Abstract
Alveolata comprises diverse taxa of single-celled eukaryotes, many of which are renowned for their ability to live inside animal cells. Notable examples are apicomplexan parasites and dinoflagellate symbionts, the latter of which power coral reef ecosystems. Although functionally distinct, they evolved from a common, free-living ancestor and must evade their host’s immune response for persistence. Both the initial cellular events that gave rise to this intracellular lifestyle and the role of host immune modulation in coral–dinoflagellate endosymbiosis are poorly understood. Here, we use a comparative approach in the cnidarian endosymbiosis model Aiptasia, which re-establishes endosymbiosis with free-living dinoflagellates every generation. We find that uptake of microalgae is largely indiscriminate, but non-symbiotic microalgae are expelled by vomocytosis, while symbionts induce host cell innate immune suppression and form a lysosomal-associated membrane protein 1-positive niche. We demonstrate that exogenous immune stimulation results in symbiont expulsion and, conversely, inhibition of canonical Toll-like receptor signalling enhances infection of host animals. Our findings indicate that symbiosis establishment is dictated by local innate immune suppression, to circumvent expulsion and promote niche formation. This work provides insight into the evolution of the cellular immune response and key steps involved in mediating endosymbiotic interactions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Raw reads of the RNA sequencing data can be accessed at the National Center for Biotechnology Information SRA with the following accession numbers: SRX7119772–7119776 (cells from aposymbiotic larvae), SRX7119782–7119787 (symbiotic cells) and SRX7119777–7119781 (aposymbiotic cells from symbiotic larvae) (combined in the SRA project SRP229372); and SRX7229078–7229080 (M. gaditana-containing cells) and SRX7229075–7229077 (microalgae-free cells from M. gaditana-containing larvae) (combined in the SRA project SRP233508). Source data are provided with this paper.
Code availability
The Knime92 workflow used for analysis, as well as other raw data and R scripts for analysis, can be found at https://doi.org/10.24433/CO.0872345.v1.
References
Saldarriaga, J. F., Taylor, F. J. R., Cavalier-Smith, T., Menden-Deuer, S. & Keeling, P. J. Molecular data and the evolutionary history of dinoflagellates. Eur. J. Protistol. 40, 85–111 (2004).
Seeber, F. & Steinfelder, S. Recent advances in understanding apicomplexan parasites. F1000Res. 5, 1369 (2016).
Kwong, W. K., del Campo, J., Mathur, V., Vermeij, M. J. A. & Keeling, P. J. A widespread coral-infecting apicomplexan with chlorophyll biosynthesis genes. Nature 568, 103–107 (2019).
De Vargas, C. et al. Eukaryotic plankton diversity in the sunlit ocean. Science 348, 1261605 (2015).
Baker, A. C. Flexibility and specificity in coral–algal symbiosis: diversity, ecology, and biogeography of Symbiodinium. Annu. Rev. Ecol. Evol. Syst. 34, 661–689 (2003).
Yellowlees, D., Rees, T. A. V. & Leggat, W. Metabolic interactions between algal symbionts and invertebrate hosts. Plant Cell Environ. 31, 679–694 (2008).
Janouškovec, J. et al. Apicomplexan-like parasites are polyphyletic and widely but selectively dependent on cryptic plastid organelles. eLife 8, e49662 (2019).
Janouškovec, J. et al. Factors mediating plastid dependency and the origins of parasitism in apicomplexans and their close relatives. Proc. Natl Acad. Sci. USA 112, 10200–10207 (2015).
Janouškovec, J., Horák, A., Oborník, M., Lukeš, J. & Keeling, P. J. A common red algal origin of the apicomplexan, dinoflagellate, and heterokont plastids. Proc. Natl Acad. Sci. USA 107, 10949–10954 (2010).
Baird, A. H., Guest, J. R. & Willis, B. L. Systematic and biogeographical patterns in the reproductive biology of scleractinian corals. Annu. Rev. Ecol. Evol. Syst. 40, 551–571 (2009).
