Abstract
BBV152 is a whole-virion inactivated vaccine based on the Asp614Gly variant. BBV152 is the first alum-imidazoquinolin-adjuvanted vaccine authorized for use in large populations. Here we characterized the magnitude, quality and persistence of cellular and humoral memory responses up to 6 months post vaccination. We report that the magnitude of vaccine-induced spike and nucleoprotein antibodies was comparable with that produced after infection. Receptor binding domain-specific antibodies declined against variants in the order of Alpha (B.1.1.7; 3-fold), Delta (B.1.617.2; 7-fold) and Beta (B.1.351; 10-fold). However, pseudovirus neutralizing antibodies declined up to 2-fold against the Delta followed by the Beta variant (1.7-fold). Vaccine-induced memory B cells were also affected by the Delta and Beta variants. The SARS-CoV-2-specific multicytokine-expressing CD4+ T cells were found in ~85% of vaccinated individuals. Only a ~1.3-fold reduction in efficacy was observed in CD4+ T cells against the Beta variant. We found that antigen-specific CD4+ T cells were present in the central memory compartment and persisted for at least up to 6 months post vaccination. Vaccine-induced CD8+ T cells were detected in ~50% of individuals. Importantly, the vaccine was capable of inducing follicular T helper cells that exhibited B-cell help potential. These findings show that inactivated vaccine BBV152 induces robust immune memory to SARS-CoV-2 and variants of concern that persists for at least 6 months after vaccination.
Similar content being viewed by others
Main
BBV152 is administered as two intramuscular doses of 6 µg of inactivated virus 4 weeks apart. Almost everyone seroconverted after 4 weeks of complete vaccination1. Recently reported phase-3 data indicated an efficacy of up to 78% against symptomatic infection, 93.4% from severe disease and 63.6% from asymptomatic disease2. Despite its global distribution and potential in providing clinically substantial protection, limited evidence is available on the mechanism of immunity and the traits of immune memory established by BBV152.
The immune memory established after SARS-CoV-2 infection is expected to stay for a long term. Many studies including ours have shown that memory T and B cells persist for several months after recovery from the infection3,4,5,6. T-cell memory after recovery from COVID-19 is mainly skewed towards CD4+ T cells3,7,8. In fact, T cells are implicated in the less severe outcome of COVID-19 and mediate protection against SARS-CoV-29,10,11, probably via limiting virus load and promoting good-quality humoral responses. Also, CD4+ T cells and B cells are detected for a long time even during rapidly waning levels of antibodies12,13. Thus, in addition to the antibody response, the cellular responses play an important role in conferring long-term protection.
Studies largely carried out with messenger RNA vaccination highlight that protection may require a low level of neutralizing antibodies along with other immune effector mechanisms, including T cells and non-neutralizing antibodies14,15. Indeed, despite a decline in antibody response, infection with virus variants often lead to mild or asymptomatic disease after complete vaccination16,17,18, which further highlight the protective contribution of cellular responses. Considering that BBV152 provides an opportunity to expose the whole virion to the human immune system and it is formulated with a first-in-human adjuvant, there is a growing interest to understand the traits of immune memory established by this vaccine and its effectiveness against the virus variants. Detailed knowledge on humoral and cellular memory will be very helpful in guiding the wider use of this vaccine to control the pandemic and decide on booster immunization.
Results
Vaccine-induced antibody response and the impact of variants
We examined the immune memory in 97 SARS-CoV-2 unexposed individuals who had received BBV152 vaccine, up to 6 months (median, 4 months) after 2-dose vaccination (Fig. 1a). The participants' characteristics are indicated in Table 1. To understand the extent of humoral and cellular memory responses induced by BBV152, we compared the vaccine-induced responses with the immune memory in 99 individuals recovered from mild COVID-19. Of note, the samples in the infection group were collected between October 2020 and January 2021, before the surge of the Delta or any other variant of concern (VOC) in India. The samples were collected 130 or 214 days (median) post the second dose of BBV152 vaccine or the diagnosis of COVID-19, respectively. We first measured anti-spike antibodies in the plasma samples and found that all vaccinated and recovered individuals had detectable antibodies, with a geometric mean end-point titre (GMT) of 2.8 × 103 and 2.3 × 103, respectively (Fig. 1b). The anti-spike immunoglobulin (IgG) titre was not significantly different between the two groups. Similar to anti-spike IgG, we found that the vaccine was capable of inducing anti-nucleoprotein IgG with no significant difference from the levels following natural infection (Fig. 1c). This observation is in line with the clinical trial data of BBV15219. Moreover, similar to natural infection, significant correlation between spike and nucleoprotein antibodies suggested that the vaccine induced coordinated immune response to different virus proteins (Fig. 1d,e). SARS-CoV-2 mRNA vaccines have been shown to induce higher production of antibodies than convalescence samples20, which seems not to be the case with BBV152. However, BBV152 has an added advantage of inducing antibodies to other proteins that may assist in reducing cell-to-cell virus spread via effector mechanisms21. We then examined the efficacy of vaccine-induced antibodies against a receptor binding domain (RBD) protein of the ancestral virus (B.1) and its variants—Alpha (B.1.1.7), Beta (B.1.351), Gamma (P.1), Kappa (B.1.617.1) and Delta (B.1.617.2). The 43/46 tested subjects showed the presence of anti-RBD IgG. The Gamma and Kappa variants showed no impact on RBD-IgG; however, the levels were significantly reduced in the case of Beta (10-fold; P < 0.0001), Delta (7-fold; P = 0.002) and Alpha (3-fold; P = 0.03). Similar reduction was observed in anti-RBD IgG from natural infection, except that the decline was modest against Delta (Fig. 1f).
a, Study design to investigate the BBV152-elicited immune memory against SARS-CoV-2 and its variants. b,c, The end-point titre (ET) of SARS-CoV-2-specific antibodies measured by in-house ELISA in plasma samples derived from BBV152 vaccinated individuals (‘BBV152’) and individuals recovered from mild COVID-19 (‘COVID-19’) for spike protein (b) and nucleoprotein (c). d,e, Correlation between anti-spike and anti-nucleoprotein IgG in vaccinated (d) and COVID-19 (e) subjects. f, Comparison of anti-RBD IgG levels measured as AUC in response to ancestral SARS-CoV-2 and its Alpha, Beta, Gamma, Kappa and Delta variants in vaccinated and mild COVID-19-recovered individuals. The fold reduction in IgG levels in variant-RBD with respect to ancestral SARS-CoV-2-RBD is represented at the top of the dataset for each variant. g,h, NT50 estimated using the SARS-CoV-2 pseudovirus (PSV) expressing full-length spike protein of the ancestral virus and its Alpha, Beta and Delta variants in BBV152 (g) (n = 52) and mild COVID-19 (h) (n = 33). The fold reduction in PSV neutralizing antibody titres with respect to ancestral virus is represented at the top of the dataset for each variant. Black bars indicate geometric mean ± s.d. in b and c, and median in f. Dotted lines represent the cut-off for positivity generated using the pre-pandemic samples from healthy donors. Schematic in a was created using Biorender. Statistics by two-tailed Mann-Whitney test (b,c), two-tailed Spearman correlation coefficient (d,e), mixed-effect analysis followed by Tukey’s multiple comparisons (f) and one-way ANOVA followed by Dunn’s multiple comparisons (g,h). NS, non-significant.
Next we measured the RBD–ACE2 inhibiting antibodies using the surrogate virus neutralization assay (sVNT)22. Following the assay protocol, 1:10 dilution of plasma samples was examined. The antibodies showed neutralization potential in ~92% (22/24) of samples from the vaccine group and all the tested samples in the infection group (Extended Data Fig. 1a). We further tested the neutralizing ability of antibodies using the pseudovirus neutralization assay against SARS-CoV-2 and its variants (Extended Data Fig. 2a,b). Similar to sVNT, we found 48/52 (92%) of the vaccinated and 33/33 (100%) of the recovered individuals with the detectable SARS-CoV-2 pseudovirus neutralizing antibodies, with no significant difference in titres between the two groups (GMT; BBV152: 167 (95% CI, 92–305); COVID-19: 243 (95% CI, 158–373); Extended Data Fig. 1b). The neutralizing potential of vaccine-induced antibodies declined significantly against the Delta (2-fold vs ancestral; P < 0.0001) and Beta (1.7-fold vs ancestral; P = 0.03) variants, with no significant impact on the Alpha variant (1.2-fold vs ancestral; Fig. 1g). However, significant 3- to 4-fold reduction was observed in the neutralizing ability of the antibodies against these variants in the infection group (Fig. 1h). The highest reduction in neutralizing ability against the Beta and Delta followed by the Alpha variant was also observed in the mRNA and the AstraZeneca vaccines23,24,25,26.
The vaccine-induced spike- and RBD-IgG correlated with the pseudovirus neutralizing antibody titres (Extended Data Fig. 1c,d). Similar correlation was observed with infection-acquired antibodies, although RBD-IgG marginally correlated with neutralizing antibodies (Extended Data Fig. 1e,f). Similar to natural infection, the anti-spike, anti-nucleoprotein IgG and neutralizing antibodies were detectable for up to at least 6 months after the 2-dose vaccination (Extended Data Fig. 1g–l). Altogether, antibody analyses establish that BBV152 elicits potent antibodies against virus spike and nucleoprotein, these antibodies having the capability to neutralize virus entry, and persist for at least up to 6 months. Clearly, RBD is more vulnerable to the effect of VOCs mutations and antibodies against other parts of spike protein are also contributing to the inhibition of virus entry. The impact of VOCs was more prominent on infection-acquired neutralizing antibodies than on those acquired by vaccination. The antibody maturation over the time after recovery from infection or mRNA vaccination influences neutralization potency and breadth6,27,28. Thus, it is not clear whether the longer duration post-infection in the infection group is the reason for higher reduction in neutralizing antibodies or BBV152-induced antibodies are just more effective. Further investigations are necessary to understand the antibody evolution in BBV152 vaccination and to determine the contribution of low-titre neutralizing antibodies against VOCs.
