Abstract
Infection of mammalian cells with viruses activates NF-κB to induce the expression of cytokines and chemokines and initiate an antiviral response. Here, we show that a vaccinia virus protein mimics the transactivation domain of the p65 subunit of NF-κB to inhibit selectively the expression of NF-κB-regulated genes. Using co-immunoprecipitation assays, we found that the vaccinia virus protein F14 associates with NF-κB co-activator CREB-binding protein (CBP) and disrupts the interaction between p65 and CBP. This abrogates CBP-mediated acetylation of p65, after which it reduces promoter recruitment of the transcriptional regulator BRD4 and diminishes stimulation of NF-κB-regulated genes CXCL10 and CCL2. Recruitment of BRD4 to the promoters of NFKBIA and CXCL8 remains unaffected by either F14 or JQ1 (a competitive inhibitor of BRD4 bromodomains), indicating that BRD4 recruitment is acetylation-independent. Unlike other viral proteins that are general antagonists of NF-κB, F14 is a selective inhibitor of NF-κB-dependent gene expression. An in vivo model of infection demonstrated that F14 promotes virulence. Molecular mimicry of NF-κB may be conserved because other orthopoxviruses, including variola, monkeypox and cowpox viruses, encode orthologues of F14.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The authors declare that the main data supporting the findings of this study are available within the article and its Supplementary Information. Poxvirus nucleotide sequences mentioned in this study are publicly available on NCBI GenBank: VACV Western Reserve (NC_006998.1), VACV Copenhagen (M35027.1), VACV Modified Virus Ankara (AY603355.1), horsepox virus isolate MNR-76 (DQ792504.1), monkeypox virus Zaire-96-I-16 (NC_003310.10, cowpox virus Brighton Red (NC_003663.2), variola virus India-1967 (NC_001611.1), camelpox virus CMS (AY009089.1), taterapox virus Dahomey 1968 (DQ437594.1), ectromelia virus Moscow (AF012825.2), raccoonpox virus Herman (NC_027213.1), premodern variola virus (LR800247.1, LR800244.1, LR800245.1, LR800246.1) and variola virus VD21 (KY358055.1). Source data are provided with this paper.
References
Iwasaki, A. A virological view of innate immune recognition. Annu. Rev. Microbiol. 66, 177–196 (2012).
Vallabhapurapu, S. & Karin, M. Regulation and function of NF-kappaB transcription factors in the immune system. Annu. Rev. Immunol. 27, 693–733 (2009).
Chen, L. F. & Greene, W. C. Shaping the nuclear action of NF-kappaB. Nat. Rev. Mol. Cell Biol. 5, 392–401 (2004).
Tian, B., Nowak, D. E. & Brasier, A. R. A TNF-induced gene expression program under oscillatory NF-kappaB control. BMC Genom. 6, 137 (2005).
Zhao, M. et al. Transcriptional outcomes and kinetic patterning of gene expression in response to NF-kappaB activation. PLoS Biol. 16, e2006347 (2018).
Kaikkonen, M. U. et al. Remodeling of the enhancer landscape during macrophage activation is coupled to enhancer transcription. Mol. Cell 51, 310–325 (2013).
Chen, L. F. et al. NF-kappaB RelA phosphorylation regulates RelA acetylation. Mol. Cell. Biol. 25, 7966–7975 (2005).
Zhong, H., Voll, R. E. & Ghosh, S. Phosphorylation of NF-kappa B p65 by PKA stimulates transcriptional activity by promoting a novel bivalent interaction with the coactivator CBP/p300. Mol. Cell 1, 661–671 (1998).
Huang, B. et al. Brd4 coactivates transcriptional activation of NF-kappaB via specific binding to acetylated RelA. Mol. Cell. Biol. 29, 1375–1387 (2009).
Huang, S. M. & McCance, D. J. Down regulation of the interleukin-8 promoter by human papillomavirus type 16 E6 and E7 through effects on CREB binding protein/p300 and P/CAF. J. Virol. 76, 8710–8721 (2002).