Schwarz, J. A., Krupp, D. A. & Weis, V. M. Late larval development and onset of symbiosis in the scleractinian coral Fungia scutaria. Biol. Bull. 196, 70–79 (1999).
Mansfield, K. M. & Gilmore, T. D. Innate immunity and cnidarian–Symbiodiniaceae mutualism. Dev. Comp. Immunol. 90, 199–209 (2018).
Poole, A. Z. & Weis, V. M. TIR-domain-containing protein repertoire of nine anthozoan species reveals coral-specific expansions and uncharacterized proteins. Dev. Comp. Immunol. 46, 480–488 (2014).
Buchmann, K. Evolution of innate immunity: clues from invertebrates via fish to mammals. Front. Immunol. 5, 459 (2014).
Ghosh, J. et al. Invertebrate immune diversity. Dev. Comp. Immunol. 35, 959–974 (2011).
Detournay, O., Schnitzler, C. E., Poole, A. & Weis, V. M. Regulation of cnidarian–dinoflagellate mutualisms: evidence that activation of a host TGFβ innate immune pathway promotes tolerance of the symbiont. Dev. Comp. Immunol. 38, 525–537 (2012).
Mansfield, K. M. et al. Transcription factor NF-κB is modulated by symbiotic status in a sea anemone model of cnidarian bleaching. Sci. Rep. 7, 16025 (2017).
Matthews, J. L. et al. Optimal nutrient exchange and immune responses operate in partner specificity in the cnidarian–dinoflagellate symbiosis. Proc. Natl Acad. Sci. USA 114, 13194–13199 (2017).
Wolfowicz, I. et al. Aiptasia sp. larvae as a model to reveal mechanisms of symbiont selection in cnidarians. Sci. Rep. 6, 32366 (2016).
Berthelier, J. et al. Implication of the host TGFβ pathway in the onset of symbiosis between larvae of the coral Fungia scutaria and the dinoflagellate Symbiodinium sp. (clade C1f). Coral Reefs 36, 1263–1268 (2017).
Mansfield, K. M. et al. Varied effects of algal symbionts on transcription factor NF-κB in a sea anemone and a coral: possible roles in symbiosis and thermotolerance. Preprint at bioRxiv https://doi.org/10.1101/640177 (2019).
Mohamed, A. R. et al. Deciphering the nature of the coral–Chromera association. ISME J. 12, 776–790 (2018).
Chen, M.-C. et al. ApRab11, a cnidarian homologue of the recycling regulatory protein Rab11, is involved in the establishment and maintenance of the Aiptasia–Symbiodinium endosymbiosis. Biochem. Biophys. Res. Commun. 338, 1607–1616 (2005).
Chen, M.-C., Cheng, Y.-M., Sung, P.-J., Kuo, C.-E. & Fang, L.-S. Molecular identification of Rab7 (ApRab7) in Aiptasia pulchella and its exclusion from phagosomes harboring zooxanthellae. Biochem. Biophys. Res. Commun. 308, 586–595 (2003).
Davy, S. K., Allemand, D. & Weis, V. M. Cell biology of cnidarian–dinoflagellate symbiosis. Microbiol. Mol. Biol. Rev. 76, 229–261 (2012).
Mohamed, A. R. et al. The transcriptomic response of the coral Acropora digitifera to a competent Symbiodinium strain: the symbiosome as an arrested early phagosome. Mol. Ecol. 25, 3127–3141 (2016).
Voolstra, C. R. et al. The host transcriptome remains unaltered during the establishment of coral–algal symbioses. Mol. Ecol. 18, 1823–1833 (2009).
Dunn, S. R. & Weis, V. M. Apoptosis as a post-phagocytic winnowing mechanism in a coral–dinoflagellate mutualism. Environ. Microbiol. 11, 268–276 (2009).
Baumgarten, S. et al. The genome of Aiptasia, a sea anemone model for coral symbiosis. Proc. Natl Acad. Sci. USA 112, 11893–11898 (2015).
