Abstract
Anthropogenic climate change threatens ecosystem functioning. Soil biodiversity is essential for maintaining the health of terrestrial systems, but how climate change affects the richness and abundance of soil microbial communities remains unresolved. We examined the effects of warming, altered precipitation and annual biomass removal on grassland soil bacterial, fungal and protistan communities over 7 years to determine how these representative climate changes impact microbial biodiversity and ecosystem functioning. We show that experimental warming and the concomitant reductions in soil moisture play a predominant role in shaping microbial biodiversity by decreasing the richness of bacteria (9.6%), fungi (14.5%) and protists (7.5%). Our results also show positive associations between microbial biodiversity and ecosystem functional processes, such as gross primary productivity and microbial biomass. We conclude that the detrimental effects of biodiversity loss might be more severe in a warmer world.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The DNA sequences of the 16S rRNA gene, 18S rRNA gene and ITS amplicons are deposited in the National Center for Biotechnology Information (NCBI) under project accession number PRJNA331185. Raw shotgun metagenomic sequences are deposited in the European Nucleotide Archive (http://www.ebi.ac.uk/ena) under study no. PRJNA533082. Silva 138.1 Ref NR database is available at https://www.arb-silva.de/documentation/release-138/. Protist Ribosomal Reference database (PR2) databases are available at https://github.com/pr2database/pr2database. The ASV table and ASV representative sequences, soil physical and chemical attributes, and plant biomass and richness are downloadable online at http://www.ou.edu/ieg/publications/datasets. Source data are provided with this paper.
Code availability
R scripts for statistical analyses are available on GitHub at https://github.com/Linwei-Wu/warming_soil_biodiversity.
References
Rands, M. R. et al. Biodiversity conservation: challenges beyond 2010. Science 329, 1298–1303 (2010).
Diaz, S., Fargione, J., Chapin, F. S. III & Tilman, D. Biodiversity loss threatens human well-being. PLoS Biol. 4, e277 (2006).
Barnosky, A. D. et al. Has the Earth’s sixth mass extinction already arrived? Nature 471, 51–57 (2011).
Pecl, G. T. et al. Biodiversity redistribution under climate change: impacts on ecosystems and human well-being. Science https://doi.org/10.1126/science.aai9214 (2017).
IPCC Climate Change 2013: The Physical Science Basis (eds Stocker, T. F. et al.) (Cambridge Univ. Press, 2013).
Cardinale, B. J. et al. Biodiversity loss and its impact on humanity. Nature 486, 59–67 (2012).
Hautier, Y. et al. Eutrophication weakens stabilizing effects of diversity in natural grasslands. Nature 508, 521–525 (2014).
Bascompte, J., García, M. B., Ortega, R., Rezende, E. L. & Pironon, S. Mutualistic interactions reshuffle the effects of climate change on plants across the tree of life. Sci. Adv. 5, eaav2539 (2019).
Blois, J. L., Zarnetske, P. L., Fitzpatrick, M. C. & Finnegan, S. Climate change and the past, present, and future of biotic interactions. Science 341, 499–504 (2013).
Tylianakis, J. M., Didham, R. K., Bascompte, J. & Wardle, D. A. Global change and species interactions in terrestrial ecosystems. Ecol. Lett. 11, 1351–1363 (2008).
Fei, S. et al. Divergence of species responses to climate change. Sci. Adv. 3, e1603055 (2017).
Li, D., Miller, J. E. D. & Harrison, S. Climate drives loss of phylogenetic diversity in a grassland community. Proc. Natl Acad. Sci. USA 116, 19989–19994 (2019).
Bay, R. A. et al. Genomic signals of selection predict climate-driven population declines in a migratory bird. Science 359, 83–86 (2018).
Xue, K. et al. Annual removal of aboveground plant biomass alters soil microbial responses to warming. mBio https://doi.org/10.1128/mBio.00976-16 (2016).
Zhou, J. et al. Microbial mediation of carbon-cycle feedbacks to climate warming. Nat. Clim. Change 2, 106–110 (2012).
Steidinger, B. S. et al. Climatic controls of decomposition drive the global biogeography of forest-tree symbioses. Nature 569, 404–408 (2019).
Blankinship, J. C., Niklaus, P. A. & Hungate, B. A. A meta-analysis of responses of soil biota to global change. Oecologia 165, 553–565 (2011).
Guo, X. et al. Climate warming leads to divergent succession of grassland microbial communities. Nat. Clim. Change 8, 813–818 (2018).
Guo, X. et al. Climate warming accelerates temporal scaling of grassland soil microbial biodiversity. Nat. Ecol. Evol. 3, 612–619 (2019).
