Abstract
Swift recruitment of phagocytic leucocytes is critical in preventing infection when bacteria breach through the protective layers of the skin. According to canonical models, this occurs via an indirect process that is initiated by contact of bacteria with resident skin cells and which is independent of the pathogenic potential of the invader. Here we describe a more rapid mechanism of leucocyte recruitment to the site of intrusion of the important skin pathogen Staphylococcus aureus that is based on direct recognition of specific bacterial toxins, the phenol-soluble modulins (PSMs), by circulating leucocytes. We used a combination of intravital imaging, ear infection and skin abscess models, and in vitro gene expression studies to demonstrate that this early recruitment was dependent on the transcription factor EGR1 and contributed to the prevention of infection. Our findings refine the classical notion of the non-specific and resident cell-dependent character of the innate immune response to bacterial infection by demonstrating a pathogen-specific high-alert mechanism involving direct recruitment of immune effector cells by secreted bacterial products.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Microarray data have been deposited in NCBI’s Gene Expression Omnibus and are accessible through GEO Series accession number GSE103779. All other data generated or analysed during this study are included in this paper or in the Supplementary Information. Source data are provided with this paper.
References
Grice, E. A. & Segre, J. A. The skin microbiome. Nat. Rev. Microbiol. 9, 244–253 (2011).
Naik, S. et al. Commensal–dendritic-cell interaction specifies a unique protective skin immune sig. Nature 520, 104–108 (2015).
Klevens, R. M. et al. Invasive methicillin-resistant Staphylococcus aureus infections in the United States. JAMA 298, 1763–1771 (2007).
Moran, G. J. et al. Methicillin-resistant S. aureus infections among patients in the emergency department. N. Engl. J. Med. 355, 666–674 (2006).
Kolaczkowska, E. & Kubes, P. Neutrophil recruitment and function in health and inflammation. Nat. Rev. Immunol. 13, 159–175 (2013).
Krishna, S. & Miller, L. S. Innate and adaptive immune responses against Staphylococcus aureus skin infections. Semin. Immunopathol. 34, 261–280 (2012).
Kumar, H., Kawai, T. & Akira, S. Pathogen recognition by the innate immune system. Int. Rev. Immunol. 30, 16–34 (2011).
Sadik, C. D. & Luster, A. D. Lipid-cytokine-chemokine cascades orchestrate leukocyte recruitment in inflammation. J. Leukoc. Biol. 91, 207–215 (2012).
Wang, R. et al. Identification of novel cytolytic peptides as key virulence determinants for community-associated MRSA. Nat. Med. 13, 1510–1514 (2007).
Surewaard, B. G. et al. Staphylococcal alpha-phenol soluble modulins contribute to neutrophil lysis after phagocytosis. Cell. Microbiol. 15, 1427–1437 (2013).
Cassat, J. E. et al. A secreted bacterial protease tailors the Staphylococcus aureus virulence repertoire to modulate bone remodeling during osteomyelitis. Cell Host Microbe 13, 759–772 (2013).
Kobayashi, S. D. et al. Comparative analysis of USA300 virulence determinants in a rabbit model of skin and soft tissue infection. J. Infect. Dis. 204, 937–941 (2011).
Nakagawa, S. et al. Staphylococcus aureus virulent PSMalpha peptides induce keratinocyte alarmin release to orchestrate IL-17-dependent skin inflammation. Cell Host Microbe 22, 667–677.e5 (2017).
Weiss, E. et al. Formyl-peptide receptor 2 governs leukocyte influx in local Staphylococcus aureus infections. FASEB J. https://doi.org/10.1096/fj.201700441R (2017).
Kretschmer, D. et al. Human formyl peptide receptor 2 senses highly pathogenic Staphylococcus aureus. Cell Host Microbe 7, 463–473 (2010).
Peschel, A. & Otto, M. Phenol-soluble modulins and staphylococcal infection. Nat. Rev. Microbiol. 11, 667–673 (2013).
