Abstract
Candida auris is among the most important emerging fungal pathogens, yet mechanistic insights into its immune recognition and control are lacking. Here, we integrate transcriptional and functional immune-cell profiling to uncover innate defence mechanisms against C. auris. C. auris induces a specific transcriptome in human mononuclear cells, a stronger cytokine response compared with Candida albicans, but a lower macrophage lysis capacity. C. auris-induced innate immune activation is mediated through the recognition of C-type lectin receptors, mainly elicited by structurally unique C. auris mannoproteins. In in vivo experimental models of disseminated candidiasis, C. auris was less virulent than C. albicans. Collectively, these results demonstrate that C. auris is a strong inducer of innate host defence, and identify possible targets for adjuvant immunotherapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Requests for materials should be addressed to the corresponding author. The datasets generated in this study are accessible through GEO Series accession number GSE154911. Source data are provided with this paper.
References
Meis, J. F. & Chowdhary, A. Candida auris: a global fungal public health threat. Lancet Infect. Dis. 18, 1298–1299 (2018).
Clancy, C. J. & Nguyen, M. H. Emergence of Candida auris: an international call to arms. Clin. Infect. Dis. 64, 141–143 (2017).
de Groot, T., Puts, Y., Berrio, I., Chowdhary, A. & Meis, J. F. Development of Candida auris short tandem repeat typing and its application to a global collection of isolates. mBio 11, e02971-19 (2020).
Lockhart, S. R. et al. Simultaneous emergence of multidrug-resistant Candida auris on 3 continents confirmed by whole-genome sequencing and epidemiological analyses. Clin. Infect. Dis. 64, 134–140 (2017).
Chow, N. A. et al. Potential fifth clade of Candida auris, Iran, 2018. Emerg. Infect. Dis. 25, 1780–1781 (2019).
Welsh, R. M., Sexton, D. J., Forsberg, K., Vallabhaneni, S. & Litvintseva, A. Insights into the unique nature of the East Asian clade of the emerging pathogenic yeast Candida auris. J. Clin. Microbiol. 57, e00007–19 (2019).
Szekely, A., Borman, A. M. & Johnson, E. M. Candida auris isolates of the Southern Asian and South African lineages exhibit different phenotypic and antifungal susceptibility profiles in vitro. J. Clin. Microbiol. 57, e02055-18 (2019).
Borman, A. M., Szekely, A. & Johnson, E. M. Comparative pathogenicity of United Kingdom isolates of the emerging pathogen Candida auris and other key pathogenic Candida species. mSphere 1, e00189-16 (2016).
Kathuria, S. et al. Multidrug-resistant Candida auris misidentified as Candida haemulonii: characterization by matrix-assisted laser desorption ionization-time of flight mass spectrometry and DNA sequencing and its antifungal susceptibility profile variability by vitek 2, CLSI broth microdilution, and Etest method. J. Clin. Microbiol. 53, 1823–1830 (2015).
Mizusawa, M. et al. Can multidrug-resistant Candida auris be reliably identified in clinical microbiology laboratories? J. Clin. Microbiol. 55, 638–640 (2017).
Ruiz-Gaitan, A. et al. An outbreak due to Candida auris with prolonged colonisation and candidaemia in a tertiary care European hospital. Mycoses 61, 498–505 (2018).
Schelenz, S. et al. First hospital outbreak of the globally emerging Candida auris in a European hospital. Antimicrob. Resist. Infect. Control 5, 35 (2016).
Vallabhaneni, S. et al. Investigation of the first seven reported cases of Candida auris, a globally emerging invasive, multidrug-resistant fungus—United States, May 2013–August 2016. Am. J. Transplant. 17, 296–299 (2017).
Lee, W. G. et al. First three reported cases of nosocomial fungemia caused by Candida auris. J. Clin. Microbiol. 49, 3139–3142 (2011).
Welsh, R. M. et al. Survival, persistence, and isolation of the emerging multidrug-resistant pathogenic yeast Candida auris on a plastic health care surface. J. Clin. Microbiol. 55, 2996–3005 (2017).
Rudramurthy, S. M. et al. Candida auris candidaemia in Indian ICUs: analysis of risk factors. J. Antimicrob. Chemother. 72, 1794–1801 (2017).
Chowdhary, A., Sharma, C. & Meis, J. F. Candida auris: a rapidly emerging cause of hospital-acquired multidrug-resistant fungal infections globally. PLoS Pathog. 13, e1006290 (2017).
