Abstract
The emergence of multidrug-resistant bacteria poses a threat to global health and necessitates the development of additional in vivo active antibiotics with diverse modes of action. Directly targeting menaquinone (MK), which plays an important role in bacterial electron transport, is an appealing, yet underexplored, mode of action due to a dearth of MK-binding molecules. Here we combine sequence-based metagenomic mining with a motif search of bioinformatically predicted natural product structures to identify six biosynthetic gene clusters that we predicted encode MK-binding antibiotics (MBAs). Their predicted products (MBA1–6) were rapidly accessed using a synthetic bioinformatic natural product approach, which relies on bioinformatic structure prediction followed by chemical synthesis. Among these six structurally diverse MBAs, four make up two new MBA structural families. The most potent member of each new family (MBA3, MBA6) proved effective at treating methicillin-resistant Staphylococcus aureus infection in a murine peritonitis-sepsis model. The only conserved feature present in all MBAs is the sequence ‘GXLXXXW’, which we propose represents a minimum MK-binding motif. Notably, we found that a subset of MBAs were active against Mycobacterium tuberculosis both in vitro and in macrophages. Our findings suggest that naturally occurring MBAs are a structurally diverse and untapped class of mechanistically interesting, in vivo active antibiotics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The biosynthetic gene clusters for MBA1–6 have been deposited in GenBank under accession nos. MZ146900 and MZ146905, respectively. All other data are available in the main text or as supplementary information. The following five publicly available resources were used in this study: antiSMASH (https://antismash-db.secondarymetabolites.org/); MIBiG (https://mibig.secondarymetabolites.org/); Norine (https://bioinfo.lifl.fr/norine/); SANDPUMA (https://bitbucket.org/chevrm/sandpuma); and PubChem databases (https://pubchem.ncbi.nlm.nih.gov/). Source data are provided with this paper.
References
Tackling Drug-resistant Infections Globally: Final Report and Recommendations. The Review on Antimicrobial Resistance (Wellcome Trust and UK Government, 2016).
de Kraker, M. E. A., Stewardson, A. J. & Harbarth, S. Will 10 million people die a year due to antimicrobial resistance by 2050? PLoS Med. 13, e1002184 (2016).
Brown, E. D. & Wright, G. D. Antibacterial drug discovery in the resistance era. Nature 529, 336–343 (2016).
Niu, G. & Li, W. Next-generation drug discovery to combat antimicrobial resistance. Trends Biochem. Sci. 44, 961–972 (2019).
Lewis, K. The science of antibiotic discovery. Cell 181, 29–45 (2020).
Johnston, J. M. & Bulloch, E. M. Advances in menaquinone biosynthesis: sublocalisation and allosteric regulation. Curr. Opin. Struct. Biol. 65, 33–41 (2020).
Boersch, M., Rudrawar, S., Grant, G. & Zunk, M. Menaquinone biosynthesis inhibition: a review of advancements toward a new antibiotic mechanism. RSC Adv. 8, 5099–5105 (2018).
Paudel, A., Hamamoto, H., Panthee, S. & Sekimizu, K. Menaquinone as a potential target of antibacterial agents. Drug Discov. Ther. 10, 123–128 (2016).
Le, P. et al. Repurposing human kinase inhibitors to create an antibiotic active against drug-resistant Staphylococcus aureus, persisters and biofilms. Nat. Chem. 12, 145–158 (2020).
Hamamoto, H. et al. Lysocin E is a new antibiotic that targets menaquinone in the bacterial membrane. Nat. Chem. Biol. 11, 127–133 (2015).
Itoh, H. et al. Total synthesis and biological mode of action of WAP-8294A2: a menaquinone-targeting antibiotic. J. Org. Chem. 83, 6924–6935 (2018).
Sang, M. et al. Identification of an anti-MRSA cyclic lipodepsipeptide, WBP-29479A1, by genome mining of Lysobacter antibioticus. Org. Lett. 21, 6432–6436 (2019).
Santiago, M. et al. Genome-wide mutant profiling predicts the mechanism of a Lipid II binding antibiotic. Nat. Chem. Biol. 14, 601–608 (2018).
Butler, M. S., Blaskovich, M. A. & Cooper, M. A. Antibiotics in the clinical pipeline in 2013. J. Antibiot. (Tokyo) 66, 571–591 (2013).