Vaccine-induced SARS-CoV-2-specific memory B cells
B-cell memory is crucial for the durability of vaccine-induced protection. We thus examined the SARS-CoV-2-specific memory B cells induced in response to BBV152 (Fig. 2a). To set up the threshold of response, we also analysed the pre-pandemic peripheral blood mononuclear cells (PBMCs) from healthy donors. SARS-CoV-2 RBD-specific memory B cells were detected in majority of the subjects, in vaccination (IgG- antibody secreting cells (ASCs): 35/39; IgA-ASC: 21/39; IgM-ASC: 31/39) and infection (IgG-ASC: 24/24; IgA-ASC: 15/24; IgM-ASC: 15/24) (Fig. 2b–d). The vaccine-induced IgG+ B cells represent around 0.44 ± 0.1% (median, 0.13%) of total IgG+ cells, which was comparable to the proportion in the infection group (0.48 ± 0.2%; median, 0.28%) (Fig. 2b). The vaccine-induced IgA+ B cells were also present in similar frequency (0.24 ± 0.05%, median, 0.17) as in natural infection (0.14 ± 0.02%, median, 0.11) (Fig. 2c). The proportion of IgM+ cells was higher with vaccination, albeit non-significant, in the range of 1.3 ± 0.16% of the total IgM+ B cells (Fig. 2d). The results corroborate a previous study showing similar range of IgG+ and IgA+ B cells in infection13. These results also establish that BBV152 is capable of inducing the memory B cells. Interestingly, noticeable cross-reactivity exists among the IgA antibody secreting cells (ASCs) in pre-pandemic PBMCs, which was also observed previously with IgA cross-reactivity to SARS-CoV-2 RBD29. We next determined the breadth of RBD-specific B cells in their reactivity to the SARS-CoV-2 variants (Extended data Fig. 3a,b). The Kappa and Gamma variants showed no impact on reactivity of vaccine-induced IgG+ B cells (Fig. 2e). The modest reduction of 1.5 to 2-fold was observed against the Alpha and Beta variants over the ancestral virus RBD-specific IgG+ B cells (Fig. 2e). While we were concluding the B-cell analyses, the Delta variant emerged as a VOC. Thus, we utilized an additional subset of samples to examine the impact of the Delta variant on memory B cells. A ~2-fold reduction in vaccine-induced IgG+ B cells was observed against the Delta variant (Fig. 2k). Interestingly, no significant impact on IgA+ or IgM+ B cells was observed in any of the variants tested (Fig. 2f,g,k, middle and right panel). IgG+ B cells established after the mild infection showed a similar trend in their reactivity towards the SARS-CoV-2 variants, except that a higher decrease was observed against the Alpha, Beta and Delta variants (Fig. 2h,l). Similar to BBV152, no significant impact of variants was observed in the infection-acquired IgA+ and IgM+ B cells (Fig. 2i,j,l, middle and right panels). Clearly, BBV152-induced memory B cells were mostly sustained against the VOCs as compared with the circulating memory B cells acquired after natural infection (Fig. 2m). It is plausible that the longer duration post-infection showed higher reduction in the circulating memory B cells. Certainly, more investigations are warranted to probe this observation. Moreover, the role of the adjuvant in this vaccine-acquired wider breadth in memory B cells cannot be overlooked and also remains to be defined.
a, Experimental design for measuring SARS-CoV-2-specific memory B cells and their isotype in FluoroSpot assay utilizing the PBMCs from vaccinated (BBV152, n = 39) and recovered (COVID-19, n = 24) individuals. b–d, Proportion of SARS-CoV-2-RBD-specific ASCs calculated in the total corresponding isotype of ASCs in 1 million PBMCs for IgG+ ASCs (b), IgA+ ASCs (c) and IgM+ ASCs (d) in vaccination and infection. Black bars indicate median. e–j, Comparison of the frequency of ancestral SARS-CoV-2 RBD-specific memory B cells and their reactivity to RBD protein of variants Alpha, Beta, Gamma and Kappa for IgG+ ASCs (e), IgA+ ASCs (f) and IgM+ ASCs (g) in BBV152, and for IgG+ ASCs (h), IgA+ ASCs (i) and IgM+ ASCs (j) in COVID-19. k,l, In a subset of samples, reactivity of RBD-specific memory B cells of all three isotypes was determined against the Delta variant in BBV152 (k) (n = 17) and COVID-19 (l) (n = 7). Fold reduction in RBD-reactive B cells against variants over the RBD-specific B cells to ancestral virus is depicted at the top of the graph for each variant. m, Comparison of the quantitative reduction in IgG+ memory B cells against multiple variants between vaccination and natural infection. Dotted lines represent the cut-off over the threshold of memory B cell isotypes observed in PBMCs from pre-pandemic healthy donors. Schematic in a was created using Biorender. Statistics by two-tailed Mann-Whitney test (b–d), one-way ANOVA followed by Dunnett’s multiple comparisons (e–j) and two-tailed Wilcoxon signed-rank test (k,l).
Vaccine-induced IgG+ B cells significantly correlated with the RBD-IgG levels and the neutralizing antibody titre, with a notable persistence of at least up to 6 months (Extended Data Fig. 3c–e). The infection-acquired IgG+ B cells showed no correlation with antibodies but persisted in circulation (Extended Data Fig. 3f–h). No impact of VOCs on the BBV152- or infection-induced IgA+ and IgM+ B cells is encouraging. It may be due to the presence of broadly reactive IgA and IgM ASCs in both groups. Dimeric IgA antibodies are more potent than IgG in neutralizing SARS-CoV-229 and IgA+ B cells with mucosal homing traits were found in SARS-CoV-2 infection30. Thus, it is also possible that IgA and IgM antibodies can bind more efficiently to VOCs-RBD due to their stable dimeric and pentameric immunoglobin structures compared with IgG. Certainly, more investigations are necessary to determine the traits and localization of BBV152-induced IgA+ B cells.
SARS-CoV-2-specific T-cell response established by BBV152
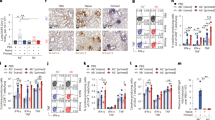
T cells are vital mediators in the protective response to SARS-CoV-2. We thus investigated the SARS-CoV-2-specific memory CD4+ T cells induced in response to BBV152 vaccination. For PBMCs stimulation, overlapping peptides megapool (MP) spanning the SARS-CoV-2 antigens of ancestral virus and VOCs were utilized7,31. We performed the activation-induced marker (AIM) assay (Fig. 3a,b and Extended Data Fig. 4a), which showed significant correlation with the IFN-γ intracellular cytokine staining (ICS) assay in detecting the antigen-specific CD4+ T cells (Extended data Fig. 5). The magnitude of total CD4+ T cells was not different between the vaccinated and recovered individuals (Fig. 3c). BBV152 induced a robust spike-specific CD4+ T-cell response (0.01 to 1.5%; median, 0.18%), which was similar to the response to infection (0.01 to 1.35%; median, 0.29) (Fig. 3d). We found that spike-specific T cells were present in detectable levels in ~85% of the vaccinated (38/45) and ~90% of the infected individuals (36/40) (Fig. 3d). In IFN-γ FluoroSpot assay, we confirmed that the vaccine-induced spike-specific T cells were capable of effector function in secreting IFN-γ during the antigen recall response (IFN-γ spot-forming cells (SFC), dimethylsulfoxide (DMSO): 22 ± 8; spike: 98 ± 21; P = 0.002) (Extended data Fig. 6a,b). We further analysed the levels of T-cell-specific cytokines in culture supernatant of spike peptideMP-stimulated PBMCs and found that the cytokine levels were not significantly different between vaccination and infection (Extended data Fig. 7a–f). Upon antigen stimulation, the cells produced the highest amount of TNF-α, followed by IL-2, IFN-γ and IL-17, with a minimal contribution of IL-13 and IL-4 (Extended Data Fig. 7g). Similar to infection, a Th1 (T helper type 1)-skewed multifunctional profile was observed for vaccine-induced CD4+ T cells, which might be contributing to the protective response32,33. Similar profiles of multifunctional T cells secreting high TNF-α > IL-2 > IFN-γ were also seen in SARS-CoV-2 mRNA vaccination and adenoviral-vectored vaccines20,34,35. It is possible that the adjuvant in the BBV152 formulation mediated this potent effector function to the vaccine-induced CD4+ T cells36. It remains to be established whether multifunctional T cells can be used as the cellular correlates-of-protection for COVID-19 vaccines, as described for Leishmania infection37. After the spike protein, nucleoprotein is the second most targeted virus protein by infection-induced CD4+ T cells38. Similar to spike T cells, we found a high frequency of nucleoprotein-specific T cells in vaccinated individuals, which was comparable with the response to infection (median, BBV152: 0.12%; COVID-19: 0.18%) (Fig. 3e). This is encouraging as broadly directed T cells may supplement spike-specific T-cell response in the protection against SARS-CoV-2 and its variants. No significant difference was observed between vaccination and infection for T cells to an unrelated antigen, Cytomegalovirus (CMV) (Fig. 3f).