Grawunder, D. et al. Induction of gametogenesis in the cnidarian endosymbiosis model Aiptasia sp. Sci. Rep. 5, 15677 (2015).
Hambleton, E. A. et al. Sterol transfer by atypical cholesterol-binding NPC2 proteins in coral–algal symbiosis. eLife 8, e43923 (2019).
Bucher, M., Wolfowicz, I., Voss, P. A., Hambleton, E. A. & Guse, A. Development and symbiosis establishment in the cnidarian endosymbiosis model Aiptasia sp. Sci. Rep. 6, 19867 (2016).
Hambleton, E. A., Guse, A. & Pringle, J. R. Similar specificities of symbiont uptake by adults and larvae in an anemone model system for coral biology. J. Exp. Biol. 217, 1613–1619 (2014).
LaJeunesse, T. C. et al. Systematic revision of Symbiodiniaceae highlights the antiquity and diversity of coral endosymbionts. Curr. Biol. 28, 2570–2580 (2018).
Xiang, T., Hambleton, E. A., DeNofrio, J. C., Pringle, J. R. & Grossman, A. R. Isolation of clonal axenic strains of the symbiotic dinoflagellate Symbiodinium and their growth and host specificity. J. Phycol. 49, 447–458 (2013).
Cumbo, V. R. et al. Chromera velia is endosymbiotic in larvae of the reef corals Acropora digitifera and A. tenuis. Protist 164, 237–244 (2013).
Budiša, A. et al. Marine microagae Microchoropsis gaditana and Pseudochoris wilhelmii cultivated in oil refinery wastewater—a perspective on remediation and biodiesel production. Fresenius Environ. Bull. 28, 7888–7897 (2019).
Ma, X.-N., Chen, T.-P., Yang, B., Liu, J. & Chen, F. Lipid production from Nannochloropsis. Mar. Drugs 14, 61 (2016).
Boulais, J. et al. Molecular characterization of the evolution of phagosomes. Mol. Syst. Biol. 6, 423 (2010).
Flannagan, R. S., Jaumouillé, V. & Grinstein, S. The cell biology of phagocytosis. Annu. Rev. Pathol. 7, 61–98 (2012).
Luzio, J. P., Hackmann, Y., Dieckmann, N. M. G. & Griffiths, G. M. The biogenesis of lysosomes and lysosome-related organelles. Cold Spring Harb. Perspect. Biol. 6, a016840 (2014).
Bojarczuk, A. et al. Cryptococcus neoformans intracellular proliferation and capsule size determines early macrophage control of infection. Sci. Rep. 6, 21489 (2016).
Smith, L. M. & May, R. C. Mechanisms of microbial escape from phagocyte killing. Biochem. Soc. Trans. 41, 475–490 (2013).
Seoane, P. I. & May, R. C. Vomocytosis: what we know so far. Cell. Microbiol. 22, e13145 (2020).
Watkins, R. A. et al. Cryptococcus neoformans escape from Dictyostelium amoeba by both WASH-mediated constitutive exocytosis and vomocytosis. Front. Cell. Infect. Microbiol. 8, 108 (2018).
Ma, H., Croudace, J. E., Lammas, D. A. & May, R. C. Expulsion of live pathogenic yeast by macrophages. Curr. Biol. 16, 2156–2160 (2006).
Johnston, S. A. & May, R. C. The human fungal pathogen Cryptococcus neoformans escapes macrophages by a phagosome emptying mechanism that is inhibited by Arp2/3 complex-mediated actin polymerisation. PLoS Pathog. 6, e1001041 (2010).
Gilbert, A. S. et al. Vomocytosis of live pathogens from macrophages is regulated by the atypical MAP kinase ERK5. Sci. Adv. 3, e1700898 (2017).
Gazzinelli, R. T., Mendonça-Neto, R., Lilue, J., Howard, J. & Sher, A. Innate resistance against Toxoplasma gondii: an evolutionary tale of mice, cats, and men. Cell Host Microbe 15, 132–138 (2014).
