Abstract
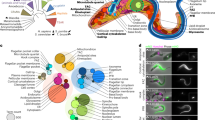
Protein kinases regulate fundamental aspects of eukaryotic cell biology, making them attractive chemotherapeutic targets in parasites like Plasmodium spp. and Toxoplasma gondii. To systematically examine the parasite kinome, we developed a high-throughput tagging (HiT) strategy to endogenously label protein kinases with an auxin-inducible degron and fluorophore. Hundreds of tagging vectors were assembled from synthetic sequences in a single reaction and used to generate pools of mutants to determine localization and function. Examining 1,160 arrayed clones, we assigned 40 protein localizations and associated 15 kinases with distinct defects. The fitness of tagged alleles was also measured by pooled screening, distinguishing delayed from acute phenotypes. A previously unstudied kinase, associated with a delayed phenotype, was shown to be a regulator of invasion and egress. We named the kinase Store Potentiating/Activating Regulatory Kinase (SPARK), based on its impact on intracellular Ca2+ stores. Despite homology to mammalian 3-phosphoinositide-dependent protein kinase-1 (PDK1), SPARK lacks a lipid-binding domain, suggesting a rewiring of the pathway in parasites. HiT screening extends genome-wide approaches into complex cellular phenotypes, providing a scalable and versatile platform to dissect parasite biology.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All oligos used in this study are available in Supplementary Table 1. All plasmids used or generated in this study are listed with their appropriate GenBank or PMID accession numbers in Supplementary Table 1. Minimally processed pooled and arrayed CRISPR screen sequencing results are available in Supplementary Table 2. Localization assignments, microscopy phenotypes, lytic assay results, and UMAP coordinates and clusters are likewise available in Supplementary Table 2. Source data are provided with this paper. Data from experimental results is available in the source data files. Additional unprocessed data is available from the corresponding author upon request.
Code availability
All code is described in the methods section and available from the corresponding author upon request.
References
Cabrera, D. G. et al. Plasmodial kinase inhibitors: License to cure? J. Med. Chem. 61, 8061–8077 (2018).
Sauvey, C., Ehrenkaufer, G., Shi, D., Debnath, A. & Abagyan, R. Antineoplastic kinase inhibitors: A new class of potent anti-amoebic compounds. PLoS Negl. Trop. Dis. 15, e0008425 (2021).
Merritt, C., Silva, L. E., Tanner, A. L., Stuart, K. & Pollastri, M. P. Kinases as druggable targets in trypanosomatid protozoan parasites. Chem. Rev. 114, 11280–11304 (2014).
Talevich, E., Mirza, A. & Kannan, N. Structural and evolutionary divergence of eukaryotic protein kinases in Apicomplexa. BMC Evol. Biol. 11, 321 (2011).
Peixoto, L. et al. Integrative genomic approaches highlight a family of parasite-specific kinases that regulate host responses. Cell Host Microbe 8, 208–218 (2010).
Gaji, R. Y., Sharp, A. K. & Brown, A. M. Protein kinases in Toxoplasma gondii. Int. J. Parasitol. 51, 415–429 (2021).
Beraki, T. et al. Divergent kinase regulates membrane ultrastructure of the Toxoplasma parasitophorous vacuole. Proc. Natl Acad. Sci. USA 116, 6361–6370 (2019).
Taylor, S. et al. A secreted serine-threonine kinase determines virulence in the eukaryotic pathogen Toxoplasma gondii. Science 314, 1776–1780 (2006).
Fox, B. A. et al. The Toxoplasma gondii rhoptry kinome is essential for chronic infection. mBio 7, e00193-16 (2016).
Fleckenstein, M. C. et al. A Toxoplasma gondii pseudokinase inhibits host IRG resistance proteins. PLoS Biol. 10, e1001358 (2012).
Niedelman, W. et al. The rhoptry proteins ROP18 and ROP5 mediate Toxoplasma gondii evasion of the murine, but not the human, interferon-gamma response. PLoS Pathog. 8, e1002784 (2012).
Sidik, S. M. et al. A genome-wide CRISPR screen in Toxoplasma identifies essential apicomplexan genes. Cell 166, 1423–1435.e12 (2016).
Bushell, E. et al. Functional profiling of a Plasmodium genome reveals an abundance of essential genes. Cell 170, 260–272.e8 (2017).