Xing, J. et al. Herpes simplex virus 1-encoded tegument protein VP16 abrogates the production of beta interferon (IFN) by inhibiting NF-kappaB activation and blocking IFN regulatory factor 3 to recruit its coactivator CBP. J. Virol. 87, 9788–9801 (2013).
Bahar, M. W. et al. How vaccinia virus has evolved to subvert the host immune response. J. Struct. Biol. 175, 127–134 (2011).
Smith, G. L. et al. Vaccinia virus immune evasion: mechanisms, virulence and immunogenicity. J. Gen. Virol. 94, 2367–2392 (2013).
Albarnaz, J. D., Torres, A. A. & Smith, G. L. Modulating vaccinia virus immunomodulators to improve immunological memory. Viruses 10, 101 (2018).
Sumner, R. P. et al. Vaccinia virus inhibits NF-kappaB-dependent gene expression downstream of p65 translocation. J. Virol. 88, 3092–3102 (2014).
Perkus, M. E. et al. Deletion of 55 open reading frames from the termini of vaccinia virus. Virology 180, 406–410 (1991).
Assarsson, E. et al. Kinetic analysis of a complete poxvirus transcriptome reveals an immediate-early class of genes. Proc. Natl Acad. Sci. USA 105, 2140–2145 (2008).
Yang, Z. et al. Simultaneous high-resolution analysis of vaccinia virus and host cell transcriptomes by deep RNA sequencing. Proc. Natl Acad. Sci. USA 107, 11513–11518 (2010).
Wente, S. R. & Rout, M. P. The nuclear pore complex and nuclear transport. Cold Spring Harb. Perspect. Biol. 2, a000562 (2010).
Gubser, C. et al. Poxvirus genomes: a phylogenetic analysis. J. Gen. Virol. 85, 105–117 (2004).
Ferguson, B. J. et al. Vaccinia virus protein N2 is a nuclear IRF3 inhibitor that promotes virulence. J. Gen. Virol. 94, 2070–2081 (2013).
Chen, R. A. et al. Inhibition of IkappaB kinase by vaccinia virus virulence factor B14. PLoS Pathog. 4, e22 (2008).
Stuart, J. H. et al. Vaccinia virus protein C6 inhibits type I IFN signalling in the nucleus and binds to the transactivation domain of STAT2. PLoS Pathog. 12, e1005955 (2016).
Torres, A. A. et al. Multiple Bcl-2 family immunomodulators from vaccinia virus regulate MAPK/AP-1 activation. J. Gen. Virol. 97, 2346–2351 (2016).
Yang, Z. et al. Deciphering poxvirus gene expression by RNA sequencing and ribosome profiling. J. Virol. 89, 6874–6886 (2015).
Yang, Z. et al. Genome-wide analysis of the 5′ and 3′ ends of vaccinia virus early mRNAs delineates regulatory sequences of annotated and anomalous transcripts. J. Virol. 85, 5897–5909 (2011).
Unterholzner, L. et al. Vaccinia virus protein C6 is a virulence factor that binds TBK-1 adaptor proteins and inhibits activation of IRF3 and IRF7. PLoS Pathog. 7, e1002247 (2011).
Soday, L. et al. Quantitative temporal proteomic analysis of vaccinia virus infection reveals regulation of histone deacetylases by an interferon antagonist. Cell Rep. 27, 1920–1933 (2019).
Parrish, S. & Moss, B. Characterization of a vaccinia virus mutant with a deletion of the D10R gene encoding a putative negative regulator of gene expression. J. Virol. 80, 553–561 (2006).
Ehlers, A. et al. Poxvirus orthologous clusters (POCs). Bioinformatics 18, 1544–1545 (2002).
Tamosiunaite, A. et al. What a difference a gene makes: identification of virulence factors of cowpox virus. J. Virol. 94, e01625–19 (2020).
Muhlemann, B. et al. Diverse variola virus (smallpox) strains were widespread in northern Europe in the Viking Age. Science 369, eaaw8977 (2020).
Duggan, A. T. et al. 17th century variola virus reveals the recent history of smallpox. Curr. Biol. 26, 3407–3412 (2016).
