as a result of multidrug-resistant pathogens being transplanted72. In these cases, additional precautions, such as implementing multidrug-resistant strain screening or reassessing microbiota components in donor stool, must be immediately developed to avoid such unacceptable risks. A further illustration of microbiome stewardship for the greater good is manipulation of the bacterial symbiont Wolbachia in insect vectors for human disease control (for example, to prevent the spread of dengue, yellow fever or malaria by mosquitoes or other insects)73,74. Despite the need for improvements, all these examples had known and unknown risks at the time of application, but their benefits outweighed the concerns, and inaction would have left a heavy toll in terms of human mortality and morbidity.
Ethical considerations and the inherent risk of inaction
Recent climate-related events, including extremely high temperatures in the Arctic and Antarctic regions75, devastating fires in the Amazon Rainforest, Australia and North America76,77,78, and the massive loss of coral reefs23 underscore the fact that the duty of environmental stewardship is not an abstract ideal but a concrete imperative for humankind. The stewardship of biodiversity is a collective duty as the planet’s ecosystems are strongly interdependent and the integrity of each of these systems is a necessary condition for sustaining life. Therefore, the highest priorities are to restore and conserve threatened ecosystems.
Some may consider microbiome management ethically challenging given the potential effects on other ecosystem members being modified or on the downstream food chain. For example, would using probiotic strains on honey bees result in irreversible alterations to flowering plant microbiota, or would the application of microbes to restore coral reefs affect the fish food chain and, ultimately, humans? Could environmental damage be triggered by such manipulation?
Such ecosystem interventions imply major challenges for assessing risks and benefits, with substantial implications for decision making, responsibility, accountability and governance. One potential risk is the loss of native diversity (for example, if a single probiotic strain takes over an inherent function). This is minimized by niche opportunity and environmental traits that shape microbial diversity (that is, probiotics establishment is rare and seems to be controlled by the host13 or by the availability of nutrients and conditions79 that triggered impact mitigation). However, this potential drawback should not be ignored and must be evaluated on the basis of the available alternatives.
It is also important to consider that, in some cases, not using probiotic applications may still result in losing native diversity and permit the spread of pathogens, causing major environmental damage. Engineering the environment to support a ‘healthy’ microbiome is an alternative approach to administering selected probiotics. This might include using prebiotics or beneficial bacteria isolated from the target environment, cultivated at scale and re-introduced into the system. These approaches might lower the risk of losing native diversity compared with administering genetically modified or exogenous strains41. Another potentially innovative approach would be to use bacteriophages to replace antibiotics and target specific, non-beneficial, microbes80,81.
One critical question that remains is who should decide on the initiation of field trials to test probiotics for environmental applications? For instance, the local or regional communities should be fully informed and involved during the early stages of an intervention, especially if an application remains local (for example, due to species-specific constraints). However, any alterations could ultimately spread globally to all members of that host species, creating an ethical conundrum: who makes the initial decision and can thus be held responsible? How can ecosystem interventions be justified? How can balanced information be provided? These complex ethical questions must be part of a global debate with all relevant stakeholders. Decisions must be based on broad scientific assessments (for example, peer-reviewed data, scientific networks and so on) and community consultations aligned with international regulations agreed upon in a binding manner, ideally with control instances established (for example, in the form of independent scientific entities that inform governments).
Ethical justifications for novel interventions have a stronger case if the intervention is a last resort, underscoring the ethical imperative to avoid these desperate situations. In addition, we should consider whether traditional interventions or alternative treatments, such as antibiotics, fertilizers, pesticides and other potentially harmful agents, may be more hazardous than probiotics.
An evidence-based framework for microbiome stewardship
We argue that clear ethical considerations and environmental safety management are necessary to advance research on probiotics applied to the environment and their primary host target. We propose an evidence-based framework (Fig. 2) for implementing environmental and wildlife probiotics (live microbes administered to species living in the wild) using elements discussed above, with steps that could be followed sequentially or in combination.
First, a detailed initial assessment of the problem must be undertaken similar to environmental impact assessments, ideally including the ecological and evolutionary understanding of the foundational species or ecosystem functions that can help elucidate the mechanistic basis of the environmental disruption. This assessment should include an examination of existing alternative treatments and the resulting risk–benefit and effort–benefit ratios. For example, if the problem is sewage outflow, well-studied alternatives, such as wastewater treatment plants using membranes, flocculants and activated sludge, could be assessed82,83,84,85. Likewise, the widespread application of amoxicillin (a potent, broad-spectrum antibiotic) to control the rapidly spreading stony coral tissue loss disease in the Caribbean86,87 could be compared with the harm it can cause versus the outcome of using probiotics.