Zhang, M. et al. Uncovering the essential genes of the human malaria parasite Plasmodium falciparum by saturation mutagenesis. Science 360, eaap7847 (2018).
Tewari, R. et al. The systematic functional analysis of Plasmodium protein kinases identifies essential regulators of mosquito transmission. Cell Host Microbe 8, 377–387 (2010).
Meissner, M., Brecht, S., Bujard, H. & Soldati, D. Modulation of myosin A expression by a newly established tetracycline repressor-based inducible system in Toxoplasma gondii. Nucleic Acids Res. 29, E115 (2001).
Meissner, M., Schlüter, D. & Soldati, D. Role of Toxoplasma gondii myosin A in powering parasite gliding and host cell invasion. Science 298, 837–840 (2002).
van Poppel, N. F. J., Welagen, J., Duisters, R. F. J. J., Vermeulen, A. N. & Schaap, D. Tight control of transcription in Toxoplasma gondii using an alternative tet repressor. Int. J. Parasitol. 36, 443–452 (2006).
Pieperhoff, M. S. et al. Conditional U1 gene silencing in toxoplasma gondii. PLoS ONE 10, e0130356 (2015).
Andenmatten, N. et al. Conditional genome engineering in Toxoplasma gondii uncovers alternative invasion mechanisms. Nat. Methods 10, 125–127 (2013).
Hunt, A. et al. Differential requirements for cyclase-associated protein (CAP) in actin-dependent processes of Toxoplasma gondii. eLife 8, e50598 (2019).
Nishimura, K., Fukagawa, T., Takisawa, H., Kakimoto, T. & Kanemaki, M. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nat. Methods 6, 917–922 (2009).
Brown, K. M., Long, S. & Sibley, L. D. Plasma membrane association by N-acylation governs PKG function in Toxoplasma gondii. mBio 8, e00375-17 (2017).
Long, S. et al. Calmodulin-like proteins localized to the conoid regulate motility and cell invasion by Toxoplasma gondii. PLoS Pathog. 13, e1006379 (2017).
Huynh, M.-H. & Carruthers, V. B. Tagging of endogenous genes in a Toxoplasma gondii strain lacking Ku80. Eukaryot. Cell. 8, 530–539 (2009).
Fox, B. A., Ristuccia, J. G., Gigley, J. P. & Bzik, D. J. Efficient gene replacements in Toxoplasma gondii strains deficient for nonhomologous end joining. Eukaryot. Cell 8, 520–529 (2009).
Krogan, N. J. et al. Global landscape of protein complexes in the yeast Saccharomyces cerevisiae. Nature 440, 637–643 (2006).
Huh, W.-K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003).
Weill, U. et al. Genome-wide SWAp-Tag yeast libraries for proteome exploration. Nat. Methods 15, 617–622 (2018).
Leonetti, M. D., Sekine, S., Kamiyama, D., Weissman, J. S. & Huang, B. A scalable strategy for high-throughput GFP tagging of endogenous human proteins. Proc. Natl Acad. Sci. USA 113, E3501–E3508 (2016).
Cho, N. H. et al. OpenCell: Endogenous tagging for the cartography of human cellular organization. Science 6585, eabi6983 (2022).
Dean, S., Sunter, J. D. & Wheeler, R. J. TrypTag.org: A trypanosome genome-wide protein localisation resource. Trends Parasitol. 33, 80–82 (2017).
Sidik, S. M., Hackett, C. G., Tran, F., Westwood, N. J. & Lourido, S. Efficient genome engineering of Toxoplasma gondii using CRISPR/Cas9. PLoS ONE 9, e100450 (2014).
Sathyan, K. M. et al. An improved auxin-inducible degron system preserves native protein levels and enables rapid and specific protein depletion. Genes Dev. 33, 1441–1455 (2019).
Long, S., Anthony, B., Drewry, L. L. & Sibley, L. D. A conserved ankyrin repeat-containing protein regulates conoid stability, motility and cell invasion in Toxoplasma gondii. Nat. Commun. 8, 2236 (2017).
Lourido, S. et al. Calcium-dependent protein kinase 1 is an essential regulator of exocytosis in Toxoplasma. Nature 465, 359–362 (2010).
Lourido, S., Tang, K. & Sibley, L. D. Distinct signalling pathways control Toxoplasma egress and host-cell invasion. EMBO J. 31, 4524–4534 (2012).