Al Maani, A. et al. Ongoing challenges with healthcare-associated Candida auris outbreaks in Oman. J. Fungi 5, 101 (2019).
Arendrup, M. C., Chowdhary, A., Astvad, K. M. T. & Jorgensen, K. M. APX001A in vitro activity against contemporary blood isolates and Candida auris determined by the EUCAST reference method. Antimicrob. Agents Chemother. 62, e01225-18 (2018).
Hager, C. L. et al. In vitro and in vivo evaluation of the antifungal activity of APX001A/APX001 against Candida auris. Antimicrob. Agents Chemother. 62, e02319-17 (2018).
Arendrup, M. C., Jorgensen, K. M., Hare, R. K. & Chowdhary, A. EUCAST in vitro activity of Ibrexafungerp (SCY-078) against C. auris isolates; comparison with activity against C. albicans and C. glabrata and with that of six comparators. Antimicrob. Agents Chemother. 64, e02136-19 (2019).
Berkow, E. L., Angulo, D. & Lockhart, S. R. In vitro activity of a novel glucan synthase inhibitor, SCY-078, against clinical isolates of Candida auris. Antimicrob. Agents Chemother. 61, e00435-17 (2017).
Larkin, E. et al. The emerging pathogen Candida auris: growth phenotype, virulence factors, activity of antifungals, and effect of SCY-078, a novel glucan synthesis inhibitor, on growth morphology and biofilm formation. Antimicrob. Agents Chemother. 61, e02396-16 (2017).
Hager, C. L., Larkin, E. L., Long, L. A. & Ghannoum, M. A. Evaluation of the efficacy of rezafungin, a novel echinocandin, in the treatment of disseminated Candida auris infection using an immunocompromised mouse model. J. Antimicrob. Chemother. 73, 2085–2088 (2018).
Lepak, A. J., Zhao, M. & Andes, D. R. Pharmacodynamic evaluation of rezafungin (CD101) against Candida auris in the neutropenic mouse invasive candidiasis model. Antimicrob. Agents Chemother. 62, e01572-18 (2018).
Richardson, J. P. & Moyes, D. L. Adaptive immune responses to Candida albicans infection. Virulence 6, 327–337 (2015).
Erwig, L. P. & Gow, N. A. Interactions of fungal pathogens with phagocytes. Nat. Rev. Microbiol. 14, 163–176 (2016).
Gow, N. A., van de Veerdonk, F. L., Brown, A. J. & Netea, M. G. Candida albicans morphogenesis and host defence: discriminating invasion from colonization. Nat. Rev. Microbiol. 10, 112–122 (2011).
Pathirana, R. U. et al. Fluconazole-resistant Candida auris is susceptible to salivary histatin 5 killing and to intrinsic host defenses. Antimicrob. Agents Chemother. 62, e01872-17 (2018).
Johnson, C. J., Davis, J. M., Huttenlocher, A., Kernien, J. F. & Nett, J. E. Emerging fungal pathogen Candida auris evades neutrophil attack. mBio 9, e01403-18 (2018).
Navarro-Arias, M. J. et al. Differential recognition of Candida tropicalis, Candida guilliermondii, Candida krusei, and Candida auris by human innate immune cells. Infect. Drug Resist. 12, 783–794 (2019).
Munoz, J. F. et al. Genomic insights into multidrug-resistance, mating and virulence in Candida auris and related emerging species. Nat. Commun. 9, 5346 (2018).
Brown, G. D. et al. Hidden killers: human fungal infections. Sci. Transl. Med. 4, 165rv113 (2012).
Hall, R. A. & Gow, N. A. Mannosylation in Candida albicans: role in cell wall function and immune recognition. Mol. Microbiol. 90, 1147–1161 (2013).
Gow, N. A. et al. Immune recognition of Candida albicans β-glucan by dectin-1. J. Infect. Dis. 196, 1565–1571 (2007).
Netea, M. G. et al. Immune sensing of Candida albicans requires cooperative recognition of mannans and glucans by lectin and Toll-like receptors. J. Clin. Invest. 116, 1642–1650 (2006).
Klebanoff, S. J. Myeloperoxidase: friend and foe. J. Leukoc. Biol. 77, 598–625 (2005).