Second, selecting probiotic strains on the basis of specific traits or functions is crucial to optimize the chance of success and minimize the risks. Any microbial species or strain known to be potentially pathogenic should be automatically excluded unless a convincing argument can be made to consider it. Native, commonly found and abundant commensal species should be preferentially selected unless a compelling case can be made to apply a non-native species. Strain properties should be assessed on the basis of what they are expected to achieve in the target environment. For example, the expected result could be to displace specific pathogens, function as keystone species to establish a base for the recovery of communities, improve host functions (development, immune system and so on) or remove pollutants from a habitat or holobiont system. The interaction of a probiotic organism with existing indigenous species and the surroundings should also be considered.
Third, an effective dosage of the probiotic should be used to achieve the desired effect or outcome with minimal degree of microbiome manipulation (MDMM). Whether this should exceed the native abundance of the beneficial microorganisms that exist when the host site is ‘healthy’ remains to be determined.
Fourth, the probiotic delivery system must be determined. Considerations include shelf life, storage and handling, dispersion when applied to water or other surfaces, time to reach metabolic activities essential for success, target uptake, carrier dissolution or degradation, and the broader effects on other organisms in this niche, plus assessing the overall function of the environment. For example, applying probiotics to a honeybee hive to improve resistance against pathogens should not result in lower pollination rates or damage to certain flowering plants.
Fifth, how an MDMM affects different life stages, non-target organisms and environments should be assessed. This assessment could be performed in vitro, accounting for spatial and temporal dilution, using model organisms or, when possible, as pilot experiments in natural ecosystems88,89. On the basis of the data, risks are defined through a combination of measured consequences and the likelihood of occurrence. Consequences must be defined in the context of economic, environmental, cultural and societal values41. In any case, a definitive, closely monitored pilot study is mandatory before full-scale environmental application. Finally, if the benefits of the target organism are greater than the risks to non-target organisms or the environment, then the application of probiotics should be recommended for full in-field assessment. Both scientists and stakeholders must address regulatory aspects, use a case-by-case approach with the proposed framework and acknowledge that the criteria for application in damaged ecosystems cannot be guided by the same concerns as those applied for pristine areas.
Conclusions and future perspectives
Despite the need for well-exercised caution, there is an increasing understanding that time is of the essence. Microbiome stewardship is dependent on specific traits, abiotic conditions and goals (for example, whether specific pathogens or beneficial microbes must be eliminated or promoted, respectively). Its scope includes (1) targeted disruption of specific microbes and their metabolic activity, (2) supplementing the host or ecosystem with native or non-native microbes, (3) changing the microbiome by manipulating substrates and (4) reducing host exposure to factors such as antimicrobial toxins or pollutants. Ensuring a sound scientific basis for overseeing these investigations forms an important part of microbiome stewardship, but this takes time, resources and willpower. Unfortunately, time is running out, which is why commitments must be made now, while robust investigation on macro and (especially) micro biodiversity losses, as well as ecosystem function and mechanism, should also be prioritized and supported by targeted funding opportunities.
Numerous challenges are shared across different hosts and environments. In many cases, the current technology is not safer than a microbiome modulation per se, for example when the current treatment is antibiotics, which comes with potential risk of failure or multidrug-resistance spread. We argue that the administration of strain-specific microbes, risk-assessed by the best means possible and scientifically characterized, is a safe option with potentially profound benefits.
In conclusion, we emphasize that it is imperative to address the nature and extent of the consequences of continued inaction. Many examples exist where we have failed to treat diseases with a chemical approach and have consequently caused major environmental disruption. The proposed use of beneficial microbes as an alternative is currently hindered by the lack of appropriate risk assessment or ethical frameworks that consider the dynamics of the expected benefits, current alternatives and unknown risks. We suggest joint microbiome stewardship involving existing scientific networks (for example, the beneficial microorganisms for marine organisms network) for the rapid exchange of protocols, results, strategies and resources, together with microbiota repositories (inspired by microbiota vaults90), which could catalyse the rapid development of probiotic applications to the environment. Carefully crafted production and safety policies in a system that stipulates speed without bureaucratic bottlenecks would provide companies with a clear framework to develop products that meet regulatory standards. Pilot-scale experiments could be proposed to provide baseline evidence before full-scale field tests. With sufficient data in place, legislators could work with researchers and companies to plan more widespread applications. Ultimately, we need to be able to reflect on our actions and know that we did not miss an opportunity to save our environment and the species critical to our survival.
References
Rockström, J. et al. Planetary boundaries: exploring the safe operating space for humanity. Ecol. Soc. 461, 472–475 (2009).