McCoy, J. M., Whitehead, L., van Dooren, G. G. & Tonkin, C. J. TgCDPK3 regulates calcium-dependent egress of Toxoplasma gondii from host cells. PLoS Pathog. 8, e1003066 (2012).
Brown, K. M., Long, S. & Sibley, L. D. Conditional knockdown of proteins using auxin-inducible degron (AID) fusions in Toxoplasma gondii. Bio Protoc. 8, e2728 (2018).
Bullen, H. E. et al. Phosphatidic acid-mediated signaling regulates microneme secretion in Toxoplasma. Cell Host Microbe 19, 349–360 (2016).
Uboldi, A. D. et al. Protein kinase A negatively regulates Ca2+ signalling in Toxoplasma gondii. PLoS Biol. 16, e2005642 (2018).
Farrell, A. et al. A DOC2 protein identified by mutational profiling is essential for apicomplexan parasite exocytosis. Science 335, 218–221 (2012).
Jia, Y. et al. Crosstalk between PKA and PKG controls pH-dependent host cell egress of Toxoplasma gondii. EMBO J. 36, 3250–3267 (2017).
Harding, C. R. et al. Gliding associated proteins play essential roles during the formation of the inner membrane complex of Toxoplasma gondii. PLoS Pathog. 12, e1005403 (2016).
Barylyuk, K. et al. A comprehensive subcellular atlas of the Toxoplasma proteome via hyperlopit provides spatial context for protein functions. Cell Host Microbe 28, 752–766.e9 (2020).
Alvarez, C. A. & Suvorova, E. S. Checkpoints of apicomplexan cell division identified in Toxoplasma gondii. PLoS Pathog. 13, e1006483 (2017).
Varberg, J. M., Coppens, I., Arrizabalaga, G. & Gaji, R. Y. TgTKL1 is a unique plant-like nuclear kinase that plays an essential role in acute toxoplasmosis. mBio 9, e00301–e00318 (2018).
Silljé, H. H., Takahashi, K., Tanaka, K., Van Houwe, G. & Nigg, E. A. Mammalian homologues of the plant Tousled gene code for cell-cycle-regulated kinases with maximal activities linked to ongoing DNA replication. EMBO J. 18, 5691–5702 (1999).
Pilyugin, M., Demmers, J., Verrijzer, C. P., Karch, F. & Moshkin, Y. M. Phosphorylation-mediated control of histone chaperone ASF1 levels by Tousled-like kinases. PLoS ONE 4, e8328 (2009).
Silljé, H. H. & Nigg, E. A. Identification of human Asf1 chromatin assembly factors as substrates of Tousled-like kinases. Curr. Biol. 11, 1068–1073 (2001).
Eckert, D. et al. Prp4 kinase grants the license to splice: Control of weak splice sites during spliceosome activation. PLoS Genet. 12, e1005768 (2016).
Naumov, A. et al. The Toxoplasma centrocone houses cell cycle regulatory factors. mBio 8, e00579-17 (2017).
Gubbels, M.-J. et al. Forward genetic analysis of the apicomplexan cell division cycle in Toxoplasma gondii. PLoS Pathog. 4, e36 (2008).
Chen, C.-T. & Gubbels, M.-J. The Toxoplasma gondii centrosome is the platform for internal daughter budding as revealed by a Nek1 kinase mutant. J. Cell Sci. 126, 3344–3355 (2013).
Suvorova, E. S., Francia, M., Striepen, B. & White, M. W. A novel bipartite centrosome coordinates the apicomplexan cell cycle. PLoS Biol. 13, e1002093 (2015).
Hu, X., O’Shaughnessy, W. J., Beraki, T. G. & Reese, M. L. Loss of the conserved alveolate kinase MAPK2 decouples Toxoplasma cell growth from cell division. mBio 11, e02517-20 (2020).
Martín Moyano, P., Němec, V. & Paruch, K. Cdc-Like Kinases (CLKs): Biology, chemical probes, and therapeutic potential. Int. J. Mol. Sci. 21, 7549 (2020).
Berto, G., Ferreira-Cerca, S. & De Wulf, P. The Rio1 protein kinases/ATPases: conserved regulators of growth, division, and genomic stability. Curr. Genet. 65, 457–466 (2019).
Mallari, J. P., Oksman, A., Vaupel, B. & Goldberg, D. E. Kinase-associated endopeptidase 1 (Kae1) participates in an atypical ribosome-associated complex in the apicoplast of Plasmodium falciparum. J. Biol. Chem. 289, 30025–30039 (2014).