Yuan, M. M. et al. Climate warming enhances microbial network complexity and stability. Nat. Clim. Change 11, 343–348 (2021).
Thakur, M. P. et al. Climate warming promotes species diversity, but with greater taxonomic redundancy, in complex environments. Sci. Adv. 3, e1700866 (2017).
Xu, X., Sherry, R. A., Niu, S., Li, D. & Luo, Y. Net primary productivity and rain‐use efficiency as affected by warming, altered precipitation, and clipping in a mixed‐grass prairie. Glob. Change Biol. 19, 2753–2764 (2013).
Luo, Y., Sherry, R., Zhou, X. & Wan, S. Terrestrial carbon‐cycle feedback to climate warming: experimental evidence on plant regulation and impacts of biofuel feedstock harvest. Glob. Change Biol. Bioenergy 1, 62–74 (2009).
Chen, M.-M. et al. Effects of soil moisture and plant interactions on the soil microbial community structure. Eur. J. Soil Biol. 43, 31–38 (2007).
Zhou, J. et al. Temperature mediates continental-scale diversity of microbes in forest soils. Nat. Commun. 7, 12083 (2016).
Rousk, J. et al. Soil bacterial and fungal communities across a pH gradient in an arable soil. ISME J. 4, 1340–1351 (2010).
DeBruyn, J. M., Nixon, L. T., Fawaz, M. N., Johnson, A. M. & Radosevich, M. Global biogeography and quantitative seasonal dynamics of Gemmatimonadetes in soil. Appl. Environ. Microbiol. 77, 6295–6300 (2011).
Van Horn, D. J. et al. Soil microbial responses to increased moisture and organic resources along a salinity gradient in a polar desert. Appl. Environ. Microbiol. 80, 3034–3043 (2014).
Van Nuland, M. E. et al. Warming and disturbance alter soil microbiome diversity and function in a northern forest ecotone. FEMS Microbiol. Ecol. 96, fiaa108 (2020).
Reimer, L. C. et al. Bac Dive in 2022: the knowledge base for standardized bacterial and archaeal data. Nucleic Acids Res. 50, D741–D746 (2022).
Nguyen, N. H. et al. FUNGuild: an open annotation tool for parsing fungal community datasets by ecological guild. Fungal Ecol. 20, 241–248 (2016).
Delgado-Baquerizo, M. et al. Microbial diversity drives multifunctionality in terrestrial ecosystems. Nat. Commun. 7, 10541 (2016).
Banerjee, S. et al. Agricultural intensification reduces microbial network complexity and the abundance of keystone taxa in roots. ISME J. 13, 1722–1736 (2019).
Ning, D. et al. A quantitative framework reveals ecological drivers of grassland microbial community assembly in respoÿnse to warming. Nat. Commun. 11, 4717 (2020).
Tiedje, J. M. et al. Microbes and climate change: a research prospectus for the future. MBio, e00800-22 (2022). doi:10.1128/mbio.00800-22 (2022).
Maestre, F. T. et al. Increasing aridity reduces soil microbial diversity and abundance in global drylands. Proc. Natl Acad. Sci. USA 112, 15684–15689 (2015).
Li, D., Zhou, X., Wu, L., Zhou, J. & Luo, Y. Contrasting responses of heterotrophic and autotrophic respiration to experimental warming in a winter annual‐dominated prairie. Glob. Change Biol. 19, 3553–3564 (2013).
Sherry, R. A. et al. Lagged effects of experimental warming and doubled precipitation on annual and seasonal aboveground biomass production in a tallgrass prairie. Glob. Change Biol. 14, 2923–2936 (2008).
Catchpole, W. & Wheeler, C. Estimating plant biomass: a review of techniques. Aust. J. Ecol. 17, 121–131 (1992).
McLean, E. in Methods of Soil Analysis. Part 2. Chemical and Microbiological Properties (ed Page, A. L.) 199–224 (1982).
Buyer, J. S. & Sasser, M. High throughput phospholipid fatty acid analysis of soils. Appl. Soil Ecol. 61, 127–130 (2012).
Zhou, J., Bruns, M. A. & Tiedje, J. M. DNA recovery from soils of diverse composition. Appl. Environ. Microbiol. 62, 316–322 (1996).
Wu, L. et al. Phasing amplicon sequencing on Illumina Miseq for robust environmental microbial community analysis. BMC Microbiol. 15, 125 (2015).
Zhou, J. et al. Reproducibility and quantitation of amplicon sequencing-based detection. ISME J. 5, 1303–1313 (2011).