Steffen, W. et al. Planetary boundaries: guiding human development on a changing planet. Science 347, 1259855 (2015).
Pimm, S. L. et al. The biodiversity of species and their rates of extinction, distribution, and protection. Science 344, 1246752 (2014).
Wake, D. B. & Vredenburg, V. T. Are we in the midst of the sixth mass extinction? A view from the world of amphibians. Proc. Natl Acad. Sci. USA 105, 11466–11473 (2008).
Sweet, M., Burian, A. & Bulling, M. Corals as canaries in the coalmine: towards the incorporation of marine ecosystems into the ‘One Health’ concept. J. Invertebr. Pathol. 186, 107538 (2021).
Flandroy, L. et al. The impact of human activities and lifestyles on the interlinked microbiota and health of humans and of ecosystems. Sci. Total Environ. 627, 1018–1038 (2018).
Cardinale, B. J. et al. Biodiversity loss and its impact on humanity. Nature 486, 59–67 (2012).
Oliver, T. H. et al. Declining resilience of ecosystem functions under biodiversity loss. Nat. Commun. 6, 10122 (2015).
Loreau, M. & de Mazancourt, C. Biodiversity and ecosystem stability: a synthesis of underlying mechanisms. Ecol. Lett. 16, 106–115 (2013).
Doering, T. et al. Towards enhancing coral heat tolerance: a ‘microbiome transplantation’ treatment using inoculations of homogenized coral tissues. Microbiome 9, 102 (2021).
Rosado, P. M. et al. Marine probiotics: increasing coral resistance to bleaching through microbiome manipulation. ISME J. 13, 921–936 (2019).
Santos, H. F. et al. Impact of oil spills on coral reefs can be reduced by bioremediation using probiotic microbiota. Sci. Rep. 5, 18268 (2015).
Santoro, E. P. et al. Coral microbiome manipulation elicits metabolic and genetic restructuring to mitigate heat stress and evade mortality. Sci. Adv. 7, eabg3088 (2021).
Silva, D. P. et al. Multi-domain probiotic consortium as an alternative to chemical remediation of oil spills at coral reefs and adjacent sites. Microbiome 9, 118 (2021).
Hoyt, J. R. et al. Field trial of a probiotic bacteria to protect bats from white-nose syndrome. Sci. Rep. 9, 9158 (2019).
Bletz, M. C. et al. Mitigating amphibian chytridiomycosis with bioaugmentation: characteristics of effective probiotics and strategies for their selection and use. Ecol. Lett. 16, 807–820 (2013).
Daisley, B. A. et al. Lactobacillus spp. attenuate antibiotic-induced immune and microbiota dysregulation in honey bees. Commun. Biol. 3, 534 (2020).
Powell, J. E., Carver, Z., Leonard, S. P. & Moran, N. A. Field-realistic tylosin exposure impacts honey bee microbiota and pathogen susceptibility, which is ameliorated by native gut probiotics. Microbiol. Spectr. 9, e0010321 (2021).
Borges, D., Guzman-Novoa, E. & Goodwin, P. H. Effects of prebiotics and probiotics on honey bees (Apis mellifera) infected with the microsporidian parasite Nosema ceranae. Microorganisms 9, 481 (2021).
Daisley, B. A. et al. Novel probiotic approach to counter Paenibacillus larvae infection in honey bees. ISME J. 14, 476–491 (2020).
Trinder, M. et al. Probiotic Lactobacillus rhamnosus reduces organophosphate pesticide absorption and toxicity to Drosophila melanogaster. Appl. Environ. Microbiol. 82, 6204–6213 (2016).
Enquist, B. J., Abraham, A. J., Harfoot, M. B. J., Malhi, Y. & Doughty, C. E. The megabiota are disproportionately important for biosphere functioning. Nat. Commun. 11, 699 (2020).
Knowlton, N. et al. Rebuilding Coral Reefs: A Decadal Grand Challenge. (International Coral Reef Society, Future Earth Coasts, 2021).
Cavicchioli, R. et al. Scientists’ warning to humanity: microorganisms and climate change. Nat. Rev. Microbiol. 17, 569–586 (2019).
Jaspers, C. et al. Resolving structure and function of metaorganisms through a holistic framework combining reductionist and integrative approaches. Zoology 133, 81–87 (2019).
Bosch, T. C. G. & McFall-Ngai, M. J. Metaorganisms as the new frontier. Zoology 114, 185–190 (2011).
Wilkins, L. G. E. et al. Host-associated microbiomes and their roles in marine ecosystem functions. PLoS Biol. 17, e3000533 (2019).