Cheung, G. Y., Joo, H. S., Chatterjee, S. S. & Otto, M. Phenol-soluble modulins–critical determinants of staphylococcal virulence. FEMS Microbiol. Rev. 38, 698–719 (2014).
Joo, H. S., Cheung, G. Y. & Otto, M. Antimicrobial activity of community-associated methicillin-resistant Staphylococcus aureus is caused by phenol-soluble modulin derivatives. J. Biol. Chem. 286, 8933–8940 (2011).
Hanzelmann, D. et al. Toll-like receptor 2 activation depends on lipopeptide shedding by bacterial surfactants. Nat. Commun. 7, 12304 (2016).
Queck, S. Y. et al. RNAIII-independent target gene control by the agr quorum-sensing system: insight into the evolution of virulence regulation in Staphylococcus aureus. Mol. Cell 32, 150–158 (2008).
Cheung, G. Y. et al. Insight into structure–function relationship in phenol-soluble modulins using an alanine screen of the phenol-soluble modulin (PSM) alpha3 peptide. FASEB J. 28, 153–161 (2014).
Cheung, G. Y. et al. Staphylococcus epidermidis strategies to avoid killing by human neutrophils. PLoS Pathog. 6, e1001133 (2010).
Prat, C., Bestebroer, J., de Haas, C. J., van Strijp, J. A. & van Kessel, K. P. A new staphylococcal anti-inflammatory protein that antagonizes the formyl peptide receptor-like 1. J. Immunol. 177, 8017–8026 (2006).
Brown, T. J., Rowe, J. M., Liu, J. W. & Shoyab, M. Regulation of IL-6 expression by oncostatin M. J. Immunol. 147, 2175–2180 (1991).
Schutyser, E., Struyf, S. & Van Damme, J. The CC chemokine CCL20 and its receptor CCR6. Cytokine Growth Factor Rev. 14, 409–426 (2003).
Kobayashi, Y. The role of chemokines in neutrophil biology. Front. Biosci. 13, 2400–2407 (2008).
Bertheloot, D. & Latz, E. HMGB1, IL-1alpha, IL-33 and S100 proteins: dual-function alarmins. Cell. Mol. Immunol. 14, 43–64 (2017).
Greenblatt, M. B., Aliprantis, A., Hu, B. & Glimcher, L. H. Calcineurin regulates innate antifungal immunity in neutrophils. J. Exp. Med. 207, 923–931 (2010).
Abdel-Latif, M. M. et al. Helicobacter pylori activates the early growth response 1 protein in gastric epithelial cells. Infect. Immun. 72, 3549–3560 (2004).
Cargnello, M. & Roux, P. P. Activation and function of the MAPKs and their substrates, the MAPK-activated protein kinases. Microbiol. Mol. Biol. Rev. 75, 50–83 (2011).
Sundqvist, M. et al. Staphylococcus aureus-derived PSMalpha peptides activate neutrophil FPR2 but lack the ability to mediate beta-arrestin recruitment and chemotaxis. J. Immunol. https://doi.org/10.4049/jimmunol.1900871 (2019).
Migeotte, I., Communi, D. & Parmentier, M. Formyl peptide receptors: a promiscuous subfamily of G protein-coupled receptors controlling immune responses. Cytokine Growth Factor Rev. 17, 501–519 (2006).
Vuong, C. et al. Regulated expression of pathogen-associated molecular pattern molecules in Staphylococcus epidermidis: quorum-sensing determines pro-inflammatory capacity and production of phenol-soluble modulins. Cell. Microbiol. 6, 753–759 (2004).
Panaro, M. A. & Mitolo, V. Cellular responses to FMLP challenging: a mini-review. Immunopharmacol. Immunotoxicol. 21, 397–419 (1999).