Kelley, L. A. et al. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Lecoq, L. et al. Structural characterization of interactions between transactivation domain 1 of the p65 subunit of NF-kappaB and transcription regulatory factors. Nucleic Acids Res. 45, 5564–5576 (2017).
Schmitz, M. L. & Baeuerle, P. A. The p65 subunit is responsible for the strong transcription activating potential of NF-kappa B. EMBO J. 10, 3805–3817 (1991).
Schmitz, M. L. et al. Interaction of the COOH-terminal transactivation domain of p65 NF-kappa B with TATA-binding protein, transcription factor IIB, and coactivators. J. Biol. Chem. 270, 7219–7226 (1995).
Schmitz, M. L. et al. Structural and functional analysis of the NF-kappa B p65 C terminus: an acidic and modular transactivation domain with the potential to adopt an alpha-helical conformation. J. Biol. Chem. 269, 25613–25620 (1994).
Blair, W. S. et al. Mutational analysis of the transcription activation domain of RelA: identification of a highly synergistic minimal acidic activation module. Mol. Cell. Biol. 14, 7226–7234 (1994).
Yang, F. et al. IKK beta plays an essential role in the phosphorylation of RelA/p65 on serine 536 induced by lipopolysaccharide. J. Immunol. 170, 5630–5635 (2003).
Richards, K. H. et al. The human papillomavirus (HPV) E7 protein antagonises an Imiquimod-induced inflammatory pathway in primary human keratinocytes. Sci. Rep. 5, 12922 (2015).
Spitkovsky, D. et al. The human papillomavirus oncoprotein E7 attenuates NF-kappa B activation by targeting the Ikappa B kinase complex. J. Biol. Chem. 277, 25576–25582 (2002).
Vandermark, E. R. et al. Human papillomavirus type 16 E6 and E 7 proteins alter NF-kB in cultured cervical epithelial cells and inhibition of NF-kB promotes cell growth and immortalization. Virology 425, 53–60 (2012).
Shi, J. & Vakoc, C. R. The mechanisms behind the therapeutic activity of BET bromodomain inhibition. Mol. Cell 54, 728–736 (2014).
Devaiah, B. N., Gegonne, A. & Singer, D. S. Bromodomain 4: a cellular Swiss army knife. J. Leukoc. Biol. 100, 679–686 (2016).
Filippakopoulos, P. et al. Selective inhibition of BET bromodomains. Nature 468, 1067–1073 (2010).
Hargreaves, D. C., Horng, T. & Medzhitov, R. Control of inducible gene expression by signal-dependent transcriptional elongation. Cell 138, 129–145 (2009).
Mochizuki, K. et al. The bromodomain protein Brd4 stimulates G1 gene transcription and promotes progression to S phase. J. Biol. Chem. 283, 9040–9048 (2008).
Pallett, M. A. et al. Vaccinia Virus BBK E3 Ligase Adaptor A55 Targets Importin-Dependent NF-kappaB Activation and Inhibits CD8(+) T-Cell Memory. J. Virol. 93, e00051–19 (2019).
Eaglesham, J. B. et al. Viral and metazoan poxins are cGAMP-specific nucleases that restrict cGAS-STING signalling. Nature 566, 259–263 (2019).
Arenzana-Seisdedos, F. et al. Inducible nuclear expression of newly synthesized I kappa B alpha negatively regulates DNA-binding and transcriptional activities of NF-kappa B. Mol. Cell. Biol. 15, 2689–2696 (1995).
Arenzana-Seisdedos, F. et al. Nuclear localization of I kappa B alpha promotes active transport of NF-kappa B from the nucleus to the cytoplasm. J. Cell Sci. 110, 369–378 (1997).
Kanno, T. et al. BRD4 assists elongation of both coding and enhancer RNAs by interacting with acetylated histones. Nat. Struct. Mol. Biol. 21, 1047–1057 (2014).
Mukherjee, S. P. et al. Analysis of the RelA:CBP/p300 interaction reveals its involvement in NF-kappaB-driven transcription. PLoS Biol. 11, e1001647 (2013).