Kerrigan, A. M. & Brown, G. D. Syk-coupled C-type lectin receptors that mediate cellular activation via single tyrosine based activation motifs. Immunol. Rev. 234, 335–352 (2010).
Gringhuis, S. I. et al. Dectin-1 directs T helper cell differentiation by controlling noncanonical NF-κB activation through Raf-1 and Syk. Nat. Immunol. 10, 203–213 (2009).
Yan, L. et al. Unique cell surface mannan of yeast pathogen Candida auris with selective binding to IgG. ACS Infect. Dis. 6, 1018–1031 (2020).
Marakalala, M. J. et al. Differential adaptation of Candida albicans in vivo modulates immune recognition by dectin-1. PLoS Pathog. 9, e1003315 (2013).
Keppler-Ross, S., Douglas, L., Konopka, J. B. & Dean, N. Recognition of yeast by murine macrophages requires mannan but not glucan. Eukaryot. Cell 9, 1776–1787 (2010).
McKenzie, C. G. J. et al. Contribution of Candida albicans cell wall components to recognition by and escape from murine macrophages. Infect. Immun. 78, 1650–1658 (2010).
Tucey, T. M. et al. Glucose homeostasis is important for immune cell viability during Candida challenge and host survival of systemic fungal infection. Cell Metab. 27, 988–1006 (2018).
Fakhim, H. et al. Comparative virulence of Candida auris with Candida haemulonii, Candida glabrata and Candida albicans in a murine model. Mycoses 61, 377–382 (2018).
Ben-Ami, R. et al. Multidrug-resistant Candida haemulonii and C. auris, Tel Aviv, Israel. Emerg. Infect. Dis. 23, 195–203 (2017).
Urban, C. F. & Nett, J. E. Neutrophil extracellular traps in fungal infection. Semin. Cell Dev. Biol. 89, 47–57 (2019).
Urban, C. F., Reichard, U., Brinkmann, V. & Zychlinsky, A. Neutrophil extracellular traps capture and kill Candida albicans yeast and hyphal forms. Cell Microbiol. 8, 668–676 (2006).
Wiederhold, N. P. et al. Efficacy of delayed therapy with fosmanogepix (APX001) in a murine model of Candida auris invasive candidiasis. Antimicrob. Agents Chemother. 63, e01120-19 (2019).
Oosting, M. et al. Borrelia-induced cytokine production is mediated by spleen tyrosine kinase (Syk) but is dectin-1 and dectin-2 independent. Cytokine 76, 465–472 (2015).
Gaus, H., Miller, C. M., Seth, P. P. & Harris, E. N. Structural determinants for the interactions of chemically modified nucleic acids with the stabilin-2 clearance receptor. Biochemistry 57, 2061–2064 (2018).
Sherrington, S. L. et al. Adaptation of Candida albicans to environmental pH induces cell wall remodelling and enhances innate immune recognition. PLoS Pathog. 13, e1006403 (2017).
Kruppa, M., Greene, R. R., Noss, I., Lowman, D. W. & Williams, D. L. C. albicans increases cell wall mannoprotein, but not mannan, in response to blood, serum and cultivation at physiological temperature. Glycobiology 21, 1173–1180 (2011).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq-a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Zhu, A., Ibrahim, J. G. & Love, M. I. Heavy-tailed prior distributions for sequence count data: removing the noise and preserving large differences. Bioinformatics 35, 2084–2092 (2018).
Kamburov, A., Wierling, C., Lehrach, H. & Herwig, R. ConsensusPathDB—a database for integrating human functional interaction networks. Nucleic Acids Res. 37, D623–D628 (2009).
Kanehisa, M. & Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27–30 (2000).
Fabregat, A. et al. The reactome pathway knowledgebase. Nucleic Acids Res. 46, D649–D655 (2018).
Lowman, D. W. et al. Mannan structural complexity is decreased when Candida albicans is cultivated in blood or serum at physiological temperature. Carbohydr. Res. 346, 2752–2759 (2011).
Graus, M. S. et al. Mannan molecular substructures control nanoscale glucan exposure in Candida. Cell Rep. 24, e2435 (2018).
Smith, A. J. et al. Immunoregulatory activity of the natural product laminarin varies widely as a result of its physical properties. J. Immunol. 200, 788–799 (2018).