Magoč, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Edgar, R. C. Updating the 97% identity threshold for 16S ribosomal RNA OTUs. Bioinformatics 34, 2371–2375 (2018).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2 – approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Munoz, R. et al. Release LTPs104 of the all-species living tree. Syst. Appl. Microbiol. 34, 169–170 (2011).
Nuccio, E. E. et al. Climate and edaphic controllers influence rhizosphere community assembly for a wild annual grass. Ecology 97, 1307–1318 (2016).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 73, 5261–5267 (2007).
Nilsson, R. H. et al. The UNITE database for molecular identification of fungi: handling dark taxa and parallel taxonomic classifications. Nucleic Acids Res. 47, D259–D264 (2018).
Guillou, L. et al. The Protist Ribosomal Reference database (PR2): a catalog of unicellular eukaryote small sub-unit rRNA sequences with curated taxonomy. Nucleic Acids Res. 41, D597–D604 (2013).
Oliverio, A. M. et al. The global-scale distributions of soil protists and their contributions to belowground systems. Sci. Adv. 6, eaax8787 (2020).
Andrews, S. FastQC: A Quality Control Tool For High Throughput Sequence Data (Babraham Bioinformatics, 2010).
Li, W. & Godzik, A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659 (2006).
Patel, R. K. & Jain, M. NGS QC Toolkit: a toolkit for quality control of next generation sequencing data. PLoS ONE 7, e30619 (2012).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2015).
Huson, D. H., Auch, A. F., Qi, J. & Schuster, S. C. MEGAN analysis of metagenomic data. Genome Res. 17, 377–386 (2007).
Kembel, S. W. et al. Picante: R tools for integrating phylogenies and ecology. Bioinformatics 26, 1463–1464 (2010).
R Core Team. R: A Language And Environment For Statistical Computing (R Foundation for Statistical Computing, 2014).
Oksanen, J. et al. Package ‘vegan’. Community Ecology Package v.2 (2013).
Bates, D., Mächler, M., Bolker, B. & Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Softw. 67, 1–48 (2015).
Fox, J. & Weisberg, S. An R Companion to Applied Regression (Sage Publications, 2018).
Barton, K. & Barton, M. K. Package ‘MuMIn’ v.1.18 (2015).
Carver, R. Practical Data Analysis with JMP (SAS Institute, 2019).
Kuhn, M. Building predictive models in R using the caret package. J. Stat. Softw. 28, 1–26 (2008).
Rosseel, Y. lavaan: An R Package for Structural Equation Modeling and More v.0.4-9 (BETA) (Ghent University, 2011).
Acknowledgements
We thank the acting director of the KAEFS field site, M. Bomgraars, and the numerous former and current members of the Institute for Environmental Genomics for their help in maintaining the long-term field experiment. This work was supported by the US Department of Energy, Office of Science, Genomic Science Program under Award Number DE-SC0004601 and DE-SC0010715, and the Office of the Vice President for Research at the University of Oklahoma to J.Z. X.G. and X.Z. were generously supported by the China Scholarship Council (CSC) to visit the University of Oklahoma. The data analyses performed by X.G. were also supported by the China Postdoctoral Science Foundation (2018M641327 and 2019T120101).
Author information
Authors and Affiliations
Contributions
All authors contributed intellectual input and assistance to this study. The original concepts were conceived by J.Z. and J.M.T. Field management was carried out by Linwei Wu, Y.Z., X.G., J.F., M.M.Y., J.K., Y.F., A.Z., D.N., J.M., S.J., S.H., Z.Y., Y.O. and Liyou Wu. Sampling collection, soil chemical and microbial characterization were carried out by M.M.Y., X.G., Linwei Wu, J.G., Z.G. and X.Z. Data analysis were done by Linwei Wu, Y.Z., X.G., S.L. and N.X. with assistance provided by D.N. and J.Z. All data analysis and integration were guided by J.Z. The manuscript was prepared by J.Z., Linwei Wu, Y.Z. and X.G., with significant input from J.M.T., Y.Y. and X.L. Considering their contributions in terms of site management, data collection, analyses and/or integration, Linwei Wu, Y. Z. and X.G. are listed as co-first authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Pankaj Trivedi, Kirsten Hofmockel and Noah Fierer for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 A schematic map of the field experimental treatments.