Staller, M. V. et al. A high-throughput mutational scan of an intrinsically disordered acidic transcriptional activation domain. Cell Syst. 6, 444–455 (2018).
Neidel, S. et al. NF-kappaB activation is a turn on for vaccinia virus phosphoprotein A49 to turn off NF-kappaB activation. Proc. Natl Acad. Sci. USA 116, 5699–5704 (2019).
Mansur, D. S. et al. Poxvirus targeting of E3 ligase beta-TrCP by molecular mimicry: a mechanism to inhibit NF-kappaB activation and promote immune evasion and virulence. PLoS Pathog. 9, e1003183 (2013).
Bravo Cruz, A. G. & Shisler, J. L. Vaccinia virus K1 ankyrin repeat protein inhibits NF-kappaB activation by preventing RelA acetylation. J. Gen. Virol. 97, 2691–2702 (2016).
Diel, D. G. et al. A nuclear inhibitor of NF-kappaB encoded by a poxvirus. J. Virol. 85, 264–275 (2011).
O’Connor, M. J. et al. Characterization of an E1A-CBP interaction defines a novel transcriptional adapter motif (TRAM) in CBP/p300. J. Virol. 73, 3574–3581 (1999).
Marzio, G. et al. HIV-1 tat transactivator recruits p300 and CREB-binding protein histone acetyltransferases to the viral promoter. Proc. Natl Acad. Sci. USA 95, 13519–13524 (1998).
Patel, D. et al. The E6 protein of human papillomavirus type 16 binds to and inhibits co-activation by CBP and p300. EMBO J. 18, 5061–5072 (1999).
Cook, J. L. et al. Role of the E1A Rb-binding domain in repression of the NF-kappa B-dependent defense against tumor necrosis factor-alpha. Proc. Natl Acad. Sci. USA 99, 9966–9971 (2002).
Naar, A. M. et al. Composite co-activator ARC mediates chromatin-directed transcriptional activation. Nature 398, 828–832 (1999).
Richardson, P. M. & Gilmore, T. D. vRel is an inactive member of the Rel family of transcriptional activating proteins. J. Virol. 65, 3122–3130 (1991).
Lin, J. R. et al. Minimalist ensemble algorithms for genome-wide protein localization prediction. BMC Bioinf. 13, 157 (2012).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014).
Gloeckner, C. J. et al. A novel tandem affinity purification strategy for the efficient isolation and characterisation of native protein complexes. Proteomics 7, 4228–4234 (2007).
Maluquer de Motes, C. et al. Vaccinia virus virulence factor N1 can be ubiquitylated on multiple lysine residues. J. Gen. Virol. 95, 2038–2049 (2014).
Falkner, F. G. & Moss, B. Transient dominant selection of recombinant vaccinia viruses. J. Virol. 64, 3108–3111 (1990).
Tscharke, D. C. & Smith, G. L. A model for vaccinia virus pathogenesis and immunity based on intradermal injection of mouse ear pinnae. J. Gen. Virol. 80, 2751–2755 (1999).
Williamson, J. D. et al. Biological characterization of recombinant vaccinia viruses in mice infected by the respiratory route. J. Gen. Virol. 71, 2761–2767 (1990).
Harris, D. P. et al. Tumor necrosis factor (TNF)-alpha induction of CXCL10 in endothelial cells requires protein arginine methyltransferase 5 (PRMT5)-mediated nuclear factor (NF)-kappaB p65 methylation. J. Biol. Chem. 289, 15328–15339 (2014).
Xu, X. et al. EVI1 acts as an inducible negative-feedback regulator of NF-kappaB by inhibiting p65 acetylation. J. Immunol. 188, 6371–6380 (2012).