Humphreys, C. P. et al. Mutualistic mycorrhiza-like symbiosis in the most ancient group of land plants. Nat. Commun. 1, 103 (2010).
Koskella, B. & Bergelson, J. The study of host-microbiome (co)evolution across levels of selection. Phil. Trans. R. Soc. Lond. B 375, 20190604 (2020).
Keller-Costa, T. et al. Metagenomic insights into the taxonomy, function, and dysbiosis of prokaryotic communities in octocorals. Microbiome 9, 72 (2021).
Guerra, C. A. et al. Global projections of the soil microbiome in the Anthropocene. Glob. Ecol. Biogeogr. 30, 987–999 (2021).
Weinbauer, M. G. & Rassoulzadegan, F. Extinction of microbes: evidence and potential consequences. Endanger. Species Res. 3, 205–215 (2007).
Petersen, C. & Round, J. L. Defining dysbiosis and its influence on host immunity and disease. Cell. Microbiol. 16, 1024–1033 (2014).
Hanski, I. et al. Environmental biodiversity, human microbiota, and allergy are interrelated. Proc. Natl Acad. Sci. USA 109, 8334–8339 (2012).
Blaser, M. J. The theory of disappearing microbiota and the epidemics of chronic diseases. Nat. Rev. Immunol. 17, 461–463 (2017).
Balbín-Suárez, A. et al. Root exposure to apple replant disease soil triggers local defense response and rhizoplane microbiome dysbiosis. FEMS Microbiol. Ecol. 97, fiab031 (2021).
Erlacher, A., Cardinale, M., Grosch, R., Grube, M. & Berg, G. The impact of the pathogen Rhizoctonia solani and its beneficial counterpart Bacillus amyloliquefaciens on the indigenous lettuce microbiome. Front. Microbiol. 5, 175 (2014).
Shahi, F., Redeker, K. & Chong, J. Rethinking antimicrobial stewardship paradigms in the context of the gut microbiome. JAC Antimicrob. Resist. 1, dlz015 (2019).
Voolstra, C. R. & Ziegler, M. Adapting with microbial help: microbiome flexibility facilitates rapid responses to environmental change. Bioessays 42, e2000004 (2020).
McBurney, M. I. et al. Establishing what constitutes a healthy human gut microbiome: state of the science, regulatory considerations, and future directions. J. Nutr. 149, 1882–1895 (2019).
Voolstra, C. R. et al. Extending the natural adaptive capacity of coral holobionts. Nat. Rev. Earth Environ. 2, 747–762 (2021).
Woodhams, D. C. et al. Prodigiosin, violacein, and volatile organic compounds produced by widespread cutaneous bacteria of amphibians can inhibit two Batrachochytrium fungal pathogens. Microb. Ecol. 75, 1049–1062 (2018).
Voyles, J. et al. Shifts in disease dynamics in a tropical amphibian assemblage are not due to pathogen attenuation. Science 359, 1517–1519 (2018).
Harris, R. N. et al. Skin microbes on frogs prevent morbidity and mortality caused by a lethal skin fungus. ISME J. 3, 818–824 (2009).
Peixoto, R. S., Harkins, D. M. & Nelson, K. E. Advances in microbiome research for animal health. Annu. Rev. Anim. Biosci. 9, 289–311 (2021).
Blanck, H. & Wängberg, S.-Å. Induced community tolerance in marine periphyton established under arsenate stress. Can. J. Fish. Aquat. Sci. 45, 1816–1819 (1988).
French, E., Kaplan, I., Iyer-Pascuzzi, A., Nakatsu, C. H. & Enders, L. Emerging strategies for precision microbiome management in diverse agroecosystems. Nat. Plants 7, 256–267 (2021).
Borges, N. et al. Bacteriome structure, function, and probiotics in fish larviculture: the good, the bad, and the gaps. Annu. Rev. Anim. Biosci. 9, 423–452 (2021).
De Schryver, P. & Vadstein, O. Ecological theory as a foundation to control pathogenic invasion in aquaculture. ISME J. 8, 2360–2368 (2014).
Sonnenschein, E. C., Jimenez, G., Castex, M. & Gram, L. The Roseobacter-group bacterium Phaeobacter as a safe probiotic solution for aquaculture. Appl. Environ. Microbiol. 87, e0258120 (2021).
Berg, G. et al. Microbiome definition re-visited: old concepts and new challenges. Microbiome 8, 103 (2020).
Peixoto, R. S., Sweet, M. & Bourne, D. G. Customized medicine for corals. Front. Mar. Sci. 6, 686 (2019).