Rutledge, P. J. & Challis, G. L. Discovery of microbial natural products by activation of silent biosynthetic gene clusters. Nat. Rev. Microbiol. 13, 509–523 (2015).
Crits-Christoph, A., Diamond, S., Butterfield, C. N., Thomas, B. C. & Banfield, J. F. Novel soil bacteria possess diverse genes for secondary metabolite biosynthesis. Nature 558, 440–444 (2018).
Libis, V. et al. Uncovering the biosynthetic potential of rare metagenomic DNA using co-occurrence network analysis of targeted sequences. Nat. Commun. 10, 3848 (2019).
Kalkreuter, E., Pan, G., Cepeda, A. J. & Shen, B. Targeting bacterial genomes for natural product discovery. Trends Pharmacol. Sci. 41, 13–26 (2020).
Nayfach, S. et al. A genomic catalog of Earth’s microbiomes. Nat. Biotechnol. 39, 499–509 (2021).
Chu, J. et al. Discovery of MRSA active antibiotics using primary sequence from the human microbiome. Nat. Chem. Biol. 12, 1004–1006 (2016).
Chu, J., Vila-Farres, X. & Brady, S. F. Bioactive synthetic-bioinformatic natural product cyclic peptides inspired by nonribosomal peptide synthetase gene clusters from the human microbiome. J. Am. Chem. Soc. 141, 15737–15741 (2019).
Hover, B. M. et al. Culture-independent discovery of the malacidins as calcium-dependent antibiotics with activity against multidrug-resistant Gram-positive pathogens. Nat. Microbiol. 3, 415–422 (2018).
Peek, J. et al. Rifamycin congeners kanglemycins are active against rifampicin-resistant bacteria via a distinct mechanism. Nat. Commun. 9, 4147 (2018).
Charlop-Powers, Z. et al. Global biogeographic sampling of bacterial secondary metabolism. eLife 4, e05048 (2015).
Lemetre, C. et al. Bacterial natural product biosynthetic domain composition in soil correlates with changes in latitude on a continent-wide scale. Proc. Natl Acad. Sci. USA 114, 11615–11620 (2017).
Reddy, B. V., Milshteyn, A., Charlop-Powers, Z. & Brady, S. F. eSNaPD: a versatile, web-based bioinformatics platform for surveying and mining natural product biosynthetic diversity from metagenomes. Chem. Biol. 21, 1023–1033 (2014).
Stachelhaus, T., Mootz, H. D. & Marahiel, M. A. The specificity-conferring code of adenylation domains in nonribosomal peptide synthetases. Chem. Biol. 6, 493–505 (1999).
Blin, K. et al. antiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res. 47, W81–W87 (2019).
Chen, H. T., Olson, A. S., Su, W., Dussault, P. H. & Du, L. Fatty acyl incorporation in the biosynthesis of WAP-8294A, a group of potent anti-MRSA cyclic lipodepsipeptides. RCS Adv. 5, 105753–105759 (2015).
Zhang, W. et al. Identification and characterization of the anti-methicillin-resistant Staphylococcus aureus WAP-8294A2 biosynthetic gene cluster from Lysobacter enzymogenes OH11. Antimicrob. Agents Chemother. 55, 5581–5589 (2011).
Chen, D., Tian, L., Po, K. H. L., Chen, S. & Li, X. Total synthesis and a systematic structure-activity relationship study of WAP-8294A2. Bioorg. Med. Chem. 28, 115677 (2020).
Bionda, N., Pitteloud, J.-P. & Cudic, P. Cyclic lipodepsipeptides: a new class of antibacterial agents in the battle against resistant bacteria. Future Med. Chem. 5, 1311–1330 (2013).
Kato, A. et al. A new anti-MRSA antibiotic complex, WAP-8294A. I. Taxonomy, isolation and biological activities. J. Antibiot. (Tokyo) 51, 929–935 (1998).
Yagi, A. et al. Anti-Mycobacterium activity of microbial peptides in a silkworm infection model with Mycobacterium smegmatis. J. Antibiot. (Tokyo) 70, 685–690 (2017).
Collins, M. D. & Jones, D. The distribution of isoprenoid quinones in streptococci of serological groups D and N. J. Gen. Microbiol. 114, 27–33 (1979).