Acknowledgements
We thank T. Jansen for performing the initial pilot experiments and I. Curfs-Breuker and D. Faro for their support at the CWZ hospital; C. Kaffa for her technical assistance; V. Kumar for his input during the transcriptomic data analysis; M. Gresnigt for in vitro experimental suggestions; and A. Becker for help during the revision experiments. A.J.P.B. and N.A.R.G. thank the Medical Research Council (MRC) (MR/M026663/1) and Wellcome for support and the MRC Centre for Medical Mycology at the University of Aberdeen (MR/N006364/1). A.H. and S.K. were supported by the Radboud Institute for Molecular Life Sciences. Part of the study was supported by the Hellenic Institute for the Study of Sepsis. D.L.W. was supported by National Institutes of Health grant nos NIH GM083016, GM119197 and C06RR0306551. M.G.N. was supported by an ERC Advanced Grant (no. 833247) and a Spinoza Grant from the Netherlands Organization for Scientific Research.
Author information
Authors and Affiliations
Contributions
Conceptualization: J.F.M., D.L.W. and M.G.N.; methodology: M.B., S.K., J.M.B., M.J., D.R., M.D.K., D.W.L., Z.M., Y.N.J., A.C., A.H., N.A.R.G., A.J.P.B., J.F.M., D.L.W. and M.G.N.; investigation: M.B., S.K., J.M.B., M.J, D.R., D.W.L., P.J.R., B.G., F.L.v.d.V., B.-J.K., E.J.G.-B., G.R., A.H., N.A.R.G., A.J.P.B., D.L.W. and M.G.N.; writing the original draft: M.B., S.K. and M.G.N.; review and editing the manuscript: M.B., S.K., J.M.B., M.J., M.D.K., D.W.L., Z.M., Y.N.J., A.C., F.L.v.d.V., E.J.G.-B., A.H., N.A.R.G., A.J.P.B., J.F.M., D.L.W. and M.G.N.; supervision: M.J., F.L.v.d.V., A.H., N.A.R.G., A.J.P.B. and M.G.N.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Transcriptomic profiling PBMCs stimulated with live C. albicans or C. auris and respective cell wall components β-glucans and mannans for 4 and 24 hours.
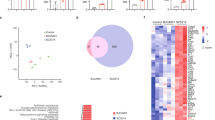
a, Principal component analysis (PCA) performed on normalized count data (normTransform, DESeq2) demonstrates the main component introducing variance in the dataset is time (40%), as indicated by a clear split between the early 4-hour host induced response (left, triangle) and the late 24-hour response (right, circle). To a lower extent (15%), the second component introducing variance appears to be inherent to the stimulus (color). b, At 4 h, PCA reveals a clear donor clustering (shape) irrespective of stimulus (color), indicating the main variance in the early host response reflects inter-individual differences (left). PCA on the late response, 24 h, is predominantly influenced by the respective stimulus (38%, color), and to a lower extent by the donors (19%, shape), indicated by the scattering of stimuli together with a rough clustering amongst donors. c, Pathway enrichment plot displaying the top 20 enriched pathways for both C. albicans live and C. auris live (color) at 24 h. Enrichment determined using Consensus PathDB, including pathways as defined by KEGG and Reactome (shape), considering a p-adjusted value < 0.01 (indicated as ‘q-value’) significant. Size of the geometric points indicates the amount of DEG in relation to the pathways’ size. The exact q values and the data used to make this figure can be found in Source Data Extended Data Fig. 1.
Extended Data Fig. 2 Comparative LDH secretion, LDH cytokine gene expression and phagocytosis dynamics between C. albicans and C. auris.