Ghosh, D. & Stumhofer, J. S. Do you see what I see: recognition of protozoan parasites by Toll-like receptors. Curr. Immunol. Rev. 9, 129–140 (2013).
Kanehisa, M. & Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27–30 (2000).
Botos, I., Segal, D. M. & Davies, D. R. The structural biology of Toll-like receptors. Structure 19, 447–459 (2011).
Bosch, T. C. G. et al. Uncovering the evolutionary history of innate immunity: the simple metazoan Hydra uses epithelial cells for host defence. Dev. Comp. Immunol. 33, 559–569 (2009).
Loiarro, M. et al. Pivotal advance: inhibition of MyD88 dimerization and recruitment of IRAK1 and IRAK4 by a novel peptidomimetic compound. J. Leukoc. Biol. 82, 801–810 (2007).
Biquand, E. et al. Acceptable symbiont cell size differs among cnidarian species and may limit symbiont diversity. ISME J. 11, 1702–1712 (2017).
Neubauer, E. F., Poole, A. Z., Weis, V. M. & Davy, S. K. The scavenger receptor repertoire in six cnidarian species and its putative role in cnidarian–dinoflagellate symbiosis. PeerJ 4, e2692 (2016).
Neubauer, E.-F. et al. A diverse host thrombospondin-type-1 repeat protein repertoire promotes symbiont colonization during establishment of cnidarian–dinoflagellate symbiosis. eLife 6, 961 (2017).
Schwarz, J. A. et al. Coral life history and symbiosis: functional genomic resources for two reef building Caribbean corals, Acropora palmata and Montastraea faveolata. BMC Genom. 9, 97 (2008).
Weis, V. M., Davy, S. K., Hoegh-Guldberg, O., Rodriguez-Lanetty, M. & Pringle, J. R. Cell biology in model systems as the key to understanding corals. Trends Ecol. Evol. 23, 369–376 (2008).
Ndungu, F. M., Urban, B. C., Marsh, K. & Langhorne, J. Regulation of immune response by Plasmodium-infected red blood cells. Parasite Immunol. 27, 373–384 (2005).
Waghabi, M. C., Keramidas, M., Feige, J.-J., Araujo-Jorge, T. C. & Bailly, S. Activation of transforming growth factor β by Trypanosoma cruzi. Cell. Microbiol. 7, 511–517 (2005).
Alvarez, M. & Casadevall, A. Phagosome extrusion and host-cell survival after Cryptococcus neoformans phagocytosis by macrophages. Curr. Biol. 16, 2161–2165 (2006).
Birmingham, C. L. et al. Listeriolysin O allows Listeria monocytogenes replication in macrophage vacuoles. Nature 451, 350–354 (2008).
Levitz, S. M. et al. Cryptococcus neoformans resides in an acidic phagolysosome of human macrophages. Infecct. Immun. 67, 885–890 (1999).
Liss, V. et al. Salmonella enterica remodels the host cell endosomal system for efficient intravacuolar nutrition. Cell Host Microbe 21, 390–402 (2017).
Madan, R., Rastogi, R., Parashuraman, S. & Mukhopadhyay, A. Salmonella acquires lysosome-associated membrane protein 1 (LAMP1) on phagosomes from Golgi via SipC protein-mediated recruitment of host Syntaxin6. J. Biol. Chem. 287, 5574–5587 (2012).
Hartenstein, V. & Martinez, P. Phagocytosis in cellular defense and nutrition: a food-centered approach to the evolution of macrophages. Cell Tissue Res. 377, 527–547 (2019).
Downs, C. A. et al. Symbiophagy as a cellular mechanism for coral bleaching. Autophagy 5, 211–216 (2009).
Titlyanov, E. et al. Degradation of zooxanthellae and regulation of their density in hermatypic corals. Mar. Ecol. Prog. Ser. 139, 167–178 (1996).
Chen, M.-C., Cheng, Y.-M., Hong, M.-C. & Fang, L.-S. Molecular cloning of Rab5 (ApRab5) in Aiptasia pulchella and its retention in phagosomes harboring live zooxanthellae. Biochem. Biophys. Res. Commun. 324, 1024–1033 (2004).