Back, P. S. et al. Ancient MAPK ERK7 is regulated by an unusual inhibitory scaffold required for Toxoplasma apical complex biogenesis. Proc. Natl Acad. Sci. USA 117, 12164–12173 (2020).
O’Shaughnessy, W. J., Hu, X., Beraki, T., McDougal, M. & Reese, M. L. Loss of a conserved MAPK causes catastrophic failure in assembly of a specialized cilium-like structure in Toxoplasma gondii. Mol. Biol. Cell 31, 881–888 (2020).
Sampels, V. et al. Conditional mutagenesis of a novel choline kinase demonstrates plasticity of phosphatidylcholine biogenesis and gene expression in Toxoplasma gondii. J. Biol. Chem. 287, 16289–16299 (2012).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR–Cas9. Nat. Biotechnol. 34, 184–191 (2016).
McInnes, L., Healy, J. & Melville, J. UMAP: Uniform manifold approximation and projection for dimension reduction. Preprint at https://arxiv.org/abs/1802.03426 (2018).
Hartigan, J. A. & Wong, M. A. Algorithm AS 136: A K-means clustering algorithm. J. R. Stat. Soc. Ser. C Appl. Stat. 28, 100–108 (1979).
Donald, R. G. K. et al. Toxoplasma gondii cyclic GMP-dependent kinase: chemotherapeutic targeting of an essential parasite protein kinase. Eukaryot. Cell 1, 317–328 (2002).
Katso, R. et al. Cellular function of phosphoinositide 3-kinases: implications for development, homeostasis, and cancer. Annu. Rev. Cell Dev. Biol. 17, 615–675 (2001).
Balla, A. & Balla, T. Phosphatidylinositol 4-kinases: old enzymes with emerging functions. Trends Cell Biol. 16, 351–361 (2006).
Dvorin, J. D. et al. A plant-like kinase in Plasmodium falciparum regulates parasite egress from erythrocytes. Science 328, 910–912 (2010).
Kato, N. et al. Gene expression signatures and small-molecule compounds link a protein kinase to Plasmodium falciparum motility. Nat. Chem. Biol. 4, 347–356 (2008).
Bansal, A. et al. Characterization of Plasmodium falciparum calcium-dependent protein kinase 1 (PfCDPK1) and its role in microneme secretion during erythrocyte invasion. J. Biol. Chem. 288, 1590–1602 (2013).
Green, J. L. et al. The motor complex of Plasmodium falciparum: phosphorylation by a calcium-dependent protein kinase. J. Biol. Chem. 283, 30980–30989 (2008).
Chen, F., Mackey, A. J., Stoeckert, C. J. Jr & Roos, D. S. OrthoMCL-DB: querying a comprehensive multi-species collection of ortholog groups. Nucleic Acids Res. 34, D363–D368 (2006).
Rosenberg, A., Luth, M. R., Winzeler, E. A., Behnke, M. & Sibley, L. D. Evolution of resistance in vitro reveals mechanisms of artemisinin activity in Toxoplasma gondii. Proc. Natl Acad. Sci. USA 116, 26881–26891 (2019).
Hitz, E. et al. The 3-phosphoinositide-dependent protein kinase 1 is an essential upstream activator of protein kinase A in malaria parasites. PLoS Biol. 19, e3001483 (2021).
Dittrich, A. C. N. & Devarenne, T. P. Perspectives in PDK1 evolution: insights from photosynthetic and non-photosynthetic organisms. Plant Signal. Behav. 7, 642–649 (2012).
Sidik, S. M. et al. Using a genetically encoded sensor to identify inhibitors of Toxoplasma gondii Ca2+ signaling. J. Biol. Chem. 291, 9566–9580 (2016).
Howard, B. L. et al. Identification of potent phosphodiesterase inhibitors that demonstrate cyclic nucleotide-dependent functions in apicomplexan parasites. ACS Chem. Biol. 10, 1145–1154 (2015).
Wiersma, H. I. et al. A role for coccidian cGMP-dependent protein kinase in motility and invasion. Int. J. Parasitol. 34, 369–380 (2004).
Brown, K. M., Lourido, S. & Sibley, L. D. Serum albumin stimulates protein kinase G-dependent microneme secretion in Toxoplasma gondii. J. Biol. Chem. 291, 9554–9565 (2016).