SARS-CoV-2-specific memory CD4+ T cells were quantified in PBMCs derived from individuals vaccinated with BBV152 (BBV152; n = 45) and individuals recovered from mild COVID-19 (COVID-19; n = 40). In AIM assay, PBMCs were stimulated with overlapping peptide MPs spanning spike glycoprotein and nucleoprotein of ancestral SARS-CoV-2 or the MP specific to the spike protein of VOCs. a,b, Representative gating strategy for measuring the antigen-specific AIM+ (OX40+CD137+) CD4+ T cells in the total CD4+ T cells from BBV152 (a) and COVID-19 (b) (see Extended Data Fig. 4 for detailed gating strategy). The numbers on the lower right corners of the contour plots represent the frequency of AIM+ cells in the total CD4+ T cells. c, Frequency of total CD4+ T cells in BBV152 and COVID-19. d,e, Comparison of background-subtracted spike-specific (d) and nucleoprotein-specific (e) CD4+ T cells in BBV152 and COVID-19 groups. f, CMV-specific CD4+ T cells in BBV152 versus COVID-19. g,h, Quantitation of spike-specific CD4+ T cells responding to ancestral SARS-CoV-2 and VOCs in individuals after vaccination (g) and recovery from COVID-19 (h). Values at the top of the graph for each variant represent fold reduction in spike-specific CD4+ T cells responding to VOCs over the ancestral virus. i, Representative FACS plots showing the memory subsets among SARS-CoV-2 spike-specific CD4+ T cells after BBV152 vaccination (blue) or recovery from COVID-19 (red) over total memory subsets (grey), determined as central memory (TCM: CD45RA−CCR7+), effector memory (TEM: CD45RA−CCR7−), naive (TN: CD45RA+CCR7+) or terminally differentiated effector memory cells re-expressing CD45RA (TEMRA: CD45RA+CCR7−). j, Comparison of the proportion of memory subsets in spike-specific CD4+ T cells between BBV152 and COVID-19. k, Persistence of spike-specific CD4+ T cells up to 6 months after BBV152 vaccination. l, Proportion of central (TCM) and effector (TEM) memory subsets in the persisted spike-specific memory CD4+ T cells after BBV152 vaccination. Black bars indicate median. Dotted lines represent the limit of positivity generated using the pre-pandemic PBMCs from healthy donors. Statistics by two-tailed Mann-Whitney test (c–f), two-tailed Wilcoxon signed-rank test (g,h), two-way ANOVA followed by Tukey’s multiple comparisons (j) and non-linear regression (k,l).
We next measured the CD4+ T-cell response directed to spike protein of SARS-CoV-2 and the VOCs. The BBV152-induced CD4+ T cells were not reduced significantly against Alpha and Delta as compared with the ancestral virus spike protein (mean %, ancestral: 0.39 ± 0.07; Alpha: 0.31 ± 0.05; Delta: 0.28 ± 0.05) (Fig. 3g). Although a significant decline was observed in the Beta variant, the median reduction was 1.3-fold over the ancestral virus (0.25 ± 0.04; P = 0.004). The reactivity profile of CD4+ T cells acquired from natural infection was comparable with that of the vaccine, except that the median decline was ~2-fold for Alpha (P = 0.004) and Beta (P = 0.0003) (Fig. 3h). Importantly, majority of the participants showed memory T cells in detectable limits even after the observed decline. A significant decline was observed against the Beta variant for the mRNA vaccine31. However, it only accounted for a ~10% decrease in the total reactivity. As most of the response was still preserved, the decline in the range of 10–20% may not pose a concern for recall response against VOCs. We further assessed the memory phenotype of virus-specific CD4+ T cells. The gating strategy is shown in Extended Data Fig. 4a and in Fig. 3i, where AIM+ cells are depicted as the antigen-specific cells in both vaccine and infection groups. Similar to infection, vaccine-induced AIM+ T cells were also enriched in both the central and effector memory compartments (Fig. 3j). Importantly, vaccine-induced SARS-CoV-2-specific CD4+ T cells persisted for at least up to 6 months (Fig. 3k), with both the central and effector memory cells (Fig. 3l). It seems that vaccine-induced central memory cells are more durable than the effector memory subset, which is an obvious biological process in the differentiation of memory CD4+ T cells39.
We also examined the capability of BBV152 in inducing CD8+ T cells. The SARS-CoV-2-specific CD8+ T cells were detected in AIM assay (Extended Data Fig. 4b) and examined against the ancestral virus and VOCs (Extended Data Fig. 8a,b). We found 15/30 and 16/20 individuals with detectable spike-specific CD8+ T cells in vaccination and infection, respectively (Extended Data Fig. 8d). Similarly, 10/24 and 8/15 individuals showed nucleoprotein-specific CD8+ T cells in response to BBV152 vaccine and recovery from mild COVID-19, respectively (Extended Data Fig. 8e). However, no significant difference was observed in the frequency of CD8+ T cells between vaccination and infection. Also, the CMV-specific CD8+ T cells were comparable between the two groups (Extended Data Fig. 8f). Interestingly, unlike CD4+ T cells, a significant impact of VOCs was observed on the CD8+ T cells induced after vaccination or acquired from infection (Extended Data Fig. 8g,h). It is plausible that BBV152 induced low levels of CD8+ T cells that declined below detection levels in circulation with time4. However, the reduction in CD8+ T cells in response to VOCs is not clear and it appears that the cells available in circulation are mostly targeting the mutated epitopes. Also, the impact of VOCs was not similar to that of antibodies or CD4+ T cells, as CD8+ T-cell responses were also affected by the Alpha variant. Certainly, further investigation in a larger cohort will be helpful in probing the observed impact of VOCs on CD8 T cells both in vaccination and infection. These results suggest that BBV152-induced CD8+ T cells are detected in a lesser fraction of participants and are more prominently impacted by VOCs than the cells acquired from infection.
B cells help the function of vaccine-induced CD4+ T cells
Follicular T helper (Tfh) cells are a vital subset of CD4+ T cells required for protective immunity against pathogens40. Tfh-cell response has been reported in SARS-CoV-2 infection10,41 and in mRNA vaccination34,42,43. We thus examined the potential of BBV152 in inducing Tfh cells. The circulating memory Tfh cells exist as the quiescent memory (CXCR5+PD1−) and activated memory Tfh cells (CXCR5+PD1+)44,45. Therefore, we measured total antigen-specific Tfh cells for spike (Fig. 4a,b) and nucleoprotein (Fig. 4c,d) by gating the CXCR5+PD1+/− cells in the AIM+CD4+ T cells4,43. Spike-specific Tfh cells were present in the range of 0.03% to 0.43% of total CD4+ T cells in the vaccinated individuals (median, 0.11%), which was marginally lower than the Tfh cells acquired after mild infection (0.06% to 0.38%, median, 0.15%; P = 0.03) (Fig. 4b). The vaccine also induced nucleoprotein-specific Tfh cells (0.03% to 0.29%; median, 0.07%), which was comparable with infection (0.06% to 0.26%; median, 0.09%) (Fig. 4d). Our observation is in line with a previous report where individuals with mild disease favoured efficient Tfh responses46. The duration of antigen persistence is a critical driver in Tfh generation47. In fact, active germinal centres have been reported in individuals 3–4 months after infection48. Thus, it is possible that an inferior persistence of inactivated virus antigens and shorter duration of germinal centre response are causing the lower magnitude of spike-Tfh cells after vaccination. Circulating Tfh (cTfh) cells are divided depending on the surface expression of CCR6/CXCR3 into Tfh1 (CXCR3+CCR6−), Tfh2 (CXCR3−CCR6−) or Tfh17 (CXCR3−CCR6+) subsets (Extended Data Fig. 9a)44. Similar to infection, BBV152-induced cTfh cells seem to comprise all three subsets, with highest enrichment of Tfh2 and Tfh1 cells (Extended Data Fig. 9b). Unlike spike-Tfh cells, nucleoprotein-Tfh cells showed preferential enrichment in the Tfh2 polarized subset for both vaccination and infection (Extended Data Fig. 9c). The functional relevance of cTfh heterogeneity is not clearly defined and it may vary in different pathological conditions49,50. It remains to be defined how Tfh heterogeneity against spike or nucleoprotein is implicated in the vaccine-induced protective immune responses. Importantly, we found significant correlation between the neutralizing antibody titre and the SARS-CoV-2-specific total CD4+ T cells (P = 0.002) as well as cTfh cells (P = 0.009) (Extended Data Fig. 10a,d), which was not prominent in infection (Extended Data Fig. 10e–h). These results further signify the implication of vaccine-induced CD4+ T cells in the establishment of antibody responses to BBV152.
a, Representative FACS plot depicting the gating strategy for quantitation of circulating Tfh (CXCR5+PD1+) cells in spike-specific CD4+ T cells. b, Comparison of spike-specific circulating Tfh-cell frequencies in total CD4+ T cells after BBV152 vaccination (n = 42) or mild COVID-19 infection (n = 37). c,d, Similar analyses for nucleoprotein-specific Tfh cells showing representative FACS plots (c) and comparison of nucleoprotein-specific Tfh cells in BBV152 (n = 25) and COVID-19 (n = 19) groups (d). e, Experimental design for the qualitative assessment of SARS-CoV-2 spike-specific memory CD4+ T cells for their potential in inducing humoral immunity, as determined by enumeration of IL-21 secreting cells and generation of plasma cells in autologous T-B co-cultures stimulated with full-length spike protein. f, Representative images of IL-21 ELIspot depicting IL-21 SFCs per well in two of the representative donors from BBV152 vaccination and COVID-19 infection. g, Comparison of the magnitude of IL-21-secreting cells between BBV152 (n = 12) and COVID-19 (n = 16) groups. h, FACS plots representing the frequency of CD20loCD38hi plasma cells gated on CD27+ memory B cells in unstimulated (unstim) or spike protein (spike) stimulation in two donors each from BBV152 and COVID-19 groups. i, Magnitude of plasma cells (as the percentage of CD27+ B cells) in unstimulated and spike-stimulated conditions for BBV152 and COVID-19 groups. j, Comparison of the frequency of background-subtracted spike-specific plasma cells between BBV152 (n = 20) and mild COVID-19 (n = 23). Black bars indicate mean ± s.e.m. Data represent the pool of 6 independent experiments. Schematic in e was created using Biorender. Statistics by two-tailed Mann-Whitney test (b,d,g,j) and two-tailed Wilcoxon signed-rank test (i).