a, Assessment of Candida induced cell death of PBMCs after 24 h stimulation without (RPMI; negative control) or with C. albicans, several C. auris strains originating from all five geographical clades or a positive control (dead cells). Lactate dehydrogenase (LDH) was detected as measure of cell death (Mean ± SEM, n = 6, pooled from two independent experiments). b, Log2Fold Change (Log2FC) of IL-6, IL-1β, and IL-1RN (encoding for IL-1Ra) gene expression in PBMCs from 3 donors stimulated for 24 h with C. albicans (1006110) and C. auris (KCTC17810, clade II) and their respective cell wall components, β-glucans (left) and mannans (right). Graphs represent Log2FC from DEG analysis. * p < 0.05, ** p < 0.01, *** p < 0.001, 1-way ANOVA with correction for multiple comparison. c, The BMDM phagocytic capacity of Thimerosal-fixed C. albicans or C. auris strains in the course of 3-hours. BMDM engulfment depicted as the percentage of macrophages having phagocytosed at least one fungal cell (left), and the phagocytic index, here considered as the number of fungal cells engulfed per 100 macrophages (right); graphs represent mean, n = 9, pooled from at least two independent experiments. d, BMDM phagocytic capacity of Thimerosal-fixed C. albicans or C. auris strains after 1 h. Engulfment is depicted as the percentage of macrophages having phagocytosed at least one fungal cell; graphs represent mean ± SEM, n = 9, pooled from at least two independent experiments. e, BMDM phagocytic capacity of live C. albicans or C. auris strains after 1 h. Engulfment is depicted as the percentage of macrophages having phagocytosed at least one fungal cell. Graphs represent mean ± SEM, n = 9 (n = 7 for C. auris 10051893), pooled from at least two independent experiments. * p < 0.05, ** p < 0.01, *** p < 0.001, d 1-way ANOVA with a Holm-Sidak’s multiple comparison test, e Kruskal Wallis test with two-sided Dunn’s multiple comparison. f, Distribution of phagocytosed Thimerosal-fixed fungal cells per macrophage in a 3-hour period, n ≥ 100 observations per condition. Data used to make this figure can be found in Source Data Extended Data Fig. 2.
Extended Data Fig. 3 Relative C. auris induced ROS production and heat-sensitivity of the cell wall components responsible for the C. auris induced cytokine production.
a, Neutrophil ROS release after 1-hour stimulation without (RPMI; negative control) or with heat-killed C. albicans, C. auris strains or zymosan (positive control), depicted in relative light units (RLU) either as time-course (left) or as area under the curve (AUC, right), n = 9. b, PBMC ROS release after 1-hour stimulation without (RPMI; negative control) or with heat-killed C. albicans, C. auris strains or zymosan (positive control), depicted in RLU either as time-course (left) or as AUC (right), n = 6. c, TNF-α, IL-6, IL-1β, and IL-1Ra levels in the supernatant of PBMCs after stimulation without (RPMI; negative control) or with heat-killed C. albicans and C. auris from all five geographical clades for 24 h, n = 8. d, PBMC production of cytokines IFN-γ (n = 10; n = 7 for C. auris 10051895), IL-10 (n = 6), IL-17 (n = 6), and IL-22 (n = 14; n = 6 for C. auris 10051893; n = 11 for C. auris 10051895) after stimulation without (RPMI; negative control) or with heat-killed C. albicans and C. auris for 7 days. Graphs represent mean ± SEM, data are pooled from at least two independent experiments. * p < 0.05, ** p < 0.01, *** p < 0.001, **** p = 0.001, a, b Time curves (left panels) were assessed for statistical differences between C. auris strains and C. albicans by a two-way ANOVA, Area Under curve (AUC) means (right panels) were compared using the two-sided Wilcoxon signed rank test, c, d two-sided Wilcoxon matched pairs signed-rank test comparing respective C. auris strains with C. albicans as control or reference species. Data used to make this figure can be found in Source Data Extended Data Fig. 3.
Extended Data Fig. 4 Transcriptional changes induced by purified cell wall components and their respective exposure on C. albicans and C. auris surface.
a, Heatmap displaying the Log2Fold change (color scale) of the top 50 DEG of C. albicans live, for both Candida species and their cell wall components, β-glucan and mannan, at 4 h (left panel) and 24 h (right panel). b, Flow cytometry plot based on forward scatter component (FSC) and side scatter component (SSC), demonstrating C. auris strains are slightly smaller and of higher complexity than C. albicans. c, Flow cytometry-based comparison of cell wall components of C. albicans and C. auris strains. Mean fluorescent intensity (MFI) of thimerosal-fixed Candida cells stained for Fc-Dectin-1, a marker for β-glucan (left), and ConA, a marker for mannans (right). Graphs represent mean ± SEM of the 3 means, each performed with three replicates in three independent measurements, * p < 0.05, Kruskall Wallis test with two-sided Dunn’s multiple comparison test was performed comparing the respective C. auris strains with the two C. albicans reference strains. Data used to make this figure can be found in Source Data Extended Data Fig. 4.