Keates, S., Keates, A. C., Nath, S., Peek, R. M. Jr. & Kelly, C. P. Transactivation of the epidermal growth factor receptor by cag+ Helicobacter pylori induces upregulation of the early growth response gene Egr-1 in gastric epithelial cells. Gut 54, 1363–1369 (2005).
de Grado, M., Rosenberger, C. M., Gauthier, A., Vallance, B. A. & Finlay, B. B. Enteropathogenic Escherichia coli infection induces expression of the early growth response factor by activating mitogen-activated protein kinase cascades in epithelial cells. Infect. Immun. 69, 6217–6224 (2001).
Hannemann, S., Gao, B. & Galan, J. E. Salmonella modulation of host cell gene expression promotes its intracellular growth. PLoS Pathog. 9, e1003668 (2013).
de Klerk, N., Saroj, S. D., Wassing, G. M., Maudsdotter, L. & Jonsson, A. B. The host cell transcription factor EGR1 is induced by bacteria through the EGFR-ERK1/2 pathway. Front. Cell. Infect. Microbiol. 7, 16 (2017).
Xu, Z. et al. Bacterial peptidoglycan-induced tnf-alpha transcription is mediated through the transcription factors Egr-1, Elk-1, and NF-kappaB. J. Immunol. 167, 6975–6982 (2001).
Coleman, D. L., Bartiss, A. H., Sukhatme, V. P., Liu, J. & Rupprecht, H. D. Lipopolysaccharide induces Egr-1 mRNA and protein in murine peritoneal macrophages. J. Immunol. 149, 3045–3051 (1992).
Rautenberg, M., Joo, H. S., Otto, M. & Peschel, A. Neutrophil responses to staphylococcal pathogens and commensals via the formyl peptide receptor 2 relates to phenol-soluble modulin release and virulence. FASEB J. 25, 1254–1263 (2011).
Chatterjee, S. S. et al. Essential Staphylococcus aureus toxin export system. Nat. Med. 19, 364–367 (2013).
Dahlgren, C., Gabl, M., Holdfeldt, A., Winther, M. & Forsman, H. Basic characteristics of the neutrophil receptors that recognize formylated peptides, a danger-associated molecular pattern generated by bacteria and mitochondria. Biochem. Pharmacol. 114, 22–39 (2016).
Forsman, H. et al. Structural changes of the ligand and of the receptor alters the receptor preference for neutrophil activating peptides starting with a formylmethionyl group. Biochim. Biophys. Acta 1853, 192–200 (2015).
Qin, L. et al. Toxin mediates sepsis caused by methicillin-resistant Staphylococcus epidermidis. PLoS Pathog. 13, e1006153 (2017).
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative C(T) method. Nat. Protoc. 3, 1101–1108 (2008).
Faust, N., Varas, F., Kelly, L. M., Heck, S. & Graf, T. Insertion of enhanced green fluorescent protein into the lysozyme gene creates mice with green fluorescent granulocytes and macrophages. Blood 96, 719–726 (2000).
Kamenyeva, O. et al. Neutrophil recruitment to lymph nodes limits local humoral response to Staphylococcus aureus. PLoS Pathog. 11, e1004827 (2015).
Acknowledgements
This study was supported by the Intramural Research Program of the National Institute of Allergy and Infectious Diseases (NIAID) and the National Cancer Institute (NCI), US National Institutes of Health (NIH), project numbers ZIA AI000904 (M.O.), ZIA AI001079 (F.R.D.), ZIA AI001171 (D.L.B.), ZIA BC010725 (J.M.W.), and by federal funds from the NCI (contract no. HSN261200800001E, to J.M.W.). F.M.F.A. received a scholarship from Shaqra University, Al Quwaiiyah, Saudi Arabia. We thank K. v. Kessel, University of Utrecht, for supplying FLIPr.