The B-cell help potential of CD4+ T cells has not yet been demonstrated in COVID-19 infection or vaccination. We recently described an efficient method for B-cell help function of SARS-CoV-2-specific CD4+ T cells51. We thus utilized this method to examine the help potential of BBV152-induced CD4+ T cells in autologous T-B (T cell-B cell) co-cultures (Fig. 4e). Similar to spike-Tfh, high frequency of IL-21-secreting cells were present in recovered individuals than in vaccinated participants (COVID-19: 367 ± 50; BBV152: 179 ± 22; P = 0.008) (Fig. 4f,g). The plasma cell output reflects the threshold of help signals received from T cells. We thus determined the plasma cell output in these co-cultures by measuring the frequency of CD20loCD38hi plasma cells in CD3−CD19+CD27+ memory B cells (Fig. 4h). As expected, plasma cell output was significantly higher in spike-stimulated conditions than in unstimulated conditions (Fig. 4i). Clearly, CD4+ T cells from vaccinated individuals were equally capable of inducing the plasma cell output similar to the T cells from recovered individuals (Fig. 4j). These data suggest that the vaccine is capable of generating adequate quality CD4+ T cells with potent help functions similar to those acquired following natural infection.
Discussion
The BBV152 vaccine has the dual advantage of inclusion of whole virus and an adjuvant, alum-adsorbed imidazoquinolin. Understanding the traits of immune memory established by this vaccine will provide relevant knowledge for future vaccine development and the application of TLR7/8 agonist molecules as adjuvant. Here, studying 97 participants after two doses of vaccination up to 6 months, we found that BBV152 induced a robust immune memory against spike and nucleoprotein that was comparable with that following SARS-CoV-2 infection in the levels of antibodies, memory B cells, memory CD4+ and CD8+ T cells. As demonstrated here, concurrent antibody and cellular memory responses were generated against spike and other proteins such as nucleoprotein of SARS-CoV-2. Almost all of the current VOCs showed major mutations in the spike protein of the virus. The immune memory against conserved nucleoprotein may provide an added advantage over spike-only responses in protecting against immune escape variants. Moreover, BBV152 induced a decent quantity of memory B cells that comprises cells with all three isotypes IgG, IgA and IgM. It is assuring that even after natural decline in antibodies, memory B cells can provide restoration of the antibodies during secondary virus exposure.
We found spike-specific memory CD4+ T cells in the majority (~85%) of the participants after complete vaccination that included both the central and effector memory T cells, and persisted for at least up to 6 months. The quantity of antigen-specific memory T cells which is comparable with that of infection, and the composition of memory subsets indicate durable vaccine-induced T-cell responses. Similar to infection, the vaccine-elicited T cells exhibited Th1-skewed anti-viral profile, with the cells expressing cytokines such as TNF-α, IL-2 and IFN-γ. These findings on the functionality of BBV152-induced memory CD4+ T cells are encouraging, providing assurance that the vaccine has potential to confer protection via T-cell-governed effector mechanisms. However, the vaccine-induced CD8 T-cell responses seem to be inferior to the CD4 responses. In general, inactivated vaccines do not induce strong memory CD8 T cells. However, considering the anti-viral role of CD8 T cells in COVID-19, it will be beneficial to conduct further investigations to understand the nature of circulating and tissue-resident CD8 T cells established in response to BBV152. The memory CD4 pool consists of a substantial fraction of Tfh cells specific to multiple virus proteins and comparable with that of natural infection. The CD4+ T cells with demonstrated potential of B-cell help further suggest that the adjuvanted vaccine has the ability to induce robust long-term humoral immunity.
The vaccine elicits robust immune memory against the virus variants, with modest impact observed for Delta and Beta variants, mostly on the neutralizing antibodies and memory B cells and well-preserved CD4+ T-cell responses. The Kappa variant that circulated in India and in the UK before the Delta surge, and the Gamma variant originating from Brazil showed no impact on humoral responses elicited by BBV152. Certainly, the wider breadth of BBV152-elicited cellular responses in protecting against the VOCs is in line with the immune memory acquired from infection. The vaccine-induced T cells appear to contribute to the protective response to VOCs, which is apparent in phase-3 effectiveness data that showed protection from asymptomatic infection and significantly lower viral load in Delta-positive vaccine recipients2. Thus, the data reported here provide an understanding of the immunological basis of vaccine effectiveness.
In summary, our study suggests that BBV152-induced immune memory consists of neutralizing antibodies and potent CD4+ T-cell response, with a wider breadth of cellular memory against the variants. The effective presence of memory T cells and antibodies in the local sites will be ideal to control not only SARS-CoV-2 infection but also disease transmission. Thus, it will be worth investigating whether booster immunization via the intranasal route can mediate mucosal localization of the protective immune memory. With the encouraging demonstration of the potential of BBV152 in generating immune memory and its effectiveness against the virus variants, our study provides a foundation for the future application of BBV152—an adjuvanted inactivated virus vaccine—in pandemic control response.
Limitations of the study
This study includes a modest sample size and limited blood volume to perform simultaneous analyses of multiple cellular responses against SARS-CoV-2 and VOCs in all the participants. Although the analyses from 4 weeks post the second dose up to 6 months provide key information on the persistence of immune memory, durability cannot be defined in the absence of analyses at the baseline and without longitudinal follow up. Also, it may be useful to examine the neutralization ability of vaccine-induced antibodies in the long-term using live virus settings. The quantitative comparison of immune responses between COVID-19 and BBV152 groups may not be accurate, as the samples in the infection group were collected at a much later timepoint that may have already caused a decline in antibody, B-cell and T-cell responses. However, the response in the infection group may serve as a reference to determine the extent and quality of immune memory established by the vaccine. In our study, the VOCs that infected the participants in the COVID-19 group were not identified. However, samples were collected before the circulation of VOCs in India (October 2020 to January 2021) and the participants were infected during the first wave of the COVID-19 pandemic. Another limitation is the investigation of basic performance characteristics for the assays used in this study. Although most of these assays have been published previously, they need further validation in a larger sample size for their widespread use. Particularly, the assay used for B-cell help quality of CD4+ T cells may be influenced by inter-individual variability in the immune cells. Certainly, a detailed immunological investigation of vaccine effectiveness in a larger prospective cohort is required to identify the immune correlates-of-protection and understand the mechanism of protective immunity conferred by BBV152.
Methods
Ethics statement
The study protocols were approved by the Human Research Ethics Committees of the National Institute of Immunology and the partner institutions. All associated procedures were carried out in accordance with approved guidelines. No compensation was given for participation in the study. All participants provided written informed consent before enrolment in the study.
Subject recruitment and sample collection
The vaccine cohort includes healthy participants who received 2-dose Covaxin/BBV152 vaccination. Individuals who reported SARS-CoV-2 positivity either by RT–PCR any time before blood collection or experienced COVID-19-related symptoms were excluded from the study. Another cohort of SARS-CoV-2-convalescent individuals was used to compare the vaccine-induced immune responses to the immune responses generated by natural infection. The individuals with confirmed diagnosis for COVID-19 by RT–PCR test were recruited in the infection cohort. At the time of infection, none of the patients required hospitalization and all patients experienced mild manifestation of the disease (WHO criteria). The demographic details and characteristics of participants in both the vaccine and infection cohorts are provided in Table 1. Whole blood was collected from the participants in tubes containing K3EDTA as anticoagulant. Plasma was separated from blood samples by centrifugation at 20 °C and stored at −80 °C until further use. PBMCs were isolated using Ficoll Paque Plus (GE Life Sciences) density gradient medium, cryopreserved in multiple aliquots in foetal bovine serum (FBS; Gibco) containing 10% DMSO (ThermoFisher) and stored in liquid nitrogen until further use. For all the cell-based assays, PBMCs were obtained with >80% viability as assessed by acridine orange and propidium iodide double staining in the LUNA-FL (Logos Biosystems) cell counter.
Recombinant protein from SARS-CoV-2 and its variants
Highly purified mammalian-system-expressed full-length spike protein and nucleoprotein of wild-type SARS-CoV-2 were procured from Native Antigen. Recombinant RBD proteins of the wild type as well as the variants of SARS-CoV-2 were either procured from a commercial source (Native Antigen) or received from Dr Tripti Shrivastava (THSTI, Faridabad; produced in a mammalian expression system52). The desirable mutations were introduced into the wild-type (B.1) RBD construct through site-directed mutagenesis one by one for the characteristic mutations in VOCs Kappa (B.1.617.1; E484Q, L452R), Gamma (P.1; K417T, E484K, N501Y), Alpha (B.1.1.7; N501Y), Beta (B.1.351; K417N, E484K, N501Y) and Delta (B.1.617.2; L452R, T478K). The sequence-verified plasmid DNA representing mutant as well as wild-type RBD sequences was further transfected to Expi293 cells using ExpiFectamine 293 transfection kit (ThermoFisher) and the secreted protein was purified from the supernatant.
Detection of SARS-CoV-2-specific antibodies
The plasma IgG level specific to SARS-CoV-2 proteins was measured by enzyme-linked immunosorbent assay (ELISA)3. The IgG response was analysed against full-length spike protein, nucleoprotein or RBD protein of the wild type and the variants Alpha, Beta, Gamma, Kappa and Delta. Briefly, ELISA microtitre plates (Nunc, Maxisorp) were coated with 100 µl per well of respective SARS-CoV-2 antigens in PBS (pH 7.4) at the final concentration of 1 µg ml−1 and incubated overnight at 4 °C. After blocking with PBS containing 3% skim milk and 0.05% Tween-20, serially diluted heat-inactivated plasma samples were added and incubated for 1.5 h at room temperature. Plates were washed before incubation with 1:4,000 dilution of horseradish peroxidase-conjugated anti-human IgG (Southern Biotech) for 1 h at room temperature. Plates were washed and developed using the o-phenylenediamine dihydrochloride substrate (Sigma Aldrich), stopped using HCl, and optical density (OD) was measured at 492 nm. The data were analysed using SkanIt v6.1. The antigen-coated wells that were added with sample diluent alone were used as the blank. The OD values obtained in test wells after subtracting the mean of blank OD values were used for calculating the end-point titres or the area under the curve (AUC), with a baseline of 0.05 for peak calculations. The cut-off for end-point titres or the limit of positivity for AUC was defined as the value above the mean plus three times the standard deviation of values obtained for the reactivity of respective proteins with pre-pandemic plasma samples of healthy donors. The fold reduction in RBD-IgG cross-reactivity to VOCs was determined by dividing the median IgG titre to wild-type RBD with the median IgG titre obtained from VOC-RBD.