McRobert, L. et al. Gametogenesis in malaria parasites is mediated by the cGMP-dependent protein kinase. PLoS Biol. 6, e139 (2008).
Collins, C. R. et al. Malaria parasite cGMP-dependent protein kinase regulates blood stage merozoite secretory organelle discharge and egress. PLoS Pathog. 9, e1003344 (2013).
Yang, L. et al. An apically located hybrid guanylate cyclase-ATPase is critical for the initiation of Ca2+ signaling and motility in Toxoplasma gondii. J. Biol. Chem. 294, 8959–8972 (2019).
Bisio, H., Lunghi, M., Brochet, M. & Soldati-Favre, D. Phosphatidic acid governs natural egress in Toxoplasma gondii via a guanylate cyclase receptor platform. Nat. Microbiol. 4, 420–428 (2019).
Brown, K. M. & Sibley, L. D. Essential cGMP signaling in Toxoplasma is initiated by a hybrid P-Type ATPase-guanylate cyclase. Cell Host Microbe 24, 804–816.e6 (2018).
Chen, T.-W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Pace, D. A., McKnight, C. A., Liu, J., Jimenez, V. & Moreno, S. N. J. Calcium entry in Toxoplasma gondii and its enhancing effect of invasion-linked traits. J. Biol. Chem. 289, 19637–19647 (2014).
Arrizabalaga, G. & Boothroyd, J. C. Role of calcium during Toxoplasma gondii invasion and egress. Int. J. Parasitol. 34, 361–368 (2004).
Alsford, S. et al. High-throughput phenotyping using parallel sequencing of RNA interference targets in the African trypanosome. Genome Res. 21, 915–924 (2011).
Young, J. et al. A CRISPR platform for targeted in vivo screens identifies Toxoplasma gondii virulence factors in mice. Nat. Commun. 10, 3963 (2019).
Tosetti, N., Dos Santos Pacheco, N., Soldati-Favre, D. & Jacot, D. Three F-actin assembly centers regulate organelle inheritance, cell–cell communication and motility in Toxoplasma gondii. eLife 8, e42669 (2019).
Brochet, M. et al. Phosphoinositide metabolism links cGMP-dependent protein kinase G to essential Ca2+ signals at key decision points in the life cycle of malaria parasites. PLoS Biol. 12, e1001806 (2014).
Carruthers, V. B., Moreno, S. N. & Sibley, L. D. Ethanol and acetaldehyde elevate intracellular [Ca2+] and stimulate microneme discharge in Toxoplasma gondii. Biochem. J. 342, 379–386 (1999).
Lovett, J. L., Marchesini, N., Moreno, S. N. J. & Sibley, L. D. Toxoplasma gondii microneme secretion involves intracellular Ca(2+) release from inositol 1,4,5-triphosphate (IP(3))/ryanodine-sensitive stores. J. Biol. Chem. 277, 25870–25876 (2002).
Bullen, H. E., Bisio, H. & Soldati-Favre, D. The triumvirate of signaling molecules controlling Toxoplasma microneme exocytosis: Cyclic GMP, calcium, and phosphatidic acid. PLoS Pathog. 15, e1007670 (2019).
Leroux, A. E., Schulze, J. O. & Biondi, R. M. AGC kinases, mechanisms of regulation and innovative drug development. Semin. Cancer Biol. 48, 1–17 (2018).
Mora, A., Komander, D., van Aalten, D. M. F. & Alessi, D. R. PDK1, the master regulator of AGC kinase signal transduction. Semin. Cell Dev. Biol. 15, 161–170 (2004).
Li, W. et al. A splitCas9 phenotypic screen in Toxoplasma gondii identifies proteins involved in host cell egress and invasion. Nat. Microbiol. https://doi.org/10.1038/s41564-022-01114-y (2022).
Markus, B. M., Bell, G. W., Lorenzi, H. A. & Lourido, S. Optimizing systems for Cas9 expression in Toxoplasma gondii. mSphere 4, e00386-19 (2019).
Paquet, D. et al. Efficient introduction of specific homozygous and heterozygous mutations using CRISPR/Cas9. Nature 533, 125–129 (2016).
Dewari, P. S. et al. An efficient and scalable pipeline for epitope tagging in mammalian stem cells using Cas9 ribonucleoprotein. eLife 7, e35069 (2018).
Bialk, P., Rivera-Torres, N., Strouse, B. & Kmiec, E. B. Regulation of gene editing activity directed by single-stranded oligonucleotides and CRISPR/Cas9 Systems. PLoS ONE 10, e0129308 (2015).