McCloskey, L. R., Cove, T. G. & Verde, E. A. Symbiont expulsion from the anemone Anthopleura elegantissima (Brandt) (Cnidaria; Anthozoa). J. Exp. Mar. Biol. Ecol. 195, 173–186 (1996).
Hoegh-Guldberg, O., McCloskey, L. R. & Muscatine, L. Expulsion of zooxanthellae by symbiotic cnidarians from the Red Sea. Coral Reefs 5, 201–204 (1987).
Hoegh-Guldberg, O. & Smith, G. J. The effect of sudden changes in temperature, light and salinity on the population density and export of zooxanthellae from the reef corals Stylophora pistillata Esper and Seriatopora hystrix Dana. J. Exp. Mar. Biol. Ecol. 129, 279–303 (1989).
Bieri, T., Onishi, M., Xiang, T., Grossman, A. R. & Pringle, J. R. Relative contributions of various cellular mechanisms to loss of algae during cnidarian bleaching. PLoS ONE 11, e0152693 (2016).
Van Treuren, W. et al. Live imaging of Aiptasia larvae, a model system for coral and anemone bleaching, using a simple microfluidic device. Sci. Rep. 9, 9275 (2019).
Baghdasarian, G. & Muscatine, L. Preferential expulsion of dividing algal cells as a mechanism for regulating algal–cnidarian symbiosis. Biol. Bull. 199, 278–286 (2000).
Steele, R. The significance of zooxanthella-containing pellets extruded by sea anemones. Bull. Mar. Sci. 27, 591–594 (1977).
Mieog, J. C., van Oppen, M. J. H., Cantin, N. E., Stam, W. T. & Olsen, J. L. Real-time PCR reveals a high incidence of Symbiodinium clade D at low levels in four scleractinian corals across the Great Barrier Reef: implications for symbiont shuffling. Coral Reefs 26, 449–457 (2007).
Thornhill, D. J., LaJeunesse, T. C., Kemp, D. W., Fitt, W. K. & Schmidt, G. W. Multi-year, seasonal genotypic surveys of coral–algal symbioses reveal prevalent stability or post-bleaching reversion. Mar. Biol. 148, 711–722 (2006).
LaJeunesse, T. C. et al. Specificity and stability in high latitude eastern Pacific coral–algal symbioses. Limnol. Oceanogr. 53, 719–727 (2008).
Silverstein, R. N., Correa, A. M. S. & Baker, A. C. Specificity is rarely absolute in coral–algal symbiosis: implications for coral response to climate change. Proc. R. Soc. B Biol. Sci. 279, 2609–2618 (2012).
Chrisman, C. J., Alvarez, M. & Casadevall, A. Phagocytosis of Cryptococcus neoformans by, and nonlytic exocytosis from, Acanthamoeba castellanii. Appl. Environ. Microbiol. 76, 6056–6062 (2010).
Hagedorn, M., Rohde, K. H., Russell, D. G. & Soldati, T. Infection by tubercular mycobacteria is spread by nonlytic ejection from their amoeba hosts. Science 323, 1729–1733 (2009).
Miao, Y., Wu, J. & Abraham, S. N. Ubiquitination of innate immune regulator TRAF3 orchestrates expulsion of intracellular bacteria by exocyst complex. Immunity 45, 94–105 (2016).
Song, J. et al. TLR4-mediated expulsion of bacteria from infected bladder epithelial cells. Proc. Natl Acad. Sci. USA 106, 14966–14971 (2009).
Smith, L. M., Dixon, E. F. & May, R. C. The fungal pathogen Cryptococcus neoformans manipulates macrophage phagosome maturation. Cell. Microbiol. 17, 702–713 (2015).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-Seq2. Nat. Protoc. 9, 171–181 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
R Core Development Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2018).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Berthold, M. R. et al. in Data Analysis, Machine Learning and Applications. Studies in Classification, Data Analysis, and Knowledge Organization (eds Preisach, C. et al.) 319–326 (Springer, 2008).