SARS-CoV-2 sVNT
The sVNT test was used to determine the potential of antibodies in blocking the RBD–ACE2 receptor interaction. The test was performed following the manufacturer’s instructions (Genscript). Briefly, 1:10 diluted plasma samples were incubated with RBD-horseradish peroxidase and the mixture was added onto a human ACE2-coated plate. The plate was developed using TMB substrate and read at 450 nm. The sample absorbance was inversely proportional to the titre of the anti-SARS-CoV-2 neutralizing antibodies. The percent neutralization was calculated using the formula: (1 − OD value of sample/OD value of negative control) × 100%. The cut-off for the positive limit of SARS-CoV-2 neutralizing antibodies was determined by manufacturer and further validated using the plasma panel of pre-pandemic healthy controls3,22.
Pseudovirus neutralization assay
The SARS-CoV-2 neutralization assay was performed using the lentivirus-based pseudovirus system53 with modifications. SARS-CoV-2 spike pseudovirus was generated using HIV-1 NL4-3 ΔEnv Vpr luciferase reporter vector (pNL4-3.Luc.R-E) provided as a kind gift by Prof. A. C. Banerjea (National Institute of Immunology, India). Briefly, 4 × 106 HEK293T cells (ATCC CRL-3216) were co-transfected with pNL4-3.Luc.R-E and SARS-CoV-2/VOCs spike plasmids (pcDNA3.1spike_del19; Addgene, 155297; pcDNA3.3-SARS2-B.1.617.2; Addgene, 172320; pUNO1-SpikeV2 B.1.1.7; Invivogen, p1-spike-v2; pUNO1-SpikeV3 B.1.351; Invivogen, p1-spike-v3) in the ratio of 1:3 using Lipofectamine 3000 (Invitrogen) in OptiMEM (Gibco) media to generate single-round replication ancestral SARS-CoV-2 (B.1) and VOCs spike pseudoviruses. Following 4 h of incubation, the medium was replaced with complete DMEM supplemented with 10% FBS. After 24 h, complete medium was replaced with serum-free medium (Gibco) and incubated for another 72 h. The supernatant was collected, filtered and stored at −80 °C. The stable Huh7 cell line (JCRB0403) expressing ACE2 + TMPRSS2 was generated to titrate the pseudoviruses. The Huh7 cells were infected with serially diluted pseudoviruses and incubated for 72 h. The cells were washed, lysed using cell lysis buffer (Promega), added with the luciferase substrate (Promega), and the relative luminescence units (RLU) were measured using a luminometer (ThermoFisher Varioskan Flash). Neutralization assay was performed using a Huh7 cell line stably expressing ACE2 + TMPRSS2. Briefly, 15,000 cells per well were seeded in 96-well plates. SARS-CoV-2 or VOCs spike pseudovirus at 106 RLU was incubated with 3-fold serially diluted plasma samples starting from a 1:10 dilution for 1 h before addition to confluent Huh7 monolayers in 96-well plates. Following incubation, cells were washed, lysed using cell lysis buffer and added with 100 μl of luciferase substrate before measuring the luminescence using a luminometer. The data were acquired using SkanIt Software 2.4.5 RE. Percentage neutralization was calculated by normalizing the test RLU with virus control and cell-only control RLU. Neutralization IC50 (half-maximal inhibitory concentration) titres (NT50) were calculated using non-linear regression curve fit ‘log(inhibitor) vs normalized response-variable slope’ in GraphPad Prism 8.4.3. The limit of positivity was determined by performing the neutralization assay on plasma samples from pre-pandemic healthy donors. The cut-off for positive response was defined as the mean plus two times the standard deviation of the NT50 obtained with the pre-pandemic samples.
Detection of antigen-specific memory B cells
To examine the quantity and phenotype of RBD-specific memory B cells and to test their cross-reactivity to the recent variants, polyclonal stimulated B cells were tested in a FluoroSpot assay utilizing RBD antigen from the ancestral strain of SARS-CoV-2 and the variants Alpha, Beta, Gamma, Kappa and Delta. Briefly, cryopreserved PBMCs were thawed and polyclonally stimulated with R848 and IL-2 for 5 d. The stimulated cells were then captured on the FluoroSpot plate (Mabtech) coated overnight with the respective RBD-antigen at 5 µg ml−1. For measuring total memory B cells of each isotype, wells were coated with anti-human IgG (MT91/145), anti-human IgM (MT11/12) and anti-human IgA (MT57) (Mabtech) at the concentration of 15 µg ml−1. Plates were washed, blocked and seeded with the 0.5 × 106 stimulated PBMCs per well for antigen-specific analysis and 20,000 cells for total B-cell analysis, followed by incubation at 37 °C for 8 h. For detection of ASC spots corresponding to IgG+, IgM+ and IgA+ isotypes, the plate was developed using detection monoclonal anti-human antibodies: IgG-550 (MT78/145), IgM−640 (MT22) and IgA-490 (MT20), diluted 1:500 in PBS containing 0.5% FBS for 2 h in the dark at room temperature. Plates were added with fluorescence enhancer (Mabtech), washed, and spots were detected using an AID vSpot Spectrum ELISpot/FluoroSpot reader system and analysed by AID ELISpot software version 7.x. ASC counts were normalized to ASCs per million of PBMCs for all analyses. The frequency of antigen-specific memory B cells was expressed as a percentage of total B cells representing the respective isotype. The median frequency of B cells specific to wild-type RBD was divided by the median frequency of B cells reacting to the VOC-RBD and depicted as the fold reduction in frequency for respective variants. The cut-off for positive response was defined as the median plus two times the standard deviation of the threshold of memory B-cell isotypes observed in PBMCs from pre-pandemic healthy donors.
Quantification of antigen-specific T cells
Antigen-specific CD4+ T cells were measured using the activation-induced markers (AIM) assay3,7,31. SARS-CoV-2-specific CD4+ T cells and CD8+ T cells were quantified as a percentage population of AIM+ (OX40+CD137+) CD4+ T cells and AIM+ (CD69+CD137+) CD8+ T cells, respectively, after stimulation of PBMCs with overlapping peptide megapools (MPs) at a concentration of 1 μg ml−1 in AIM-V media (Gibco) for 24 h at 37 °C. The MPs consist of 15-mer peptides overlapping by 10 amino acids covering the spike (S) or nucleocapsid protein (NP) of ancestral SARS-CoV-2 (B.1; GenBank: MN_908947) and MPs covering spike protein of VOCs Alpha (B.1.1.7; GISAID: EPI_ISL_601443), Beta (B.1.351; GISAID: EPI_ISL_660629) and Delta (B.1.617.2; GISAID: EPI_ISL_2020950). All peptides were synthetized as crude material (TC Peptide Lab), resuspended in DMSO, pooled and sequentially lyophilised as previously reported7,31. An equimolar concentration of DMSO was used as a negative control. Staphylococcal enterotoxin B at 0.1 µg ml−1 and CMV-specific MP were used as the positive stimulation control. After the stimulation period, for quantification of SARS-CoV-2-specific CD4+ T cells, PBMCs were washed with fluorescence-activated cell sorting (FACS) buffer (2% FBS in PBS) and surface stained with monoclonal antibody cocktail for 1 h at 4 °C in the dark: CD20, CD14, CD16, CD8a and fixable-viability dye coupled with APC eflour 780 in the dump channel, CD4−AlexaFluor 700 (RPA-T4), OX40−FITC (Ber-ACT35), CD137 PE Dazzle 594(4B4−1), CD45RA Brilliant Violet 785 (HI100) and CCR7 AlexaFluor 647 (G043H7). The cTfh cells were quantified by measuring the frequency of CXCR5+PD1+/− cells in the AIM+CD4+ T cells obtained after stimulation with spike and nucleoprotein MPs. The frequency of cTfh cells was expressed as the percentage of total CD4+ T cells. For cTfh analyses, in addition to the AIM markers, cells were also stained with CXCR5-Brilliant Violet 421 (J252D4), PD1-PE (EH12.2H7), CXCR3-Brilliant Violet 711 (G025H7) and CCR6-PE/Cy7 (G034E3). For quantification of SARS-CoV-2-specific CD8+ T cells, PBMCs were washed with FACS buffer and surface stained with monoclonal antibody cocktail for 1 h at 4 °C in the dark: CD20, CD14, CD16 and fixable-viability dye coupled with APC eflour 780 in the dump channel, CD4-AlexaFluor 700 (RPA-T4), CD8a-APC (RPA-T8), OX40−FITC (Ber-ACT35), CD137 PE Dazzle 594(4B4-1) and CD69 Brilliant Violet 510 (FN50). For all analyses, after staining, cells were washed and fixed with freshly prepared 1% paraformaldehyde (Sigma-Aldrich), followed by acquisition using BD LSRFortessa X-20 flow cytometer (BD Biosciences) and subsequent data analysis by FlowJo 10.7.1. The antigen-specific CD4+ T cells were measured as the data subtracted from DMSO conditions as the background. The positive response in the AIM assay was defined by setting up the limit of positivity above the median plus two times the standard deviation of negative controls (pre-pandemic PBMCs) stimulated with spike PeptideMP. The frequency of responders to SARS-CoV-2 spike PeptideMP stimulation was determined by applying Fisher’s exact test on the AIM+ and AIM− cells in DMSO- and PeptideMP-stimulated conditions.