Quraishi, M. N. et al. Systematic review with meta-analysis: the efficacy of faecal microbiota transplantation for the treatment of recurrent and refractory Clostridium difficile infection. Aliment. Pharmacol. Ther. 46, 479–493 (2017).
Henrick, B. M. et al. Bifidobacteria-mediated immune system imprinting early in life. Cell 184, 3884–3898.e11 (2021).
Freedman, S. B. et al. Multicenter trial of a combination probiotic for children with gastroenteritis. N. Engl. J. Med. 379, 2015–2026 (2018).
Cabana, M. D. et al. Early probiotic supplementation for eczema and asthma prevention: a randomized controlled trial. Pediatrics 140, e20163000 (2017).
Matsumoto, H. et al. Bacterial seed endophyte shapes disease resistance in rice. Nat. Plants 7, 60–72 (2021).
D’Alvise, P. W. et al. Phaeobacter gallaeciensis reduces Vibrio anguillarum in cultures of microalgae and rotifers, and prevents vibriosis in cod larvae. PLoS ONE 7, e43996 (2012).
Dittmann, K. K. et al. Changes in the microbiome of mariculture feed organisms after treatment with a potentially probiotic strain of Phaeobacter inhibens. Appl. Environ. Microbiol. 86, e00499-20 (2020).
Metchnikoff, E. The Prolongation of Life: Optimistic Studies (Heinemann, 1907).
Khanna, S., Jones, C., Jones, L., Bushman, F. & Bailey, A. Increased microbial diversity found in successful versus unsuccessful recipients of a next-generation FMT for recurrent Clostridium difficile infection. Open Forum Infect. Dis 5, 304–309(2015).
Kachrimanidou, M. & Tsintarakis, E. Insights into the role of human gut microbiota in Clostridioides difficile infection. Microorganisms 8, 200 (2020).
Aggarwala, V. et al. Precise quantification of bacterial strains after fecal microbiota transplantation delineates long-term engraftment and explains outcomes. Nat. Microbiol. 6, 1309–1318 (2021).
Zachow, C., Müller, H., Tilcher, R., Donat, C. & Berg, G. Catch the best: novel screening strategy to select stress protecting agents for crop plants. Agronomy 3, 794–815 (2013).
Berg, G., Kusstatscher, P., Abdelfattah, A., Cernava, T. & Smalla, K. Microbiome modulation-toward a better understanding of plant microbiome response to microbial inoculants. Front. Microbiol. 12, 650610 (2021).
Ehlers, R.-U. in Regulation of Biological Control Agents (ed. Ehlers, R.-U.) 3–23 (Springer Netherlands, 2011).
CDC. V-Safe After Vaccination Health Checker https://www.cdc.gov/coronavirus/2019-ncov/vaccines/safety/vsafe.html (2022).
Bok, K., Sitar, S., Graham, B. S. & Mascola, J. R. Accelerated COVID-19 vaccine development: milestones, lessons, and prospects. Immunity 54, 1636–1651 (2021).
Vestal, R. Fecal microbiota transplant. Hosp. Med. Clin. 5, 58–70 (2016).
Jansen, J. W. Fecal microbiota transplant vs oral vancomycin taper: important undiscussed limitations. Clin. Infect. Dis. 64, 1292–1293 (2017).
Basson, A. R., Zhou, Y., Seo, B., Rodriguez-Palacios, A. & Cominelli, F. Autologous fecal microbiota transplantation for the treatment of inflammatory bowel disease. Transl. Res. 226, 1–11 (2020).
DeFilipp, Z. et al. Drug-resistant E. coli bacteremia transmitted by fecal microbiota transplant. N. Engl. J. Med. 381, 2043–2050 (2019).
Slatko, B. E., Luck, A. N., Dobson, S. L. & Foster, J. M. Wolbachia endosymbionts and human disease control. Mol. Biochem. Parasitol. 195, 88–95 (2014).
Ahantarig, A. & Kittayapong, P. Endosymbiotic Wolbachia bacteria as biological control tools of disease vectors and pests. J. Appl. Entomol. 135, 479–486 (2011).
Turner, J. et al. Extreme temperatures in the Antarctic. J. Clim. 34, 2653–2668 (2021).
Schoennagel, T. et al. Adapt to more wildfire in western North American forests as climate changes. Proc. Natl Acad. Sci. USA 114, 4582–4590 (2017).
Di Virgilio, G. et al. Climate change increases the potential for extreme wildfires. Geophys. Res. Lett. 46, 8517–8526 (2019).
Liu, Y., Stanturf, J. & Goodrick, S. Trends in global wildfire potential in a changing climate. Ecol. Manage. 259, 685–697 (2010).