Liang, X., Potter, J., Kumar, S., Ravinder, N. & Chesnut, J. D. Enhanced CRISPR/Cas9-mediated precise genome editing by improved design and delivery of gRNA, Cas9 nuclease, and donor DNA. J. Biotechnol. 241, 136–146 (2017).
Burg, J. L., Perelman, D., Kasper, L. H., Ware, P. L. & Boothroyd, J. C. Molecular analysis of the gene encoding the major surface antigen of Toxoplasma gondii. J. Immunol. 141, 3584–3591 (1988).
Waldman, B. S. et al. Identification of a master regulator of differentiation in toxoplasma. Cell 180, 359–372.e16 (2020).
Starnes, G. L., Jewett, T. J., Carruthers, V. B. & Sibley, L. D. Two separate, conserved acidic amino acid domains within the Toxoplasma gondii MIC2 cytoplasmic tail are required for parasite survival. J. Biol. Chem. 281, 30745–30754 (2006).
Plattner, F. et al. Toxoplasma profilin is essential for host cell invasion and TLR11-dependent induction of an interleukin-12 response. Cell Host Microbe 3, 77–87 (2008).
Shortt, E. & Lourido, S. Plate-based quantification of stimulated Toxoplasma egress. Methods Mol. Biol. 2071, 171–186 (2020).
Acknowledgements
We thank the Whitehead Institute Bioinformatics and Research Computing Core, especially B. Yuan, for assistance implementing gRNA design pipelines; L.D. Sibley for the TIR1 strain; M. Treeck for the DiCre strain; W. Salmon and the W.M. Keck Biological Imaging Facility for confocal microscopy support; P. W. Reddien for use of the Illumina MiSeq; B.S. Waldman, E.A. Boydston, C.J. Giuliano, A.W. Chan, S. Sidik and B.M. Markus for technical support in generation of the array; VEuPathDB and all contributors to this resource. This work was supported by funds from a National Institutes of Health grant (R01AI144369) to S.L. and National Science Foundation Graduate Research Fellowships to T.A.S. (2018259980) and A.L.H. (174530).
Author information
Authors and Affiliations
Contributions
T.A.S. and S.L. designed the overall study and experiments. HiT vectors were designed, constructed and tested for their tagging efficiency by T.A.S. and G.S.L.-P. The TIR1/GCaMP6f parasite strain was constructed and validated by A.L.H. and the scarlessly tagged CDPK1 and CDPK3 parasite strains were constructed and validated by E.S. T.A.S. performed all remaining parasite strain construction and experiments. T.A.S. and S.L. wrote the manuscript and all authors reviewed, offered input and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks David Horn and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Transfected populations efficiently incorporate a variety of HiT vectors.
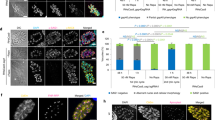
a, Fluorescence microscopy of the tagged populations displaying the correct localization of each kinase and expression levels consistent with flow cytometry (Fig. 1b). b, Live microscopy of V5-T2A-mKate2 HiT-tagged population (merged image in Fig. 1g). c, Immunofluorescence microscopy of population tagged with the HA-U1 HiT vector following treatment with rapamycin or vehicle control (merged image in Fig. 1i). d, Flow cytometry of parasite populations tagged with the V5-mNG-mAID HiT vector targeting CDPK1 or CDPK3 and treated with either IAA or vehicle control for 24 h (excerpt shown in Fig. 1l).
Extended Data Fig. 2 Arrayed screening results.
a, Results from dual-indexed sequencing of the arrayed clones. A minimum of 100 reads were required to assign a given gRNA to a particular clone. Cases where a second gRNA reached >10% the abundance of the first gRNA were classified as containing multiple integrations. b, Histogram showing the distribution of gRNAs and genes contained among single-integrated wells within the array. Genes and gRNAs with no representation are omitted from the plot.
Extended Data Fig. 3 Representative images from the arrayed screen.
a–f, Widefield microscopy of representative clones. Maximum intensity projections for IMC1-tdTomato and mNeonGreen-tagged targets are displayed for cultures treated with either IAA or vehicle for 24 hours. All images are displayed at the same scale. Localizations to the nucleus (a), daughter cell IMC (b), parasitophorous vacuole (c), perinuclear space (d), cytosol (e) or apical end (f) were assigned to a gene if half or more of single-integrated wells for that gene displayed consistent localizations.