Luo, W. & Brouwer, C. Pathview: an R/Bioconductor package for pathway-based data integration and visualization. Bioinformatics 29, 1830–1831 (2013).
Capella-Gutierrez, S., Silla-Martinez, J. M. & Gabaldon, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Wood, S. N. Generalized Additive Models: An Introduction with R (Chapman and Hall/CRC, 2017).
Keeling, P. J. & Burki, F. Progress towards the tree of eukaryotes. Curr. Biol. 29, R808–R817 (2019).
Hardiman, G., Rock, F. L., Balasubramanian, S., Kastelein, R. A. & Bazan, J. F. Molecular characterization and modular analysis of human MyD88. Oncogene 13, 2467–2475 (1996).
Vyncke, L. et al. Reconstructing the TIR side of the Myddosome: a paradigm for TIR–TIR interactions. Structure 24, 437–447 (2016).
Loiarro, M. et al. Peptide-mediated interference of TIR domain dimerization in MyD88 inhibits interleukin-1-dependent activation of NF-κB. J. Biol. Chem. 280, 15809–15814 (2005).
Loiarro, M., Ruggiero, V. & Sette, C. Targeting TLR/IL-1R signalling in human diseases. Mediators Inflamm. 2010, 674363 (2010).
Winchester, B. G. Lysosomal membrane proteins. Eur. J. Paediatr. Neurol. 5, 11–19 (2001).
Acknowledgements
We thank D. Pavlinic and V. Benes (GeneCore Facility, EMBL Heidelberg) for assistance with the Smart-Seq2 protocol and sequencing library preparation; D. Ibberson (Deepseqlab, Heidelberg University) for assistance with the Smart-Seq2 protocol; C. Rippe for access to the bioanalyzer; L. Hambleton for help with antibody purification; B. G. Bergheim for initiating live imaging of Aiptasia larvae; M. Mercker (Bionum) for advice on statistical analysis; F. Frischknecht, T. Gilmore, T. Holstein and S. Lemke for advice; and R. May for advice and comments on the manuscript. Funding was provided by the Deutsche Forschungsgemeinschaft (DFG) (Emmy Noether Program Grant GU 1128/3-1) and H2020 European Research Council (ERC Consolidator Grant 724715) to A.G., a scholarship from the CellNetworks Excellence Cluster (Heidelberg University) Postdoctoral Program to S.R. and a PhD scholarship within the graduate school Evolutionary Novelty and Adaptation by the Baden-Württemberg Landesgraduiertenförderung Program to P.A.V.
Author information
Authors and Affiliations
Contributions
M.R.J., S.R. and A.G. conceived of the study. M.R.J., S.R., P.A.V., I.M. and A.G. developed the methodology. P.A.V. and S.G.G. developed the software. M.R.J. and S.R. performed the formal analyses. M.R.J., S.R. and I.M. performed the investigation. M.R.J., S.R., P.A.V., I.M. and A.G. interpreted the data. A.G. provided the resources. P.A.V. and S.G.G. curated the data. M.R.J., S.R. and A.G. wrote the original draft of the manuscript. M.R.J., S.R., S.G.G., P.A.V. and A.G. reviewed and edited the manuscript. M.R.J. and S.R. visualized the data. A.G. supervised the project. M.R.J., S.R. and A.G. administered the project. A.G. acquired the funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Microbiology thanks Alejandro Sánchez Alvarado, Simon Davy and Christian Voolstra for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Uptake of microalgae is indiscriminate.
a, Additional microalgae screened: Isochrysis sp., Chlorella sp., D. salina, and C. parkeae. Images are DIC and red autofluorescence of microalgae photosynthetic pigments. Scale bar indicates 25 µm. b, Aiptasia larvae were infected at 4-6 days post fertilization (dpf) for 24 hours and were washed into fresh FASW. Error bars represent SEM. n=3 for all except C. parkeae with n= 1.