IFN-γ ICS assay
To validate quantification of SARS-CoV-2-specific CD4+ T cells in the AIM assay, IFN-γ ICS was performed on a subset of samples after stimulation of PBMCs with ancestral SARS-CoV-2 (B.1) spike MPs for 24 h at 37 °C. Four hours before staining, cells were treated with protein transport inhibitor BD GolgiPlug (BD Biosciences), followed by washing and surface staining with CD20, CD14, CD16 and fixable-viability dye coupled with APC eflour 780 in the dump channel and CD4-AlexaFluor 700 (RPA−T4). Later, cells were washed and treated with fixative buffer (eBioscience). After fixation, cells were washed with permeabilization buffer (eBioscience) and intracellular cellular staining was performed with IFN-γ PE-CF594 (B27) for 1 h in the dark, followed by acquisition using BD LSRFortessa X-20 flow cytometer (BD Biosciences) and subsequent data analysis by FlowJo 10.7.1.
Bead-based multiplex cytokine immunoassay
The cytokine levels were measured in the culture supernatants of PBMCs stimulated for 24 h with SARS-CoV-2 spike PeptideMP. The cell-free supernatants were stored at −80 °C until thawed for the quantification of secreted factors. The T-cell-specific secreted cytokines were quantified using the Bio-Plex human cytokine screening panel (17-Plex, Bio-Rad) on a Luminex 200 multiplex suspension array system following the manufacturer’s instruction. Positive cytokine response was defined as a response above the lowest limit of quantification and significance of P < 0.05 between unstimulated and spike PeptideMP-stimulated conditions.
Qualitative immunoassay for SARS-CoV-2-specific CD4+ T cells
The B-cell help quality of antigen-specific memory CD4+ T cells in vaccination and infection was assessed in the newly described autologous T-B co-culture assay51. Briefly, cryopreserved PBMCs were revived as mentioned previously with ≥90% cellular viability and surface stained with the following monoclonal antibodies for 15 min at 4 °C: fixable-viability dye efluor 506, anti-CD4 AF700 (RPA−T4), anti-CD45RO FITC (UCHL1), anti-CD14 PE (MΦP9) and anti-CD20 PE-Cy7 (2H7). Memory CD4+ T cells sorted as live CD14−CD20−CD4+CD45RO+ cells. Live CD20+CD14− cells and CD14+CD20− cells were sorted as B cells and monocytes, respectively. Cells were sorted on a BD FACSAria Fusion flow cytometer (BD Biosciences) at low flow rate using a 70 μm nozzle. Autologous memory CD4+ T cells and CD20+ B cells were co-cultured in a 1:1 ratio (6 × 104 cells per well) in the presence of monocytes (3 × 104 cells) in AIM-V serum-free medium (Gibco) in 96-well plates (Nunc, ThermoFisher). Cells were kept either unstimulated (no exogenous stimulation) or stimulated with SARS-CoV-2 full-length spike protein (CoV-S protein, 10 μg ml−1) (Native Antigen) for 9 d. Subsequently, enumeration of plasma cells as CD20loCD38hi in CD27+CD19+ B cells was performed by flow cytometry. Cells were surface stained with the following antibodies for 40 min at 4 °C: fixable-viability dye efluor 506 (eBioscience), anti-CD3 APC-Cy7 (HIT3a), anti-CD19 BV786 (HIB19), anti-CD27 PE-Dazzle 594 (M-T271), anti-CD20 PE-Cy7 (2H7) and anti-CD38 PE-Cy5 (HIT2). After surface staining, cells were washed with FACS buffer (2% FBS in PBS) and resuspended in FACS buffer. All samples were acquired on BD LSRFortessa X-20 flow cytometer (BD Biosciences). Data were analysed using FlowJo 10.7.1.
IL-21 ELISpot
IL-21-secreting T cells were quantified using human IL-21 ELISpot (3540; Mabtech) in the cells obtained from T-B co-cultures after 9 d of stimulation with spike protein. Briefly, a multiscreen ELISpot plate (MSIPS4510; Millipore) was coated with mAb MT216G (10 μg ml−1, Mabtech) overnight at 4 °C. The plate was washed and blocked with 10% FBS for at least 30 min at room temperature. After blocking of the wells, cells from co-cultures were seeded and restimulated with SARS-CoV-2 spike protein (2 μg ml−1) for 24 h at 37 °C. Next day, the plate was washed vigorously with PBS and developed by incubating with detection antibody mAb MT21.3m-biotin (Mabtech; dilution, 1:2,000) for 2 h, followed by the addition of streptavidin-ALP (dilution, 1:1,000) and BCIP/NBT substrate. The plate was dried and read on an AID vSpot Spectrum ELISpot/Fluorospot reader system. IL-21 SFCs were captured and enumerated using AID ELISpot software version 7.x.
IFN-γ FluoroSpot
To enumerate IFN-γ-secreting cells, FluoroSpot assay was performed according to the manufacturer’s (Mabtech) protocol. A low- fluorescence IPFL 96-well plate (Mabtech) was coated with 15 μg ml−1 of purified mAb IFN-γ (clone 1-D1K, Mabtech) overnight at 4 °C. After incubation, the plate was washed, blocked and seeded with 250 × 103 PBMCs, followed by stimulation with SARS-CoV-2 spike PeptideMP at 1 µg ml−1 for 24 h. Equimolar DMSO was used as negative control and staphylococcal enterotoxin B at 0.1 µg ml−1 was used as positive stimulation control. After incubation, IFN-γ spots were detected using BAM (bm-Amyloid-β)-conjugated monoclonal antibody against IFN-γ (7−B6-1-BAM, Mabtech; dilution, 1:200). The plate was developed by incubating with the anti-BAM 490 (Mabtech; dilution, 1:200) and addition of fluorescent enhancer (Mabtech). The plate was dried and spots were read using the AID vSpot Spectrum ELISpot/FluoroSpot reader system and quantified using AID ELISpot software version 7.x.
Statistical analyses
In all experiments, data are either represented as median, or as mentioned in the figure legends. The significance of the differences between the groups was analysed with a two-sided Mann-Whitney test, Fisher’s exact test, two-tailed Wilcoxon matched-pairs signed-rank test, two-way analysis of variance (ANOVA) or as specified in the figure legends. P values <0.05 were considered statistically significant. Statistical analyses and data visualization were performed with the GraphPad Prism software version v8.4.3.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
All data generated or analysed during this study are included in this Article. Source data are provided with this paper.
References
Ella, R. et al. Safety and immunogenicity of an inactivated SARS-CoV-2 vaccine, BBV152: interim results from a double-blind, randomised, multicentre, phase 2 trial, and 3-month follow-up of a double-blind, randomised phase 1 trial. Lancet Infect. Dis. 21, 950–961 (2021).
Ella, R. et al. Efficacy, safety, and lot-to-lot immunogenicity of an inactivated SARS-CoV-2 vaccine (BBV152): interim results of a randomised, double-blind, controlled, phase 3 trial. Lancet https://doi.org/10.1016/S0140-6736(21)02000-6 (2021).
Ansari, A. et al. Immune memory in mild COVID-19 patients and unexposed donors reveals persistent T cell responses after SARS-CoV-2 infection. Front. Immunol. 12, 636768 (2021).
Dan, J. M. et al. Immunological memory to SARS-CoV-2 assessed for up to 8 months after infection. Science https://doi.org/10.1126/science.abf4063 (2021).
Zuo, J. et al. Robust SARS-CoV-2-specific T cell immunity is maintained at 6 months following primary infection. Nat. Immunol. 22, 620–626 (2021).
Gaebler, C. et al. Evolution of antibody immunity to SARS-CoV-2. Nature 591, 639–644 (2021).
Grifoni, A. et al. Targets of T cell responses to SARS-CoV-2 coronavirus in humans with COVID-19 disease and unexposed individuals. Cell 181, 1489–1501.e15 (2020).
Sekine, T. et al. Robust T cell immunity in convalescent individuals with asymptomatic or mild COVID-19. Cell 183, 158–168.e14 (2020).
Braun, J. et al. SARS-CoV-2-reactive T cells in healthy donors and patients with COVID-19. Nature 587, 270–274 (2020).
Rydyznski Moderbacher, C. et al. Antigen-specific adaptive immunity to SARS-CoV-2 in acute COVID-19 and associations with age and disease severity. Cell 183, 996–1012.e19 (2020).
Tan, A. T. et al. Early induction of functional SARS-CoV-2-specific T cells associates with rapid viral clearance and mild disease in COVID-19 patients. Cell Rep. 34, 108728 (2021).
Bonifacius, A. et al. COVID-19 immune signatures reveal stable antiviral T cell function despite declining humoral responses. Immunity 54, 340–354.e6 (2021).
Turner, J. S. et al. SARS-CoV-2 infection induces long-lived bone marrow plasma cells in humans. Nature 595, 421–425 (2021).
Kalimuddin, S. et al. Early T cell and binding antibody responses are associated with COVID-19 RNA vaccine efficacy onset. Med 2, 682–688.e4 (2021).
Polack, F. P. et al. Safety and efficacy of the BNT162b2 mRNA Covid-19 vaccine. N. Engl. J. Med. 383, 2603–2615 (2020).
Madhi, S. A. et al. Efficacy of the ChAdOx1 nCoV-19 Covid-19 vaccine against the B.1.351 Variant. N. Engl. J. Med. 384, 1885–1898 (2021).
Altmann, D. M., Boyton, R. J. & Beale, R. Immunity to SARS-CoV-2 variants of concern. Science 371, 1103–1104 (2021).
Thiruvengadam, R. et al. Effectiveness of ChAdOx1 nCoV-19 vaccine against SARS-CoV-2 infection during the delta (B.1.617.2) variant surge in India: a test-negative, case-control study and a mechanistic study of post-vaccination immune responses. Lancet Infect Dis. 22, 473–482 (2021).
Ella, R. et al. Safety and immunogenicity of an inactivated SARS-CoV-2 vaccine, BBV152: a double-blind, randomised, phase 1 trial. Lancet Infect. Dis. 21, 637–646 (2021).
Jackson, L. A. et al. An mRNA vaccine against SARS-CoV-2 – preliminary report. N. Engl. J. Med. 383, 1920–1931 (2020).