Acknowledgements
We thank R. Seear, S. Macilwee and J. Milburn for technical support and F. Pfaff and M. Beer (Friedrich-Loeffler-Institut, Germany) for help with access to the cowpox RNA sequencing dataset. We also thank J. Doorbar and C. Crump (Department of Pathology, University of Cambridge, UK), T. Kouzarides (Department of Pathology and The Gurdon Institute, University of Cambridge, UK) and G. Blobel (University of Pennsylvania, Philadelphia, USA) for providing us with reagents. We are grateful to T. Kouzarides for helpful advice and to C. Talbot-Cooper for critical reading of the manuscript. This work was supported by grant no. 090315 from the Wellcome Trust (to G.L.S.). B.Y-W.C.’s laboratory is funded by the Medical Research Council (grant no. MR/R021821/1), the Biotechnology and Biological Sciences Research Council (grant no. BB/V017780.1) and Isaac Newton Trust (grant no. G101522). J.D.A. was a postdoctoral fellow of the Science without Borders programme from CNPq-Brazil (grant no. 235246/2014-0). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
J.D.A., A.A.T. and G.L.S. conceived the idea. J.D.A., H.R., A.A.T., E.V.S., C.A.M., A.J.B., M.P.B. and B.Y-W.C. developed the methodology. J.D.A., H.R., A.A.T. and E.V.S. validated the results. J.D.A. and H.R. undertook the formal analysis. J.D.A., H.R., A.A.T. and E.V.S. undertook investigation. A.A.T., C.A.M., A.J.B., B.Y-W.C. and G.L.S. obtained resources. J.D.A. undertook data curation. J.D.A. wrote the original draft. J.D.A., H.R., A.A.T., C.A.M., A.J.B., B.Y-W.C. and G.L.S. reviewed and edited the manuscript. J.D.A. prepared visualization. J.D.A. and G.L.S. supervised the work. J.D.A. and G.L.S. were project administrators. J.D.A. and G.L.S. obtained funding. A.A.T., C.A.M., A.J.B., B.Y-W.C. and G.L.S. contributed resources.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Microbiology thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Screen of VACV strain WR ORFs for NF-κB inhibitory activity.
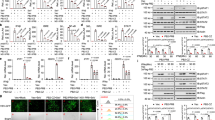
NF-κB-dependent luciferase activity in HEK 293T cells transfected with vectors expressing the indicated VACV proteins or empty vector (EV), and stimulated with TNF-α. Negative (EV, GFP, and N2) and positive (B14) controls are highlighted in the dashed black square, whilst F14 is highlighted in the dashed red square. Means + s.d. (n = 3-4 per condition) are shown.
Extended Data Fig. 2 Virulence of VACV mutant lacking F14 in the intranasal mouse model of infection.
BALB/c mice were infected intranasally with 5×103 p.f.u. of the indicated VACV strains and their body mass was measured daily. Body mass is expressed as the percentage ± s.e.m. of the mean of the same group of mice on day 0 (n = 5 mice).
Extended Data Fig. 3 Replication and spread of VACV mutant lacking F14 in cell culture.
(a,b) HeLa cells were infected with the indicated VACV strains (5 p.f.u./cell) and virus titres associated with the cells (a) and in the supernatants (b) were determined by plaque assay. Means (n = 2 per condition) are shown. (c) Plaque formation by the indicated VACV strains on BS-C-1 cells.
Extended Data Fig. 4 F14 does not inhibit the nuclear translocation of NF-κB subunit p65.
a, T-REx-293 cells inducibly expressing the empty vector (EV) or VACV proteins B14, C6, or F14, were induced with doxycycline and stimulated with TNF-α. Fixed and permeabilised cells were stained with anti-p65 antibody and DAPI, and analysed by confocal microscopy. Scale bars (50 μm) are shown in the bottom right of each micrograph. Representative micrographs of quantitative analysis shown in Fig. 1l. b, Flow cytometry analysis of T-REx-293-F14 induced with doxycycline in the absence and in the presence of the proteasome inhibitor MG132. F14 presence was detected by staining with an anti-FLAG antibody.
Extended Data Fig. 5 F14 inhibits NF-κB at or downstream of p65.