In the long-term climate change experiment, warming (+3 °C), half precipitation (−50% precipitation), and double precipitation (+100% precipitation) are primary factors, which are nested with clipping as the secondary factor (all the 24 southern subplots are under clipping treatment). Thus, this experiment has twelve single and combined treatments as follows: Control (N), Warming (W), Half precipitation (H), Double precipitation (D), Clipping (C), Warming & Half precipitation (WH), Warming & Double precipitation (WD), Warming & Clipping (WC), Half precipitation & Clipping (HC), Double precipitation & Clipping (DC), Warming & Half precipitation & Clipping (WHC), and Warming & Double precipitation & Clipping (WDC). Each of these treatments has four replicates in four different blocks. The site was established in July, 2009. Surface (0-15 cm) soil samples were collected annually from all plots at approximately the date of peak plant biomass in fall (September or October) from 2009 to 2016.
Extended Data Fig. 2 Effects of experimental treatments on soil and plant variables by linear mixed-effects models (LMMs).
a, Soil temperature; b, Soil moisture; c, Soil pH; d, Soil NO3−N; e, Soil NH4-N; f, Total plant biomass; and g, Plant richness. Data are presented as mean values ± standard errors of the estimated effect sizes. Statistical significance is based on Wald type II χ² tests (n = 360 independent soil samples). Significant effects are denoted by asterisks: *** P < 0.001, ** P < 0.01, * P < 0.05. W: Warming; P: Precipitation level; C: Clipping. Since the precipitation level is considered as a continuous variable in the LMM (0.5 for half precipitation, 1 for normal and 2 for double precipitation), only one regression coefficient of precipitation treatment would be derived by the LMM. The effect size of half precipitation (as compared to ambient precipitation) can be derived by multiplying the regression coefficient by −0.5, while the effect size of double precipitation (as compared to ambient precipitation) can be derived by multiplying the regression coefficient by 1.
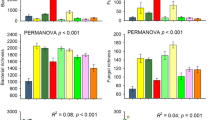
Extended Data Fig. 3 Sequencing depth among different treatments for the bacterial community (a), fungal community (b), and protistan community (c).
A total of 360 soil samples over 8 years were analyzed with 16S ribosomal RNA (rRNA) gene for bacteria and archaea, the internal transcribed spacer (ITS) between 5.8S and 28S rRNA genes for fungi, and the 18S rRNA gene for protists. For 18S sequences, only those annotated as protists were selected for the subsequent analyses. An average of 56,182 ± 27,613, 23,569 ± 16,323, and 11,146 ± 10,528 sequence reads were obtained for bacteria, fungi, and protists, respectively. There was no significant difference between treatments in the number of sequences (that is, sequencing depth) for the bacteria, fungal, and protistan communities except half precipitation (H), double precipitation (D), clipping (C), and double precipitation & clipping (DC) for protists (p = 0.002-0.021). Groups: Control (N), Warming (W), Half precipitation (H), Double precipitation (D), Clipping (C), Warming & Half precipitation (WH), Warming & Double precipitation (WD), Warming & Clipping (WC), Half precipitation & Clipping (HC), Double precipitation & Clipping (DC), Warming & Half precipitation & Clipping (WHC), and Warming & Double precipitation & Clipping (WDC). In the box plots, hinges show the 25, 50, and 75 percentiles. The upper whisker extends to the largest value no further than 1.5 * IQR from the upper hinge, where IQR is the inter-quartile range between the 25% and 75% quartiles; the lower whisker extends to the smallest value at most 1.5 * IQR from the lower hinge.
Extended Data Fig. 4 Rarefaction curves.
The number of ASVs with an increasing number of sequences (a, c, e) and accumulation curves of the number of ASVs with an increasing number of samples (b, d, f) for the bacterial community (a, b), fungal community (c, d), and protistan community (e, f). The observed number of ASVs with warming treatment was lower compared with all those without warming treatment except warming & double precipitation & clipping (WDC) versus double precipitation & clipping (DC) for fungi and warming & clipping (WC) versus clipping (C) for protists in (a, c, e) (Paired t test, p < 0.0001). The number of samples did not have a substantial influence on the differences between warming and non-warming control as shown in (b, d, f). After removing global singletons and resampling, the rarefaction curves approached asymptotes for all treatments, indicating that the sequencing depth was sufficient for assessing the effects of various climate change factors on the diversity of these soil microbial communities.
Extended Data Fig. 5 Yearly differences of bacterial (a), fungal (b), and protistan (c) richness between warmed and unwarmed samples.
Data are presented as mean values ± SEM of the mean differences (warmed -unwarmed). For each year, the treatment effects are tested with linear mixed-effects models, and the significant treatment effects (p < 0.05, Wald type II χ² tests, n = 48 soils each year) are listed in the table. W: Warming; P: Precipitation level; C: Clipping.