Buchrieser, J. et al. Syncytia formation by SARS-CoV-2-infected cells. EMBO J. 39, e106267 (2020).
Tan, C. W. et al. A SARS-CoV-2 surrogate virus neutralization test based on antibody-mediated blockage of ACE2-spike protein-protein interaction. Nat. Biotechnol. 38, 1073–1078 (2020).
Planas, D. et al. Reduced sensitivity of SARS-CoV-2 variant Delta to antibody neutralization. Nature 596, 276–280 (2021).
Wall, E. C. et al. Neutralising antibody activity against SARS-CoV-2 VOCs B.1.617.2 and B.1.351 by BNT162b2 vaccination. Lancet 397, 2331–2333 (2021).
Liu, C. et al. Reduced neutralization of SARS-CoV-2 B.1.617 by vaccine and convalescent serum. Cell 184, 4220–4236 e4213 (2021).
Zhou, D. et al. Evidence of escape of SARS-CoV-2 variant B.1.351 from natural and vaccine-induced sera. Cell 184, 2348–2361.e6 (2021).
Muecksch, F. et al. Affinity maturation of SARS-CoV-2 neutralizing antibodies confers potency, breadth, and resilience to viral escape mutations. Immunity 54, 1853–1868 e1857 (2021).
Cho, A. et al. Anti-SARS-CoV-2 receptor binding domain antibody evolution after mRNA vaccination. Nature https://doi.org/10.1038/s41586-021-04060-7 (2021).
Wang, Z. et al. Enhanced SARS-CoV-2 neutralization by dimeric IgA. Sci. Transl. Med. https://doi.org/10.1126/scitranslmed.abf1555 (2021).
Sterlin, D. et al. IgA dominates the early neutralizing antibody response to SARS-CoV-2. Sci. Transl. Med. https://doi.org/10.1126/scitranslmed.abd2223 (2021).
Tarke, A. et al. Impact of SARS-CoV-2 variants on the total CD4(+) and CD8(+) T cell reactivity in infected or vaccinated individuals. Cell Rep. Med. 2, 100355 (2021).
Chen, G. et al. Clinical and immunological features of severe and moderate coronavirus disease 2019. J. Clin. Invest. 130, 2620–2629 (2020).
Weiskopf, D. et al. Phenotype and kinetics of SARS-CoV-2-specific T cells in COVID-19 patients with acute respiratory distress syndrome. Sci. Immunol. https://doi.org/10.1126/sciimmunol.abd2071 (2020).
Mateus, J. et al. Low-dose mRNA-1273 COVID-19 vaccine generates durable memory enhanced by cross-reactive T cells. Science 374, eabj9853 (2021).
Folegatti, P. M. et al. Safety and immunogenicity of the ChAdOx1 nCoV-19 vaccine against SARS-CoV-2: a preliminary report of a phase 1/2, single-blind, randomised controlled trial. Lancet 396, 467–478 (2020).
Wille-Reece, U. et al. Toll-like receptor agonists influence the magnitude and quality of memory T cell responses after prime-boost immunization in nonhuman primates. J. Exp. Med. 203, 1249–1258 (2006).
Darrah, P. A. et al. Multifunctional TH1 cells define a correlate of vaccine-mediated protection against Leishmania major. Nat. Med. 13, 843–850 (2007).
Cohen, K. W. et al. Longitudinal analysis shows durable and broad immune memory after SARS-CoV-2 infection with persisting antibody responses and memory B and T cells. Cell Rep. Med. 2, 100354 (2021).
Sallusto, F., Geginat, J. & Lanzavecchia, A. Central memory and effector memory T cell subsets: function, generation, and maintenance. Annu. Rev. Immunol. 22, 745–763 (2004).
Crotty, S. T follicular helper cell differentiation, function, and roles in disease. Immunity 41, 529–542 (2014).
Juno, J. A. et al. Humoral and circulating follicular helper T cell responses in recovered patients with COVID-19. Nat. Med. 26, 1428–1434 (2020).
Painter, M. M. et al. Rapid induction of antigen-specific CD4(+) T cells is associated with coordinated humoral and cellular immunity to SARS-CoV-2 mRNA vaccination. Immunity 54, 2133–2142.e3 (2021).
Goel, R. R. et al. mRNA vaccines induce durable immune memory to SARS-CoV-2 and variants of concern. Science 374, abm0829 (2021).
Schmitt, N., Bentebibel, S. E. & Ueno, H. Phenotype and functions of memory Tfh cells in human blood. Trends Immunol. 35, 436–442 (2014).
Crotty, S. T follicular helper cell biology: a decade of discovery and diseases. Immunity 50, 1132–1148 (2019).
Kuri-Cervantes, L. et al. Comprehensive mapping of immune perturbations associated with severe COVID-19. Sci. Immunol. https://doi.org/10.1126/sciimmunol.abd7114 (2020).
Baumjohann, D. et al. Persistent antigen and germinal center B cells sustain T follicular helper cell responses and phenotype. Immunity 38, 596–605 (2013).
Poon, M. M. L. et al. SARS-CoV-2 infection generates tissue-localized immunological memory in humans. Sci. Immunol. https://doi.org/10.1126/sciimmunol.abl9105 (2021).
Locci, M. et al. Human circulating PD-1+CXCR3-CXCR5+ memory Tfh cells are highly functional and correlate with broadly neutralizing HIV antibody responses. Immunity 39, 758–769 (2013).
Morita, R. et al. Human blood CXCR5(+)CD4(+) T cells are counterparts of T follicular cells and contain specific subsets that differentially support antibody secretion. Immunity 34, 108–121 (2011).
Ansari, A. et al. An efficient immunoassay for the B-cell help function of SARS-CoV-2 specific memory CD4+ T cells. Cell Rep. Methods https://doi.org/10.1016/j.crmeth.2022.100224 (2022).
Shrivastava, T. et al. Comparative immunomodulatory evaluation of the receptor binding domain of the SARS-CoV-2 spike protein; a potential vaccine candidate which imparts potent humoral and Th1 type immune response in a mouse model. Front. Immunol. 12, 641447 (2021).
Hu, J. et al. Development of cell-based pseudovirus entry assay to identify potential viral entry inhibitors and neutralizing antibodies against SARS-CoV-2. Genes Dis. 7, 551–557 (2020).
Acknowledgements
We thank participants, staff of partner medical centres and DBT NCR Cluster COVID-19 Biorepository for their contributions to this study; P. Garg, THSTI for scientific discussion and reviewing the manuscript; N., J. and R. Prasad for technical help. Some schematics (Figs. 1a, 2a, 4e and ED Fig. 2a) were created with Biorender.com. This work was financially supported by SERB, DST grant (IPA/2020/000077), DBT grant (BT/PR30223/MED/2018) to N.G. and NII core grant. Further support was provided by a DBT grant (BT/PR40401/BIOBANK/03.2020) to S.B. and NIH contract 75N93021C00016 to A. Sette and 75N9301900065 to A. Sette and D.W.
Author information
Authors and Affiliations
Contributions
N.G. conceptualized and supervised the study and wrote the original draft; S.B. supervised the cohort establishment; R.V., A.A., A.R.P., S.N.J., S. Sachan, S.P., B.N. and A.K. carried out the investigation; H.A.P. and T.S. provided the RBD proteins; B.P.J., S. Singh, P.K., N.W., P.C., S.K., P.S., N.S., J.T., A.K.P., A. Sharma and R.T. supervised enrolment and sample collection; A.G., D.W. and A. Sette provided the peptide pools; A. Sette, S.B. and N.G. acquired funding. All authors edited and reviewed the final manuscript.
Corresponding author
Ethics declarations
Competing interests
N.G. and A.A. are listed as inventors on Indian patent application no. 202111003148 filed in 2021 (submitted by NII) that covers the use of ‘T-cell qualitative immunoassay’ for vaccine evaluation. A. Sette is consultant for GritstoneBio, FlowPharma, ArcturusTherapeutics, ImmunoScape, CellCarta, Avalia, Moderna, Fortress and Repertoire. LJI has filed USA patent protection for various aspects of T-cell epitope and vaccine design work. All other authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Jincun Zhao and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 BBV152-elicited neutralizing antibodies persisted at least up to 6 months after complete vaccination.
(a) The potential of antibodies in inhibiting the RBD-ACE2 interaction was measured by the SARS-CoV-2 surrogate virus neutralization test in vaccinated (n = 24) and recovered individuals (n = 24). (b) Neutralizing antibody titer in BBV152 vaccinated (n = 52) and COVID-19 recovered individuals (n = 33) as measured in the pseudovirus neutralization assay using the pseudovirus expressing full-length spike protein of the ancestral SARS-CoV-2 and its variants; Alpha, Beta and Delta. Correlation of the respective ancestral SARS-CoV-2 pseudovirus neutralizing antibody titres with (c) Spike binding IgG, (d) RBD binding IgG in vaccinated individuals, and (e) Spike binding IgG (f) RBD binding IgG in individuals recovered from mild COVID-19. Persistence of (g) anti-spike (h) anti-nucleoprotein (NP) IgG and (i) pseudovirus neutralizing antibodies over the d post vaccination, and (j) anti-spike (k) anti-nucleoprotein IgG and (l) pseudovirus neutralizing antibodies over the days post-diagnosis of COVID-19, in BBV152 vaccination and recovery from mild COVID-19, respectively. Black bars indicate geometrical mean and geometric SD in (a-b). Dotted line represents the cut-off for positivity. Statistics by (a-b) two-tail Mann-Whitney test, (c-f) two-tail Spearman correlation coefficient, (g-l) non-linear regression analysis. ns: non-significant.
Extended Data Fig. 2 Pseudovirus Neutralization Assay.
(a) Scheme (generated using Biorender) depicting the pseudovirus neutralization assay performed in BBV152 vaccinated and COVID-19 mild disease recovered individuals using pseudovirus expressing full-length spike protein of the ancestral SARS-CoV-2 and its variants; Alpha (B.1.1.7), Beta (B.1.351) and Delta (B.1.617.2). (b) Neutralization curves for plasma samples derived from the representative pre-pandemic healthy donors (n = 10), BBV152 vaccinated (n = 15) and COVID-19 recovered individuals (n = 15) corresponding to the ancestral SARS-CoV-2 and the VoCs spike pseudoviruses.