Extended Data Fig. 2 N-deglycosylation of LAMP1 in Aiptasia homogenate.
Verification of α-LAMP1 antibody used in Figs. 2a, 3d,e by Western blot. LAMP1 has been observed to run at a higher than predicted weight (38 kDa) because it is heavily glycosylated101. Deglycosylation of homogenates of symbiotic and aposymbiotic adult Aiptasia CC7 using PNGase F resulted in a shift to a lower molecular weight. As control, extracts were detected with LAMP1 antibody pre-adsorbed with the peptide used for raising the antibody.
Extended Data Fig. 3 Inhibition of actin polymerization does not affect expulsion of non-symbiotic microalgae.
a, Analysis of the effects of different concentrations of LatrunculinB (LatB) on Aiptasia larvae to determine a suitable concentration for live imaging analysis. Larvae were incubated for 6 hours in LatB, washed, fixed and the f-actin levels were assessed by fluorescence microscopy. 0.01 µM does not affect the overall levels or distribution of actin. In contrast, LatB concentrations >0.1 µM substantially decreased actin levels and impacted the integrity of Aiptasia larvae (see arrowheads pointing to ‘holes’ within the organisms). Accordingly, an intermediate concentration of 0.05 µM LatB which substantially reduced f-actin levels without compromising larval integrity was used for live imaging in Fig. 3a). b, Inhibition of actin polymerization with Latrunculin B did not affect the time to expulsion of M. gaditana from infected Aiptasia larvae.
Extended Data Fig. 4 ERK5 and MAP2K5 homologues in Aiptasia.
Phylogenetic analysis of ERK5 and MAP2K5 from Aiptasia. a + b are collapsed trees of Aiptasia MAPK (A) or MAP2K (B) in comparison to several other cnidarian and vertebrate species. Red arrowheads or writing indicate presence of an Aiptasia homolog. Both Aiptasia ERK5 and MEK5 cluster within ERK5 (MAPK7) or MAP2K5, respectively. Full tree can be accessed through Supplementary Files 1 and 2.
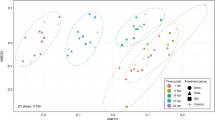
Extended Data Fig. 5 Cell-specific characterization for transcriptomic analysis.
a, Schematic of Aiptasia larvae used for cell-specific sequencing. Ectodermal cells were removed resulting in only endodermal cells that were dissociated and selected for based on contents: aposymbiotic cells from symbiotic larvae (Symbiont-Apo), symbiotic cells from symbiotic larvae (Symbiont (red)), aposymbiotic cells from aposymbiotic larvae (Apo), cells containing M. gaditana from larvae infected with M. gaditana (M. gaditana (yellow)), and aposymbiotic cells from larvae infected with M. gaditana (M. gaditana-Apo). b, Principal Component Analysis (PCA) plot of host gene expression in different conditions.
Extended Data Fig. 6 Amino acid sequence similarity between human and Aiptasia MyD88.
Human MyD88 homo-dimerizes to trigger a downstream signaling cascade leading to immune activation. It consists of three domains, the death domain (DD), the interdomain (ID) and the C-terminal TIR domain97. The human TIR domain is key for homo-dimerization with other TIR domains from MyD88 or other TIR domain containing proteins. Three distinct regions contributing to homo-dimerization have been identified by crystallography, NMR and mammalian two-hybrid analysis98. However, the so-called BB-loop within the TIR domain, a solvent-exposed stretch of 7 residues (RDLVPGT) is particularly critical for homodimerization in human MyD88. Accordingly, cell-permeable peptides mimicking the 7 residues of the BB-loop of human MyD88 interfere with homo-dimerization55,99,100. The TIR domains (black box/upper alignment) of mammals and Aiptasia are well conserved (∼50% sequence identity). Moreover, the BB-loop (red box) is almost identical and key residues (*) are conserved. Identical amino acids have black background, similar aa have gray background and aa with white background are not similar according to blosume62 scoring.