Author information
Authors and Affiliations
Contributions
M.O. conceptualized the study. D.L.B. and T.H.N. assisted in planning the leucocyte influx experiments, and F.R.D., the microarray experiments. M.O. and F.R.D. supervised experiments. G.Y.C.C. and T.H.N. designed and set up, and G.Y.C.C., T.H.N., R.L., J.S.B., P.P., S.F., J.C., M.D.P., R.L.H., F.M.F.A., V.Y.T., T.K.A., J.W.M., E.L.F. and A.J.Y. performed the leucocyte influx and intravital imaging experiments. O.K. performed confocal microscopy. J.K., O.K., G.Y.C.C. and T.H.N. analysed imaging data. K.M.R., A.R.P. and S.D.K. performed neutrophil array and corresponding control experiments. D.E.S. analysed array data. T.H.N. and A.E.V. performed RT-qPCR, and S.F. and T.H.N., pathway analysis. J.M.W. supplied FPR2−/− mouse breeding pairs. M.O. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Microbiology thanks Mia Phillipson and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Bacterial numbers and psm gene expression during early ear infection.
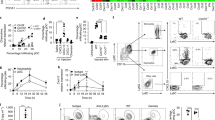
a, CFU in ears infected with wild-type LAC or Δpsm S. aureus, and with PBS as control to analyze for contaminating skin bacteria. n = 3/group. b, Expression of the psmα locus by S. aureus LAC bacteria during early ear and back skin infection. n = 4/group and time point. a,b, Error bars show the mean ± SD.
Extended Data Fig. 2 S. aureus PSM toxins are critical to the early leukocyte influx to the site of skin infection – Experiments with pure peptides.
a, Neutrophil influx measured after injection of 3 μl of a 1 μM solution of PSMα3 peptide in water over 6 h p. i. (n = 3), and influx of total leukocytes, neutrophils, and non-neutrophil leukocytes 5 h p. i. after injection of 3 μl of a 1 μM solution of PSMα3 or PSMα3K12A, or 3 μl water as control. n = 7/group (peptides); n = 4/group (water control). Statistical analysis is by 1-way ANOVA with Dunnett’s post-test versus the data obtained at 1 h p. i. (left panel) or Tukey’s post-tests (other panels). Error bars show the mean ± SD. b, Selected representative images. Green, leukocytes; red, blood vessels.
Extended Data Fig. 3 Establishment of sublytic and pro-inflammatory culture filtrate dilutions or pure PSMs and impact of FLIPr.
a, Test of S. aureus culture filtrate dilutions and PSMα3 and δ-toxin concentrations on cytolytic activity toward human neutrophils by release of LDH. n = 3/group. Error bars show the mean ± SD. b-d, Test by flow cytometry of the impact of FLIPr (green line) on blocking of induced pro-inflammatory effect in human neutrophils (expression of CD11b) by different concentrations of PSMα3 or δ-toxin. FLIPr was tested at concentrations between 0.5 and 10 μg/ml. Shown are the 0.5 μg/ml results for PSMα3 and the 1.0 μg/ml results for δ-toxin. Tests were performed ranging from n = 1 to n = 6 at concentrations close to the range of interest. A gate was set during data analysis to exclude dead cells and debris (as indicated in panel d). Error bars show the mean ± SD.
Extended Data Fig. 4 Verification of differential EGR1 expression in selected comparisons by RT-qPCR.
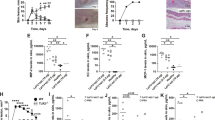
a, Comparison of the impact of diluted culture filtrates of S. aureus wild-type (WT) versus Δpsm and Δpsmα mutants on EGR1 expression in human neutrophils. Statistical analysis is by 1-way ANOVA with Tukey’s post-test. n = 14. b, Impact of FLIPr on stimulation by S. aureus WT culture filtrate. n = 5. c, Impact of FLIPr on stimulation by PSMα3. n = 3. b,c, Statistical analysis is by two-tailed unpaired Student’s t-test. a-c, Error bars show the mean ± SD.