Extended Data Fig. 5 Evaluation of cytokine production upon C. albicans and C. auris mannan stimulation.
a, PBMC production of cytokines TNF-α, IL-6, IL-1β, and IL-1Ra after 24 h stimulation without (RPMI; negative control) or with purified mannans from C. albicans and C. auris strains in the presence of 10% heat-inactivated human serum, n = 7. b, PBMC production of cytokines IFN-γ (n = 6), IL-17 (n = 9), and IL-22 (n = 9) after 7 days hours stimulation without (RPMI; negative control) or with purified mannans from C. albicans and C. auris strains in the presence of 10% human serum. Graphs represent mean ± SEM, data pooled from at least two independent experiments. * p < 0.05, two-sided Wilcoxon matched pairs signed-rank test, comparing respective C. auris strains with C. albicans as control or reference species. Data used to make this figure can be found in Source Data Extended Data Fig. 5.
Extended Data Fig. 6 Cytokine levels in plasma and organ homogenates from C.albicans and C. auris-infected mice.
a, IL-6 production in plasma and supernatants from liver homogenates. b, IFN-γ production in supernatants from kidney and spleen homogenates. c–e, IL-1β (c), IL-17 (d), and IL-10 (e) production in plasma and supernatants from liver, kidney, and spleen homogenates. Mice have been infected i.v. with 1×106 c.f.u. of C. albicans or C. auris. Graphs represent mean ± SEM, n = 6 per group per time-point pooled from two independent experiments. Data used to make this figure can be found in Source Data Extended Data Fig. 6.
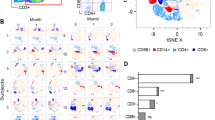
Extended Data Fig. 7 Applied gating strategies across flow cytometry experiments.
a, Gating strategy for FITC-labelled Candida in PBMCs (linked to Fig. 2b). All events were plotted based on forward scatter (FS) and side scatter (SS) characteristics. In the upper plot (2.1) the region of cells positive for FITC-Candida was highlighted (green gate) while in the bottom plot (2.2) CD14 positive cells are represented (red gate) gated within the total PBMCs population (1). Within the CD14 + cells selection, the amount of phagocytosed FITC positive Candida was examined by plotting (3) the FITC signal against the CD14-PB450 signal (blue gate) and the percentage of cells and mean fluorescent intensity (MFI) were used for analysis. b, Gating strategy for Thimerosal-fixed Candida cells stained for either β-glucan using Fc-Dectin-1 or ConA as marker for mannans (Extended Data Fig. 4c).
Supplementary information
Supplementary Information
Supplementary Tables 1–8.
Supplementary Video 1
C. auris is able to multiply within phagosomes.
Supplementary Video 2
C. auris accumulates in high numbers within macrophages and does not induce macrophage lysis.
Supplementary Video 3
C. auriscells are taken up extensively into a subpopulation of macrophages.
Supplementary Video 4
Phagocytosis of C. albicans SC5314 and macrophage lysis after 3 h.
Source data
Source Data Fig. 1
Supporting data for Fig. 1.
Source Data Fig. 2
Supporting data for Fig. 2.
Source Data Fig. 3
Supporting data for Fig. 3.
Source Data Fig. 5
Supporting data for Fig. 5.
Source Data Fig. 6
Supporting data for Fig. 6.
Source Data Extended Data Fig. 1
Supporting data for Extended Data Fig. 1.
Source Data Extended Data Fig. 2
Supporting data for Extended Data Fig. 2.
Source Data Extended Data Fig. 3
Supporting data for Extended Data Fig. 3.
Source Data Extended Data Fig. 4
Supporting data for Extended Data Fig. 4.
Source Data Extended Data Fig. 5
Supporting data for Extended Data Fig. 5.
Source Data Extended Data Fig. 6
Supporting data for Extended Data Fig. 6.
Rights and permissions
About this article
Cite this article
Bruno, M., Kersten, S., Bain, J.M. et al. Transcriptional and functional insights into the host immune response against the emerging fungal pathogen Candida auris. Nat Microbiol 5, 1516–1531 (2020). https://doi.org/10.1038/s41564-020-0780-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-020-0780-3
This article is cited by
-
Immune responses to human fungal pathogens and therapeutic prospects
Nature Reviews Immunology (2023)
-
Innate immune responses against the fungal pathogen Candida auris
Nature Communications (2022)
-
Root system architecture in rice: impacts of genes, phytohormones and root microbiota
3 Biotech (2022)
-
Dissection of the anti-Candida albicans mannan immune response using synthetic oligomannosides reveals unique properties of β-1,2 mannotriose protective epitopes
Scientific Reports (2021)