Extended Data Fig. 4 Additional representative images from the arrayed screen and comparisons to the pooled results.
a–c, Widefield microscopy of representative clones. Maximum intensity projections for IMC1-tdTomato and mNeonGreen-tagged targets are displayed for cultures treated with either IAA or vehicle for 24 hours. All images are displayed at the same scale. Localizations to puncta (a), the basal end (b), or peripheral structures (c) were assigned to a gene if half or more of single-integrated wells for that gene displayed consistent localizations. d, Representative confocal images of a sample of clones. mNeonGreen (green); IMC1-tdTomato (magenta). Images are maximum intensity projections. Genes are numbered based on the unique identifier from ToxoDB (for example, TGGT1_210830, labeled 210830). e, Comparison of relative gRNA abundances in the array compared to the pooled population that was subcloned. Spearman correlation coefficient = 0.77. f, Impact of the initial lytic cycles on gRNA abundance for genes with delayed or acute loss phenotypes in the HiT screen. The effect of the first lytic cycle from the HiT screen is plotted against the effect of the first or second lytic cycles for the genome-wide knockout screen (Sidik & Huet, et al. 2016). Genes are paired across their first and second lytic cycles within the genome-wide knockout screen.
Extended Data Fig. 5 Extended analysis of SPARK depletion.
a, Replication assay of SPARK-AID parasites. Parasites were treated with either IAA or vehicle at 3 hours post-invasion and imaged 24 hours later. The number of parasites per vacuole were counted for 100 vacuoles per sample. Mean ± S.E. graphed for n = 3 biological replicates. b, Extracellular parasites in basal Ca2+ buffer stimulated with vehicle or the Ca2+ ionophore ionomycin, following 24 h of treatment with vehicle or IAA. Cytosolic Ca2+ flux was measured in bulk as GCaMP6f fluorescence normalized to the initial and maximum fluorescence following aerolysin permeabilization in 2 mM Ca2+. Mean ± S.E. graphed for n = 3–6 biological replicates.
Supplementary information
Supplementary Information
Supplementary Figs. 1–3.
Supplementary Table 1
Oligos and plasmids used in this study.
Supplementary Table 2
Combined results from the HiT screens summarizing data from arrayed and pooled analyses.
Source data
Source Data Fig. 4
b, Competition assays of delayed-loss candidates. Provided are relative abundances of each strain relative to the WT competitor strain. Values are normalized to the starting ratio. d, Area sizes of individual plaques. Provided values are in mm2. e, Invasion efficiencies of delayed-loss candidates. Invaded parasites per nuclei for each replicate are provided, in addition to the values post-normalization to the WT vehicle-treated sample.
Source Data Fig. 5
a, Kinase domain sequences used to generate alignments and the subsequent phylogenetic tree. e,f, Egress efficiencies following either (e) zaprinast or (f) A23187 stimulation. Provided is per cent of egress relative to the final percentage of egress of the vehicle-treated sample. h, Quantification of GCaMP6f fluorescence signal following either zaprinast or A23187 treatment. Average fluorescence of each vacuole was quantified relative to initial fluorescence until egress of the vacuole or until the end of the time-course. i, Quantification of GCaMP6f fluorescence from extracellular parasites treated with either zaprinast or A23187. Provided are background-subtracted values normalized to initial fluorescence and the final maximum fluorescence following aerolysin and Ca2+ treatment.
Source Data Extended Data Fig. 5
a, Replication assays of SPARK grown in the presence or absence of IAA. 100 vacuoles were quantified for each sample and condition. Provided are the number of occurrences of each vacuole size for each replicate. b, Quantification of GCaMP6f fluorescence from extracellular parasites treated with either vehicle (DMSO) or ionomycin. Provided are background-subtracted values normalized to initial fluorescence and the final maximum fluorescence following aerolysin and Ca2+ treatment.
Rights and permissions
About this article
Cite this article
Smith, T.A., Lopez-Perez, G.S., Herneisen, A.L. et al. Screening the Toxoplasma kinome with high-throughput tagging identifies a regulator of invasion and egress. Nat Microbiol 7, 868–881 (2022). https://doi.org/10.1038/s41564-022-01104-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-022-01104-0
This article is cited by
-
A positive feedback loop controls Toxoplasma chronic differentiation
Nature Microbiology (2023)