Huycke, M. M. et al. Extracellular superoxide production by Enterococcus faecalis requires demethylmenaquinone and is attenuated by functional terminal quinol oxidases. Mol. Microbiol. 42, 729–740 (2001).
Müller, A., Klöckner, A. & Schneider, T. Targeting a cell wall biosynthesis hot spot. Nat. Prod. Rep. 34, 909–932 (2017).
Malin, J. J. & de Leeuw, E. Therapeutic compounds targeting Lipid II for antibacterial purposes. Infect. Drug Resist. 12, 2613–2625 (2019).
Sampson, B. A., Misra, R. & Benson, S. A. Identification and characterization of a new gene of Escherichia coli K-12 involved in outer membrane permeability. Genetics 122, 491–501 (1989).
Meganathan, R. Biosynthesis of menaquinone (vitamin K2) and ubiquinone (coenzyme Q): a perspective on enzymatic mechanisms. Vitam. Horm. 61, 173–218 (2001).
Meganathan, R. & Kwon, O. Biosynthesis of menaquinone (vitamin K2) and ubiquinone (coenzyme Q). EcoSal Plus https://doi.org/10.1128/ecosalplus.3.6.3.3 (2009).
Wakeman, C. A. et al. Menaquinone biosynthesis potentiates haem toxicity in Staphylococcus aureus. Mol. Microbiol. 86, 1376–1392 (2012).
Saravanan, M. et al. Review on emergence of drug-resistant tuberculosis (MDR & XDR-TB) and its molecular diagnosis in Ethiopia. Microb. Pathog. 117, 237–242 (2018).
Global Tuberculosis Report 2019 (World Health Organization, 2019).
Libardo, M. D. J., Boshoff, H. I. & Barry, C. E.3rd The present state of the tuberculosis drug development pipeline. Curr. Opin. Pharmacol. 42, 81–94 (2018).
Wellington, S. & Hung, D. T. The expanding diversity of Mycobacterium tuberculosis drug targets. ACS Infect. Dis. 4, 696–714 (2018).
Berube, B. J. et al. Novel MenA inhibitors are bactericidal against Mycobacterium tuberculosis and synergize with electron transport chain inhibitors. Antimicrob. Agents Chemother. 63, e02661-18 (2019).
Pieters, J. Mycobacterium tuberculosis and the macrophage: maintaining a balance. Cell Host Microbe 3, 399–407 (2008).
Rastogi, N., Labrousse, V. & Goh, K. S. In vitro activities of fourteen antimicrobial agents against drug susceptible and resistant clinical isolates of Mycobacterium tuberculosis and comparative intracellular activities against the virulent H37Rv strain in human macrophages. Curr. Microbiol. 33, 167–175 (1996).
Laufer, B., Chatterjee, J., Frank, A. O. & Kessler, H. Can N-methylated amino acids serve as substitutes for prolines in conformational design of cyclic pentapeptides? J. Pept. Sci. 15, 141–146 (2009).
Stark, M., Liu, L.-P. & Deber, C. M. Cationic hydrophobic peptides with antimicrobial activity. Antimicrob. Agents Chemother. 46, 3585–3590 (2002).
Kaji, T. et al. Total synthesis and functional evaluation of fourteen derivatives of lysocin E: importance of cationic, hydrophobic, and aromatic moieties for antibacterial activity. Chemistry 22, 16912–16919 (2016).
Kaupp, M. The function of photosystem I. Quantum chemical insight into the role of tryptophan–quinone interactions. Biochemistry 41, 2895–2900 (2002).
Yu, L. et al. Identification of the biosynthetic gene cluster for the anti-MRSA lysocins through gene cluster activation using strong promoters of housekeeping genes and production of new analogs in Lysobacter sp. 3655. ACS Synth. Biol. 9, 1989–1997 (2020).
Masschelein, J., Jenner, M. & Challis, G. L. Antibiotics from Gram-negative bacteria: a comprehensive overview and selected biosynthetic highlights. Nat. Prod. Rep. 34, 712–783 (2017).
Liu, Y., Ding, S., Shen, J. & Zhu, K. Nonribosomal antibacterial peptides that target multidrug-resistant bacteria. Nat. Prod. Rep. 36, 573–592 (2019).