Zhou, J. et al. Stochasticity, succession, and environmental perturbations in a fluidic ecosystem. Proc. Natl Acad. Sci. USA 111, E836–E845 (2014).
Wittebole, X., De Roock, S. & Opal, S. M. A historical overview of bacteriophage therapy as an alternative to antibiotics for the treatment of bacterial pathogens. Virulence 5, 226–235 (2014).
Sieiro, C. et al. A hundred years of bacteriophages: can phages replace antibiotics in agriculture and aquaculture? Antibiotics 9, 493 (2020).
Rulkens, W. Increasing the environmental sustainability of sewage treatment by mitigating pollutant pathways. Environ. Eng. Sci. 23, 650–665 (2006).
Obotey Ezugbe, E. & Rathilal, S. Membrane technologies in wastewater treatment: a review. Membranes 10, 89 (2020).
Lee, C. S., Robinson, J. & Chong, M. F. A review on application of flocculants in wastewater treatment. Process Saf. Environ. Prot. 92, 489–508 (2014).
Guo, W.-Q., Yang, S.-S., Xiang, W.-S., Wang, X.-J. & Ren, N.-Q. Minimization of excess sludge production by in-situ activated sludge treatment processes–a comprehensive review. Biotechnol. Adv. 31, 1386–1396 (2013).
Alvarez-Filip, L., Estrada-Saldívar, N., Pérez-Cervantes, E., Molina-Hernández, A. & González-Barrios, F. J. A rapid spread of the stony coral tissue loss disease outbreak in the Mexican Caribbean. PeerJ 7, e8069 (2019).
Meiling, S. S. et al. Variable species responses to experimental stony coral tissue loss disease (SCTLD) exposure. Front. Mar. Sci. 8, 670829 (2021).
Hunt, P. R. The C. elegans model in toxicity testing. J. Appl. Toxicol. 37, 50–59 (2017).
Tkaczyk, A., Bownik, A., Dudka, J., Kowal, K. & Ślaska, B. Daphnia magna model in the toxicity assessment of pharmaceuticals: a review. Sci. Total Environ. 763, 143038 (2021).
Microbiota Vault. A Vault for Humanity https://www.microbiotavault.org/ (2021).
Health and Nutritional Properties of Probiotics in Food Including Powder Milk with Live Lactic Acid Bacteria (FAO, WHO, 2001).
Sanders, M. E., Merenstein, D. J., Reid, G., Gibson, G. R. & Rastall, R. A. Probiotics and prebiotics in intestinal health and disease: from biology to the clinic. Nat. Rev. Gastroenterol. Hepatol. 16, 605–616 (2019).
Gibson, G. R. et al. Expert consensus document: the International Scientific Association for Probiotics and Prebiotics (ISAPP) consensus statement on the definition and scope of prebiotics. Nat. Rev. Gastroenterol. Hepatol. 14, 491–502 (2017).
Salminen, S. et al. The International Scientific Association of Probiotics and Prebiotics (ISAPP) consensus statement on the definition and scope of postbiotics. Nat. Rev. Gastroenterol. Hepatol. 18, 649–667 (2021).
Liu, A. et al. Adjunctive probiotics alleviates asthmatic symptoms via modulating the gut microbiome and serum metabolome. Microbiol. Spectr. 9, e0085921 (2021).
Bagga, D. et al. Probiotics drive gut microbiome triggering emotional brain signatures. Gut Microbes 9, 486–496 (2018).
Patel, R. M. & Underwood, M. A. Probiotics and necrotizing enterocolitis. Semin. Pediatr. Surg. 27, 39–46 (2018).
Tobias, J. et al. Bifidobacterium longum subsp. infantis EVC001 administration is associated with a significant reduction in the incidence of necrotizing enterocolitis in very low birth weight infants. J. Pediatr. https://doi.org/10.1016/j.jpeds.2021.12.070 (2022).
Koziol, L. et al. The plant microbiome and native plant restoration: the example of native mycorrhizal fungi. Bioscience 68, 996–1006 (2018).
Cabello, F. C. et al. Antimicrobial use in aquaculture re-examined: its relevance to antimicrobial resistance and to animal and human health. Environ. Microbiol. 15, 1917–1942 (2013).
Evensen, Ø. & Leong, J.-A. C. DNA vaccines against viral diseases of farmed fish. Fish. Shellfish Immunol. 35, 1751–1758 (2013).
Burridge, L., Weis, J. S., Cabello, F., Pizarro, J. & Bostick, K. Chemical use in salmon aquaculture: a review of current practices and possible environmental effects. Aquaculture 306, 7–23 (2010).