NF-κB activity in HEK 293 T cells transfected with vectors expressing p65, VACV proteins B14 or F14, or empty vector (EV). Top panel: Means + s.d. (n = 4 per condition) are shown. Statistical significance was determined by unpaired two-tailed Student’s t-test. Bottom panel: Immunoblotting. Protein molecular mass markers in kDa are shown on the left of the blots. Immunoblots of tagged proteins are labelled with the protein name followed by the epitope tag antibody in parentheses. When multiple tagged proteins are shown in the same immunoblot, each protein is indicated by a red arrowhead.
Extended Data Fig. 6 F14 is unique among known viral inhibitors of NF-κB.
(a) Top: Amino acid sequence of the TAD of HSV-1 VP16 with the acidic activation domain similar to p65 highlighted in red, and hydrophobic residues (Φ) are indicated. Middle: NF-κB-dependent luciferase activity in HEK 293T cells transfected with vectors expressing VP16, VP16 mutant, or empty vector (EV), and stimulated with TNF-α. Bottom: Immunoblotting. (b) Top: Amino acid residues 61-98 from HPV16 protein E7 encompassing a ΦXXΦΦ motif containing and preceded by negatively charged residues. Highlighted are two residues mutated to disrupt this motif. Middle: NF-κB-dependent luciferase activity in HEK 293T cells expressing E7 and two mutants as described in (a). Bottom: Immunoblotting. Means + s.d. (n = 4 per condition) are shown. c, Lysates from transfected HEK 293T cells were immunoprecipitated with anti-HA. Immunoblots are representative of two independent experiments. Protein molecular masses in kDa are shown on the left of the blots. Immunoblots of tagged proteins are labelled with the protein name followed the epitope tag antibody in parentheses. When multiple tagged proteins are shown in the same immunoblot, each protein is indicated a red arrowhead. Statistical significance was determined by the Student’s t-test.
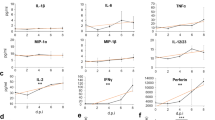
Extended Data Fig. 7 F14 suppresses expression of a subset of NF-κB-responsive genes.
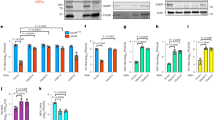
(a-d) RT-qPCR analysis of NF-κB-responsive gene expression in inducible T-REx-293 cells induced with doxycycline overnight to express VACV proteins F14 or C6, and stimulated with TNF-α. Means (n = 2 per condition) are shown. e, Immunoblotting of lysates of inducible T-REx-293 cell lines induced with doxycycline overnight. Data are representative of two independent experiments. Protein molecular masses in kDa are shown on the left of the blots.
Extended Data Fig. 8 F14 suppresses expression of CXCL10, but not CXCL8, after stimulation with TNF-α.
This shows data normalized for presentation in Fig. 5c,f. ELISA of CXCL8 (a) and CXCL10 (b) in culture supernatants from T-REx-293 cells inducibly expressing the empty vector (EV) or VACV proteins B14, C6, or F14, induced with doxycycline and stimulated with TNF-α. Means + s.d. (n = 3 per condition) are shown. Statistical significance was determined by the Student’s t-test.
Extended Data Fig. 9 JQ1 reduces BRD4 occupancy on CCND1 gene promoter.
Chromatin immunoprecipitation (ChIP) with anti-BRD4 antibody or control IgG, and qPCR for the promoters of CCND1 gene. T-REx-293 cells were treated with JQ1 and stimulated with TNF-α. Means + s.d. (n = 6 per condition from two independent experiments). Statistical significance was determined by the Student’s t-test.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2.
Source data
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Unprocessed western blots.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Unprocessed western blots.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Unprocessed western blots.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Rights and permissions
About this article
Cite this article
Albarnaz, J.D., Ren, H., Torres, A.A. et al. Molecular mimicry of NF-κB by vaccinia virus protein enables selective inhibition of antiviral responses. Nat Microbiol 7, 154–168 (2022). https://doi.org/10.1038/s41564-021-01004-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-021-01004-9
This article is cited by
-
Mpox virus infection and drug treatment modelled in human skin organoids
Nature Microbiology (2023)
-
TRIM5α restricts poxviruses and is antagonized by CypA and the viral protein C6
Nature (2023)