Urakami, T., Tamaoka, J., Suzuki, K.-I. & Komagata, K. Paracoccus alcaliphilus sp. nov., an alkaliphilic and facultatively methylotrophic bacterium. Int. J. Syst. Bacteriol. 39, 116–121 (1989).
Xie, C.-H. & Yokota, A. Dyella japonica gen. nov., sp. nov., a γ-proteobacterium isolated from soil. Int. J. Syst. Evol. Microbiol. 55, 753–756 (2005).
Proctor, R. A. et al. Small colony variants: a pathogenic form of bacteria that facilitates persistent and recurrent infections. Nat. Rev. Microbiol. 4, 295–305 (2006).
Lannergård, J. et al. Identification of the genetic basis for clinical menadione-auxotrophic small-colony variant isolates of Staphylococcus aureus. Antimicrob. Agents Chemother. 52, 4017–4022 (2008).
Cao, S., Huseby, D. L., Brandis, G. & Hughes, D. Alternative evolutionary pathways for drug-resistant small colony variant mutants in Staphylococcus aureus. mBio 8, e00358-17 (2017).
Blin, K., Shaw, S., Kautsar, S. A., Medema, M. H. & Weber, T. The antiSMASH database version 3: increased taxonomic coverage and new query features for modular enzymes. Nucleic Acids Res. 49, D639–D643 (2021).
Flissi, A. et al. Norine: update of the nonribosomal peptide resource. Nucleic Acids Res. 48, D465–D469 (2020).
Kautsar, S. A. et al. MIBiG 2.0: a repository for biosynthetic gene clusters of known function. Nucleic Acids Res. 48, D454–D458 (2020).
Chevrette, M. G., Aicheler, F., Kohlbacher, O., Currie, C. R. & Medema, M. H. SANDPUMA: ensemble predictions of nonribosomal peptide chemistry reveal biosynthetic diversity across Actinobacteria. Bioinformatics 33, 3202–3210 (2017).
Samdani, A. & Vetrivel, U. POAP: a GNU parallel based multithreaded pipeline of open babel and AutoDock suite for boosted high throughput virtual screening. Comput. Biol. Chem. 74, 39–48 (2018).
Kim, S. et al. PubChem in 2021: new data content and improved web interfaces. Nucleic Acids Res. 49, D1388–D1395 (2021).
Röttig, M. et al. NRPSpredictor2––a web server for predicting NRPS adenylation domain specificity. Nucleic Acids Res. 39, W362–W367 (2011).
Xie, F, Pei, S., Lin, X., Tian, Y. & Zhang, G. A rapid and efficient method for the extraction and identification of menaquinones from Actinomycetes in wet biomass. BMC Microbiol. 21, 175 (2021).
Acknowledgements
We thank the High Throughput Screening and Resource Center at The Rockefeller University for their assistance with the ITC experiments. We thank J. Rock at The Rockefeller University for M. smegmatis mc2 155. We thank W. Jacobs Jr at Albert Einstein College of Medicine for M. tuberculosis mc2 6206, mc2 7901 and mc2 6206 with the mLux plasmid. We thank V. Fischetti at The Rockefeller University for S. aureus Newman, D712 and O315. We thank Y. Q. Xiong at the David Geffen School of Medicine at University of California Los Angeles for S. aureus A215 and SA684. We thank E. Skaar at Vanderbilt University Medical Center for S. aureus ΔmenA and ΔmenB. We thank J. Peek and J. Burian at The Rockefeller University for careful proofreading. This work was supported by the National Institutes of Health (grant nos. 1U19AI142731 and 5R35GM122559 to S.F.B.).
Author information
Authors and Affiliations
Contributions
L.L. designed the overall experimental plan for the manuscript, performed most of the experiments presented and wrote the manuscript. B.K. performed the peptide synthesis and assisted with the mouse experiments. Y.H. performed the bioinformatics analysis. L.W.M. contributed to the NMR data analysis. M.A.T. performed the MiSeq sequencing. R.R. performed the antibacterial assay against the WT and MDR M. tuberculosis strains. S.F.B. contributed to project management, designed the overall experimental plan for the manuscript and wrote the manuscript. All authors edited the manuscript and approved the final submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Microbiology thanks Michael R. Barbachyn, Stephan A. Sieber and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Phylogenetic analysis of eSNaPD hits from six conserved A-domains found in the BGCs of the three known MBAs.