Kesarcodi-Watson, A., Kaspar, H., Lategan, M. J. & Gibson, L. Probiotics in aquaculture: the need, principles and mechanisms of action and screening processes. Aquaculture 274, 1–14 (2008).
Irianto, A. & Austin, B. Probiotics in aquaculture. J. Fish. Dis. 25, 633–642 (2002).
Assefa, A. & Abunna, F. Maintenance of fish health in aquaculture: review of epidemiological approaches for prevention and control of infectious disease of fish. Vet. Med. Int. 2018, 5432497 (2018).
Hoseinifar, S. H., Sun, Y.-Z., Wang, A. & Zhou, Z. Probiotics as means of diseases control in aquaculture, a review of current knowledge and future perspectives. Front. Microbiol. 9, 2429 (2018).
Castex, M., Leclercq, E., Lemaire, P. & Chim, L. Dietary probiotic Pediococcus acidilactici MA18/5M improves the growth, feed performance and antioxidant status of penaeid shrimp Litopenaeus stylirostris: a growth-ration-size approach. Animals 11, 3451 (2021).
Goulson, D., Nicholls, E., Botías, C. & Rotheray, E. L. Bee declines driven by combined stress from parasites, pesticides, and lack of flowers. Science 347, 1255957 (2015).
Daisley, B. A., Chmiel, J. A., Pitek, A. P., Thompson, G. J. & Reid, G. Missing microbes in bees: how systematic depletion of key symbionts erodes immunity. Trends Microbiol. 28, 1010–1021 (2020).
Chmiel, J. A., Daisley, B. A., Burton, J. P. & Reid, G. Deleterious effects of neonicotinoid pesticides on Drosophila melanogaster immune pathways. Mbio 10, e01395-19 (2019).
Daisley, B. A. et al. Microbiota-mediated modulation of organophosphate insecticide toxicity by species-dependent interactions with lactobacilli in a Drosophila melanogaster insect model. Appl. Environ. Microbiol. 84, e02820-17 (2018).
Duarte, G. A. S. et al. Heat waves are a major threat to turbid coral reefs in Brazil. Front. Mar. Sci. 7, 179 (2020).
Hughes, T. P. et al. Global warming impairs stock-recruitment dynamics of corals. Nature 568, 387–390 (2019).
Hughes, T. P. et al. Coral reefs in the Anthropocene. Nature 546, 82–90 (2017).
Barno, A. R., Villela, H. D. M., Aranda, M., Thomas, T. & Peixoto, R. S. Host under epigenetic control: a novel perspective on the interaction between microorganisms and corals. Bioessays 43, e2100068 (2021).
Welsh, R. M. et al. Alien vs. predator: bacterial challenge alters coral microbiomes unless controlled by Halobacteriovorax predators. PeerJ 5, e3315 (2017).
Peixoto, R. S. et al. Coral probiotics: premise, promise, prospects. Annu. Rev. Anim. Biosci. 9, 265–288 (2021).
Peixoto, R. S. et al. Beneficial Microorganisms for Corals (BMC): proposed mechanisms for coral health and resilience. Front. Microbiol. 8, 341 (2017).
Morgans, C. A., Hung, J. Y. & Bourne, D. G. Symbiodiniaceae probiotics for use in bleaching recovery. Restoration 28, 282–288 (2020).
Zhang, Y. et al. Shifting the microbiome of a coral holobiont and improving host physiology by inoculation with a potentially beneficial bacterial consortium. BMC Microbiol. 21, 130 (2021).
Assis, J. M. et al. Delivering beneficial microorganisms for corals: rotifers as carriers of probiotic bacteria. Front. Microbiol. 11, 608506 (2020).
Zhou, G. et al. Changes in microbial communities, photosynthesis and calcification of the coral Acropora gemmifera in response to ocean acidification. Sci. Rep. 6, 35971 (2016).
VanCompernolle, S. E. et al. Antimicrobial peptides from amphibian skin potently inhibit human immunodeficiency virus infection and transfer of virus from dendritic cells to T cells. J. Virol. 79, 11598–11606 (2005).
Scheele, B. C. et al. Amphibian fungal panzootic causes catastrophic and ongoing loss of biodiversity. Science 363, 1459–1463 (2019).
Harris, R. N., Lauer, A., Simon, M. A., Banning, J. L. & Alford, R. A. Addition of antifungal skin bacteria to salamanders ameliorates the effects of chytridiomycosis. Dis. Aquat. Organ. 83, 11–16 (2009).
Loudon, A. H. et al. Interactions between amphibians’ symbiotic bacteria cause the production of emergent anti-fungal metabolites. Front. Microbiol. 5, 441 (2014).