Supplementary information
Supplementary Data 1
Phylogenetic tree of MAPK with a focus on ERK5 (MAPK7).
Supplementary Data 2
Phylogenetic tree of MAP2K with a focus on MAP2K5.
Supplementary Video 1
Three-dimensional reconstruction of LAMP1 staining in Aiptasia larvae infected with a symbiont. LAMP1 is stained magenta, DNA is stained with Hoechst (cyan) and autofluorescence of the symbiont is shown in white.
Supplementary Video 2
Three-dimensional reconstruction of LAMP1 staining in Aiptasia larvae infected with N. oculata. LAMP1 is stained magenta, DNA is stained with Hoechst (cyan) and autofluorescence of the symbiont is shown in white.
Supplementary Video 3
Three-dimensional reconstruction of LAMP1 staining in Aiptasia larvae infected with C. velia. LAMP1 is stained magenta, DNA is stained with Hoechst (cyan) and autofluorescence of the symbiont is shown in white.
Supplementary Video 4
Intracellular/attached microalgae (symbionts) move in cohesion with the larva, whereas non-intracellular microalgae (asterisk) clearly move independently within the gastric cavity. Autofluorescence of microalgae is shown in red. The timestamp is given in hours, minutes and seconds.
Supplementary Video 5
Long-term imaging of larva infected with a symbiont. The symbiont can be seen dividing at ~8 h after the start of imaging. Autofluorescence of microalgae is shown in red. The timestamp is given in hours, minutes and seconds.
Supplementary Video 6
Z-stack of a symbiont within the gastric cavity for one time point during acquisition. Autofluorescence of microalgae is shown in red. The timestamp is given in hours, minutes and seconds.
Supplementary Video 7
Long-term imaging of larva infected with M. gaditana. M. gaditana can be seen being expelled and taken up again. Autofluorescence of microalgae is shown in red. The timestamp is given in hours, minutes and seconds.
Supplementary Video 8
Long-term imaging of larva infected with N. oculata. N. oculata can be seen being expelled and taken up again. Autofluorescence of microalgae is shown in red. The timestamp is given in hours, minutes and seconds.
Supplementary Video 9
Long-term imaging of larva infected with C. velia. C. velia can be seen being expelled and taken up again. Autofluorescence of microalgae is shown in red. The timestamp is given in hours, minutes and seconds.
Supplementary Video 10
Long-term imaging of larva infected with beads. Beads can be seen being expelled and taken up again. Autofluorescence of microalgae is shown in red. The timestamp is given in hours, minutes and seconds.
Supplementary Table 1
Live imaging statistics.
Supplementary Table 2
Statistics of transcriptional suppression of host cell immunity. Enumeration of the modulation of innate immunity genes over ten immune pathways, with some genes present in multiple pathways and multiple transcripts annotated as the same gene (for example, TRAF3).
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed western blot.
Source Data Extended Data Fig. 3
Statistical source data.
Rights and permissions
About this article
Cite this article
Jacobovitz, M.R., Rupp, S., Voss, P.A. et al. Dinoflagellate symbionts escape vomocytosis by host cell immune suppression. Nat Microbiol 6, 769–782 (2021). https://doi.org/10.1038/s41564-021-00897-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-021-00897-w
This article is cited by
-
Insights into the molecular mechanisms underlying the different heat tolerance of the scleractinian coral Pavona decussata
Coral Reefs (2024)
-
Genes possibly related to symbiosis in early life stages of Acropora tenuis inoculated with Symbiodinium microadriaticum
Communications Biology (2023)
-
Movement and storage of nematocysts across development in the nudibranch Berghia stephanieae (Valdés, 2005)
Frontiers in Zoology (2022)
-
Larval transcriptomic responses of a stony coral, Acropora tenuis, during initial contact with the native symbiont, Symbiodinium microadriaticum
Scientific Reports (2022)
-
Symbiosis maintenance in the facultative coral, Oculina arbuscula, relies on nitrogen cycling, cell cycle modulation, and immunity
Scientific Reports (2021)