Extended Data Fig. 3 BBV152-induced memory B cells correlates with antibody titers and persisted in circulation at least up to 6 months post-vaccination.
The RBD-specific memory B cells were enumerated in the R848 + IL2 stimulated B cells utilizing the antigen-specific ELIspot analysis. Representative ELIspot images of IgG+, IgM+ and IgA+ memory B cells against RBD of ancestral SARS-CoV-2 and its variants in (a) vaccinated and (b) recovered COVID-19 subjects. ‘Total’ represent the frequency of total B cells captured using anti-IgG, -IgM or -IgA antibody. Proportion of SARS-CoV-2-RBD-specific antibody secreting cells (ASCs) was measured by calculating the percentage of antigen-specific B cells in the total corresponding isotype of ASCs in 1 million PBMCs. Correlation of RBD-specific IgG+ memory B cells with (c) RBD binding IgG, (d) pseudovirus neutralizing antibodies, and (e) persistence of IgG+ memory B cells over the days post vaccination, and (f) RBD binding IgG, (g) pseudovirus neutralizing antibodies, and (h) persistence of IgG+ memory B cells over the days post-diagnosis of COVID-19, in BBV152 vaccination and recovery from mild COVID-19, respectively. Dotted line represents the cut-off for positivity. Statistics by (c-d, f-g) two-tail Pearson correlation coefficient. ns: non-significant.
Extended Data Fig. 4 Flow cytometry Gating Strategy.
(a) gating strategy for quantitation of the antigen-specific CD4+ T cells as determined by OX40+CD137+ (AIM+) cells, total and antigen-specific memory CD4+ T-cell subsets, and circulating total and SARS-CoV-2-specific follicular helper T (Tfh) cells in the PBMCs derived from individuals vaccinated with BBV152 and individuals recovered from mild COVID-19. (b) gating strategy for quantitation of the antigen-specific CD8+ T cells as determined by CD69+CD137+ (AIM+) cells in the PBMCs derived from individuals as described for (a).
Extended Data Fig. 5 Comparison of AIM and ICS assay for the quantitative measurement of SARS-CoV-2 specific CD4+ T cells.
To validate quantification of SARS-CoV-2 specific T cells by AIM assay, IFN-γ intracellular cytokine staining (ICS) and AIM was performed on the PBMCs after stimulation with ancestral SARS-CoV-2 spike MPs for 24 hours at 37 °C. In the Activation-induced markers (AIM) assay, antigen-specific CD4+ T cell were quantified as a percentage population of AIM+ (OX40+CD137+) CD4+ T cells. In the IFN-γ ICS assay, antigen-specific CD4+ T cell were quantified as a percentage population of IFN-γ+CD4+ T cells. The graph depicts the correlation between SARS-CoV-2 spike-specific AIM+ CD4+ T cells and IFN-γ+ CD4+ T cells measured in respective PBMCs (n = 8) from BBV152 vaccinated and COVID-19 recovered individuals. Statistics by two-tail Pearson correlation coefficient.
Extended Data Fig. 6 BBV152 vaccination induces IFN-γ expressing cells.
The IFN-γ spot forming cells were measured in the PBMCs derived from individuals with BBV152 vaccination (n = 10) after stimulation with overlapping peptide megapools (MP) spanning spike protein of SARS-CoV-2 in the IFN- γ Fluorospot assay. (a) Representative images of IFN-γ spot forming cells (SFCs) in PBMCs from two vaccinated donors, stimulated with DMSO (Unstim), spike peptideMP (spike), and Staphylococcal Enterotoxin B (SEB) as a positive control. (b) The magnitude of IFN-γ spot forming cells (SFCs) per million PBMCs was compared between unstimulated and the spike peptideMP stimulated conditions. Statistics by two-tail Wilcoxon signed-rank test.
Extended Data Fig. 7 BBV152 vaccination generates multifunctional spike-specific CD4+ T-cell response with the polarization towards Th1-cell phenotype.
Cytokine levels were measured by multiplex bead-based immunoassay in the cell culture supernatants of PBMCs stimulated or not with spike peptideMP for 24 h, and compared between BBV152 vaccination (n = 23) and recovery from mild COVID-19 (n = 20). The standard curve depicting the fluorescence intensity (FI) of cytokine-standard and the adjacent bar graph showing the concentration (pg/ml) of T-cell specific cytokines in vaccination and infection for (a) IFN-γ (b) IL4 (c) IL17 (d) IL13 (e) IL2 and (f) TNF-α. (g) Stacked graph showing the comparison of mean concentration of each cytokine in BBV152 vaccination and recovery from mild COVID-19. The data denotes the mean±s.e.m. The positive response was determined as the response over Lower Limit of Quantification (LLOQ) and statistical significance in spike peptideMP stimulation over unstimulated conditions. Statistics by two-tail Mann-Whitney test. ns: non-significant.
Extended Data Fig. 8 BBV152 vaccination induces SARS-CoV-2 specific CD8+ T cells.
SARS-CoV-2-specific CD8+ T cells were quantified in PBMCs derived from the individuals vaccinated with BBV152 (‘BBV152’) and individuals recovered from mild COVID-19 (‘COVID-19’). The SARS-CoV-2-specific CD8+ T cells were quantified in AIM assay as a percentage population of AIM+ (CD69+CD137+) CD8+ T cells, after stimulation of PBMCs with overlapping peptide megapools (MP) spanning spike glycoprotein of ancestral SARS-CoV-2, nucleoprotein of SARS-CoV-2 or the MP specific to the spike protein of VOCs. Equimolar DMSO was used as the negative control and CMV MP was used as the positive stimulation control. Representative gating strategy for measuring the antigen-specific AIM+ (CD69+CD137+) CD8+ T cells in the total CD8+ T cells from (a) BBV152 and (b) COVID-19 (see Extended data Fig. 4 for detailed gating strategy). The numbers on the lower right corners of the contour plots represent the frequency of AIM+ cells in the total CD8+ T cells. (c) Frequency of total CD8+ T cells in BBV152 and COVID-19, (d) Comparison of background subtracted spike-specific CD8+ T cells in BBV152 and COVID-19 groups, (e) Comparison of background subtracted nucleoprotein-specific CD8+ T cells in BBV152 and COVID-19, (f) CMV-specific CD8+ T cells in BBV152 versus COVID-19. Quantitation of spike-specific CD8+ T cells responding to ancestral SARS-CoV-2 and VOCs – Alpha (B.1.1.7), Beta (B.1.351) and Delta (B.1.617.2) in individuals after (g) vaccination and (h) recovery from COVID-19. Values in parentheses represents fold-reduction in spike-specific CD8+ T cells responding to VOCs over ancestral virus. Black bars indicate median. Dotted line represents the limit of positivity. Statistics by (c-f) two-tail Mann-Whitney test, (g-h) two-tail Wilcoxon signed-rank test. ns: non-significant.
Extended Data Fig. 9 BBV152-induced circulating Tfh cells comprised of both Tfh1 and Tfh2 phenotype.
The functional heterogeneity of spike and nucleoprotein (NP) specific circulating follicular helper T (cTfh) cells was determined based on the expression of CXCR3 and CCR6 in total Tfh cells (CXCR5+) among the AIM+ antigen-specific CD4+ T cells. The proportion of Tfh subsets was measured in the SARS-CoV-2 specific cTfh cells induced in response to BBV152 vaccination or mild COVID-19. (a) Representative gating strategy for measuring the Tfh heterogeneity in SARS-CoV-2-specific CD4+ T cells. Proportion of Tfh17 (CXCR3−CCR6+), Tfh1 (CXCR3+CCR6−), and Tfh2 (CXCR3−CCR6−) subsets among (b) spike-specific cTfh and (c) nucleoprotein-specific cTfh cells in BBV152 (spike: n = 38; NP: n = 25) and COVID-19 (spike: n = 37; NP: n = 19) groups. Black bars indicate median. Statistics by 2way ANOVA followed by Tukey’s multiple comparisons. ns: non-significant.
Extended Data Fig. 10 BBV152-induced spike specific AIM+ cTfh cells correlated with the neutralizing antibodies.
The correlation of SARS-CoV-2 spike-specific CD4+ T cells and cTfh cells with the spike-binding and neutralizing antibodies was examined in BBV152 vaccinated (top panel) and COVID-19 recovered individuals (bottom panel). The graphs depict the correlation of the AIM+CD4+ T cells with (a) Spike binding IgG, (b) pseudovirus neutralizing antibodies and correlation of the AIM+ total cTfh cells with (c) Spike binding IgG, and (d) pseudovirus neutralizing antibodies in BBV152 vaccinated individuals, and similar correlations (e-h) in the individuals recovered from mild COVID-19. Dotted line represents the cut-off for positivity. Statistics by two-tail Spearman correlation coefficient. ns: non-significant.
Supplementary information
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
About this article
Cite this article
Vikkurthi, R., Ansari, A., Pai, A.R. et al. Inactivated whole-virion vaccine BBV152/Covaxin elicits robust cellular immune memory to SARS-CoV-2 and variants of concern. Nat Microbiol 7, 974–985 (2022). https://doi.org/10.1038/s41564-022-01161-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-022-01161-5
This article is cited by
-
Polyfunctional CD4 T-cells correlating with neutralising antibody is a hallmark of COVISHIELDTM and COVAXIN® induced immunity in COVID-19 exposed Indians
npj Vaccines (2023)
-
Persistence of immunity and impact of third dose of inactivated COVID-19 vaccine against emerging variants
Scientific Reports (2022)
-
COVID-19 vaccination boosts the potency and breadth of the immune response against SARS-CoV-2 among recovered patients in Wuhan
Cell Discovery (2022)