Muletz-Wolz, C. R. et al. Inhibition of fungal pathogens across genotypes and temperatures by amphibian skin bacteria. Front. Microbiol. 8, 1551 (2017).
Jin Song, S. et al. Engineering the microbiome for animal health and conservation. Exp. Biol. Med. 244, 494–504 (2019).
Küng, D. et al. Stability of microbiota facilitated by host immune regulation: informing probiotic strategies to manage amphibian disease. PLoS ONE 9, e87101 (2014).
Micalizzi, E. W. & Smith, M. L. Volatile organic compounds kill the white-nose syndrome fungus, Pseudogymnoascus destructans, in hibernaculum sediment. Can. J. Microbiol. 66, 593–599 (2020).
Gabriel, K. T., Joseph Sexton, D. & Cornelison, C. T. Biomimicry of volatile-based microbial control for managing emerging fungal pathogens. J. Appl. Microbiol. 124, 1024–1031 (2018).
Woodhams, D. C., Bletz, M., Kueneman, J. & McKenzie, V. Managing amphibian disease with skin microbiota. Trends Microbiol. 24, 161–164 (2016).
Acknowledgements
R.S.P. acknowledges funding from King Abdullah University of Science and Technology (grants FCC/1/1973-51-01 and BAS/1/1095-01-01). C.R.V. acknowledges funding from the German Research Foundation (DFG) (grants 433042944 and 458901010). J.W. acknowledges support from the Science Foundation Ireland (SFI) through an SFI Professorship (19/RP/6853) and a Centre award (APC/SFI/12/RC/2273_P2) to the APC Microbiome Ireland. L.G. acknowledges funding from the Danish National Research Foundation (DNRF137). Funding for this work came from NSF grant no. 1924501 to R.V.T. J.S.B was supported by a Simons Foundation Early Career Investigator in Marine Microbial Ecology and Evolution award and the US National Science Foundation (NSF-OPP 1821911 and 1846837). R.C. and T.K.-C. acknowledge structural funding to iBB (grants UIDB/04565/2020 and UIDP/04565/2020) from the Portuguese Foundation for Science and Technology (FCT). R.C. acknowledges further funding from FCT and the European Regional Development Fund (ERDF) (grants PTDC/BIA-MIC/31996/2017 and ALG-01-0145-FEDER-031966). T.T. acknowledges support from the Betty and Gordon Moore Foundation. G.R. was funded by the Natural Sciences and Engineering Research Council of Canada (NSERC). B.D. acknowledges support from an NSERC Postdoctoral Fellowship (PDF-558010-2021) and the Ontario Ministry of Agriculture, Food and Rural Affairs (ND2017-3164). A.R. was funded by the Helmholtz Institute for Functional Marine Biodiversity at the University of Oldenburg, Niedersachsen, Germany and acknowledges the travel award/young investigator award from the CRC 1182 (DFG). U.H. acknowledges support from the DFG-CRC 1182 TPB01. A.S.R. acknowledges funding from King Abdullah University of Science and Technology (grant BAS/1/1096-01-01).
Author information
Authors and Affiliations
Contributions
The original discussion about the risk of inaction was initiated as part of a round table of the Beneficial Microorganisms for Corals (BMMO) network organized/developed by R.S.P., M.S., U.H., G.B, L.G, R.C and T.K.-C. at the 15th Symposium on Bacterial Genetics and Ecology (BAGECO). R.S.P, C.R.V and G.B. prepared the original draft. All authors heavily contributed with additional writing, ideas, edits and approval of the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Maria Gloria Dominguez-Bello, David Relman and Angela Sessitsch for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Fig. 1.
Rights and permissions
About this article
Cite this article
Peixoto, R.S., Voolstra, C.R., Sweet, M. et al. Harnessing the microbiome to prevent global biodiversity loss. Nat Microbiol 7, 1726–1735 (2022). https://doi.org/10.1038/s41564-022-01173-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-022-01173-1
This article is cited by
-
Disentangling direct vs indirect effects of microbiome manipulations in a habitat-forming marine holobiont
npj Biofilms and Microbiomes (2024)
-
The coral microbiome in sickness, in health and in a changing world
Nature Reviews Microbiology (2024)
-
Symbiosis, dysbiosis and the impact of horizontal exchange on bacterial microbiomes in higher fungus-gardening ants
Scientific Reports (2024)
-
Probiotics reshape the coral microbiome in situ without detectable off-target effects in the surrounding environment
Communications Biology (2024)
-
Aquaculture ecosystem microbiome at the water-fish interface: the case-study of rainbow trout fed with Tenebrio molitor novel diets
BMC Microbiology (2023)