Extended Data Fig. 6 Effects of experimental warming across microbial groups based on linear mixed-effects models.
a, Effect sizes of experimental warming on the (rescaled) phylogenetic diversity of major microbial groups based on linear mixed-effects models. b, Effect sizes of experimental warming on the (rescaled) relative abundance of major microbial groups based on linear mixed-effects models. Data are presented as mean values ± standard errors of the estimated effect sizes. Statistical significance is based on Wald type II χ² tests (n = 360 independent soil samples), which is denoted by asterisks: *** p < 0.001, ** p < 0.01, * p < 0.05.
Extended Data Fig. 7 Effects of experimental warming on sporulation families or genes of Firmicutes and Actinobacteria.
a, Number of Firmicutes families whose relative abundances increased, decreased or unchanged under warming. b, Number of Actinobacteria families whose relative abundances increased, decreased or unchanged under warming. Significant changes (p < 0.05) are based on Wald type II χ² tests (n = 360 independent soil samples) of the warming effects in linear mixed-effects models (relative abundance ~ warming × precipitation level × clipping + (1|Block) + (1 | year)). ‘Yspore’, ‘Nspore’, and ‘NAspore’ refer to known spore-formers, known non-spore-formers, and information not available on the ability to form spores, respectively. The spore-forming capability or different families within Firmicutes and Actinobacteria were mainly based on databases and literature. c, Effect sizes of experimental warming on the (rescaled) relative abundance of major sporulation genes in Firmicutes (spo0A gene) and Actinobacteria (bldD gene) based on linear mixed-effects models. The sporulation genes were retrieved from shotgun sequencing data. Data are presented as mean values ± standard errors of the estimated effect sizes. Statistical significance is based on Wald type II χ² tests (n = 64 independent soil metagenome samples).
Extended Data Fig. 8 Hypothesized conceptual models on the relationships between experimental treatments, environmental variables, and microbial diversity.
a, Bacteria and protists; b, Fungi. The environmental variables were selected based on their biological inference and collinearity, as detailed in the Methods and Supplementary Tables 3–7. We hypothesized that each experimental treatment would influence each environmental variable, and the environmental variables would all influence microbial diversity. We also assumed that microbial diversity would influence plant richness and biomass. In addition, we assumed interactions between bacteria and protists since there could be prey-predator relationships between them. In fact, the consumers, which include potential predators of bacteria, account for 84.6% of the total protist abundance. The richness of protistan consumers also highly correlated with the total protistan richness (Pearson’s r = 0.98). Protists was not included in the fungal model because the relative abundance of fungivorous protists is very low.
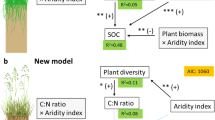
Extended Data Fig. 9 The structural equation model (SEM) showing the relationships among treatments, soil and plant variables, and fungal richness.
Blue and red arrows indicate positive and negative relationships, respectively. Solid or dashed lines indicate significant (p < 0.05) or nonsignificant relationships. Numbers near the pathway arrow indicate the standard path coefficients. R2 represents the proportion of variance explained for every dependent variable. χ2 = 28.70, df = 23, p = 0.19 (large p value indicates that the predicted model and observed data are equal, that is, good model fitting), CFI = 0.974. n = 48 biologically independent plots.
Extended Data Fig. 10 Effects of experimental warming on different ecosystem functions.
Data are presented as mean values ± standard errors of the estimated effect sizes. Statistical significance is based on Wald type II χ² tests (n = 360), which is indicated in the plot. GPP: gross primary productivity; ER: ecosystem respiration; NEE: net ecosystem exchange; Rh: heterotrophic respiration; Rs: soil total respiration.
Supplementary information
Supplementary Information
Supplementary Fig. 1, Tables 1–7 and Notes A–F.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
About this article
Cite this article
Wu, L., Zhang, Y., Guo, X. et al. Reduction of microbial diversity in grassland soil is driven by long-term climate warming. Nat Microbiol 7, 1054–1062 (2022). https://doi.org/10.1038/s41564-022-01147-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-022-01147-3
This article is cited by
-
Gut microbiota modulation enhances the immune capacity of lizards under climate warming
Microbiome (2024)
-
Microbially mediated mechanisms underlie soil carbon accrual by conservation agriculture under decade-long warming
Nature Communications (2024)
-
Relative importance of altitude shifts with plant and microbial diversity to soil multifunctionality in grasslands of north-western China
Plant and Soil (2024)
-
Experimental warming accelerates positive soil priming in a temperate grassland ecosystem
Nature Communications (2024)
-
Community composition drives siderophore dynamics in multispecies bacterial communities
BMC Ecology and Evolution (2023)