Extended Data Fig. 5 Determination of leukocyte infiltration into spleens for normalization in the adoptive transfer experiment.
a-c, Exemplary confocal pictures of spleens. Leukocytes were labeled with different dyes and visualized as follows: leukocytes from wild-type mice in magenta, from FPR2-/- mice in green, and from EGR1-/- mice in blue. Note that computation and analysis was performed in a 3D manner; the pictures only show 2D slices. The exemplary pictures shown here are from the same mice as those in Extended Data Fig. 6.
Extended Data Fig. 6 Leukocyte attraction via the FPR2-EGR1 pathway is direct and independent of resident skin cells – exemplary confocal microscopy pictures.
a, The pictures show the unprocessed images and the two steps of processing: masking of hairs and ROI determination (see methods). Leukocytes are labeled with different dyes: leukocytes from wild-type mice in magenta, from FPR2-/- mice in green, and from EGR1-/- mice in blue. Dye-labeled bacteria are in cyan. Central, non-shaded areas represent the analyzed, computed regions of interest (ROIs). Note that computation and analysis was performed in a 3D manner; the pictures only show 2D slices. b, Processed WT mouse picture in higher magnification centered on the injected bacteria. The exemplary pictures shown here are from the same mice as those in Extended Data Fig. 5 showing spleen controls.
Extended Data Fig. 7 Leukocyte attraction via the FPR2-EGR1 pathway is direct and independent of resident skin cells – FPR2-/- and EGR1-/- comparisons.
Shown are the comparisons with FPR2-/- and EGR1-/- recipient mice versus FPR2-/- and EGR1-/- donor leukocyte comparisons, in analogy to the comparisons with wild-type recipient mice versus wild-type donor leukocytes shown in Fig. 4c,d,f,g. Statistical analysis is by repeated measures ANOVA with Dunnett’s post-test versus WT (wild-type). Error bars show the mean ± SD. n = 4-6. For the comparisons with mixed group numbers, a mixed model (rather than ANOVA) was automatically employed by Prism for data analysis.
Extended Data Fig. 8 Activation of mouse neutrophils by fMLP.
Neutrophil activation was assessed by determination of Ca2+ flux. 1 × 105 neutrophils isolated from wild-type, EGR1-/-, and FPR2-/- mice were stimulated with fMLP in DMSO at the indicated concentrations. DMSO controls are shown. Error bars show the mean ± SD. n = 8/group.
Supplementary information
Source data
Source Data Fig. 1
Numerical source data and associated statistical analysis.
Source Data Fig. 2
Numerical source data and associated statistical analysis.
Source Data Fig. 3
Numerical source data and associated statistical analysis.
Source Data Fig. 4
Numerical source data and associated statistical analysis.
Source Data Fig. 5
Numerical source data and associated statistical analysis.
Source Data Extended Data Fig. 1
Numerical source data.
Source Data Extended Data Fig. 2
Numerical source data and associated statistical analysis.
Source Data Extended Data Fig. 3
Numerical source data.
Source Data Extended Data Fig. 4
Numerical source data and associated statistical analysis.
Source Data Extended Data Fig. 7
Numerical source data and associated statistical analysis.
Source Data Extended Data Fig. 8
Numerical source data.
Rights and permissions
About this article
Cite this article
Nguyen, T.H., Cheung, G.Y.C., Rigby, K.M. et al. Rapid pathogen-specific recruitment of immune effector cells in the skin by secreted toxins. Nat Microbiol 7, 62–72 (2022). https://doi.org/10.1038/s41564-021-01012-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-021-01012-9
This article is cited by
-
Characterization of immunomodulating agents from Staphylococcus aureus for priming immunotherapy in triple-negative breast cancers
Scientific Reports (2024)
-
Staphylococcus aureus PSMs are a double-edged sword
Nature Microbiology (2021)