Phylogenetic analysis of eSNaPD hits (a) and predicted peptide sequences of recovered clones (b). All the hits at an e-value ≤10–45 from A-domain analysis of l-Leu-6 encoded new MBAs and formed a separate, well-defined clade with A-domains of three known MBAs, which suggested that l-Leu-6 in the proposed minimal MK-binding motif were encoded by the most highly conserved A domain in MBA-family peptides.
Extended Data Fig. 2 Three potential MBA BGCs from eSNaPD-guided soil metagenomic mining.
Comparison of NRPS gene organization (a) as well as amino acid substrates (b) between the three known MBA BGCs and the three potential MBA BGCs we cloned from soil metagenomes. The blue residues represent building blocks that are conserved across all MBAs. The green circles represent residues that are shared between known and potential MBAs.
Extended Data Fig. 3 Predicted MBA peptide sequences identified in a motif search of the p-NRP database (a) and the BGCs associated with these predicted peptides (b).
The blue residues represent building blocks that are conserved across all MBAs.
Extended Data Fig. 4 The structures (a) and anti-bacterial activities (b) of the N-acylated peptides associated with known MBAs cyclized in two different ways.
The (R)-3-hydroxy-octanoic acid analogs of lysocin E, WBP-29479A1 and the deoxy version of WAP-8294A1 shown here were synthesized in this study. B. subtilis 168 1A1, S. aureus USA300, S. epidemidis RP62A and M. tuberculosis H37Rv were used as tested strains. The blue residues represent building blocks that are conserved across all MBAs.
Extended Data Fig. 5 Membrane depolarization activity and resistance frequency of MBAs 1 through 6.
a, The effect of each MBA on S. aureus membrane potential was measured using 3,3′-Dipropylthiadicarbocyanine iodide [DiSC3(5)]. Vancomycin (Van) and lysocin were used as the negative and positive controls, respectively. b, Resistance frequency of MBAs 1 through 6 against S. aureus USA300 in the presence of 4x the MIC of each antibiotic.
Extended Data Fig. 6
Isothermal titration of 1:1 (mol/mol) DOPC:DOPG vesicles containing MK into each MBA.
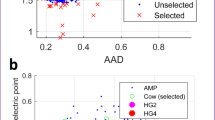
Extended Data Fig. 7 Correlation between antibiotic activity and MK binding affinity for active or inactive syn-BNP MBAs.
Extended Data Fig. 8 Antibiotic activity and MK binding of MBA3 with single point mutations in the proposed minimal MK-binding motif.
MIC in μg/mL, highest concentration tested was 64 μg/mL.
Supplementary information
Supplementary Information
Supplementary Discussion, Tables 1–5 and Figs. 1–19.
Supplementary Data 1
Numerical data for the graphs in Supplementary Fig. 3.
Supplementary Data 2
Numerical data for the graphs in Supplementary Fig. 4.
Source data
Source Data Fig. 3
Numerical data for the graphs in Fig. 3a–c.
Source Data Fig. 4
Numerical data for the graphs in Fig. 4b,c.
Source Data Fig. 6
Numerical data for the graphs in Fig. 6.
Source Data Extended Data Fig. 5
Numerical data for the graph in Extended Data Fig.5a.
Source Data Extended Data Fig. 6
Numerical data for the graphs in Extended Data Fig.6.
Source Data Extended Data Fig. 7
Numerical data for the graphs in Extended Data Fig.7a.
Source Data Extended Data Fig. 8
Numerical data for the graphs in Extended Data Fig.8b.
Rights and permissions
About this article
Cite this article
Li, L., Koirala, B., Hernandez, Y. et al. Identification of structurally diverse menaquinone-binding antibiotics with in vivo activity against multidrug-resistant pathogens. Nat Microbiol 7, 120–131 (2022). https://doi.org/10.1038/s41564-021-01013-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-021-01013-8
This article is cited by
-
Bottom-up synthetic biology approach for improving the efficiency of menaquinone-7 synthesis in Bacillus subtilis
Microbial Cell Factories (2022)