Abstract
Cell identity in eukaryotes is controlled by transcriptional regulatory networks that define cell-type-specific gene expression. In the opportunistic fungal pathogen Candida albicans, transcriptional regulatory networks regulate epigenetic switching between two alternative cell states, ‘white’ and ‘opaque’, that exhibit distinct host interactions. In the present study, we reveal that the transcription factors (TFs) regulating cell identity contain prion-like domains (PrLDs) that enable liquid–liquid demixing and the formation of phase-separated condensates. Multiple white–opaque TFs can co-assemble into complex condensates as observed on single DNA molecules. Moreover, heterotypic interactions between PrLDs support the assembly of multifactorial condensates at a synthetic locus within live eukaryotic cells. Mutation of the Wor1 TF revealed that substitution of acidic residues in the PrLD blocked its ability to phase separate and co-recruit other TFs in live cells, as well as its function in C. albicans cell fate determination. Together, these studies reveal that PrLDs support the assembly of TF complexes that control fungal cell identity and highlight parallels with the ‘super-enhancers’ that regulate mammalian cell fate.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Data that support the findings of this study are available from the corresponding author upon request. Source data are provided with this paper.
References
Wilkinson, A. C., Nakauchi, H. & Gottgens, B. Mammalian transcription factor networks: recent advances in interrogating biological complexity. Cell Syst. 5, 319–331 (2017).
Moris, N., Pina, C. & Arias, A. M. Transition states and cell fate decisions in epigenetic landscapes. Nat. Rev. Genet. 17, 693–703 (2016).
Sabari, B. R. et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science 361, eaar3958 (2018).
Plys, A. J. & Kingston, R. E. Dynamic condensates activate transcription. Science 361, 329–330 (2018).
Patel, A. et al. A liquid-to-solid phase transition of the ALS protein FUS accelerated by disease mutation. Cell 162, 1066–1077 (2015).
Kato, M. et al. Cell-free formation of RNA granules: low complexity sequence domains form dynamic fibers within hydrogels. Cell 149, 753–767 (2012).
Burke, K. A., Janke, A. M., Rhine, C. L. & Fawzi, N. L. Residue-by-residue view of in vitro FUS granules that bind the C-terminal domain of RNA polymerase II. Mol. Cell 60, 231–241 (2015).
Chong, S. et al. Imaging dynamic and selective low-complexity domain interactions that control gene transcription. Science 361, eaar2555 (2018).
Hnisz, D., Shrinivas, K., Young, R. A., Chakraborty, A. K. & Sharp, P. A. A phase separation model for transcriptional control. Cell 169, 13–23 (2017).
Nair, S. J. et al. Phase separation of ligand-activated enhancers licenses cooperative chromosomal enhancer assembly. Nat. Struct. Mol. Biol. 26, 193–203 (2019).
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013).
Mansour, M. R. et al. An oncogenic super-enhancer formed through somatic mutation of a noncoding intergenic element. Science 346, 1373–1377 (2014).
Pott, S. & Lieb, J. D. What are super-enhancers? Nat. Genet. 47, 8–12 (2015).
Whyte, W. A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Parker, S. C. et al. Chromatin stretch enhancer states drive cell-specific gene regulation and harbor human disease risk variants. Proc. Natl Acad. Sci. USA 110, 17921–17926 (2013).
Niederriter, A. R., Varshney, A., Parker, S. C. & Martin, D. M. Super enhancers in cancers, complex disease, and developmental disorders. Genes 6, 1183–1200 (2015).
Loven, J. et al. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell 153, 320–334 (2013).
Varshney, A. et al. Cell specificity of human regulatory annotations and their genetic effects on gene expression. Genetics 211, 549–562 (2018).
Deitsch, K. W., Lukehart, S. A. & Stringer, J. R. Common strategies for antigenic variation by bacterial, fungal and protozoan pathogens. Nat. Rev. Microbiol. 7, 493–503 (2009).
Noble, S. M., Gianetti, B. A. & Witchley, J. N. Candida albicans cell-type switching and functional plasticity in the mammalian host. Nat. Rev. Microbiol. 15, 96–108 (2017).
Norman, T. M., Lord, N. D., Paulsson, J. & Losick, R. Stochastic switching of cell fate in microbes. Annu. Rev. Microbiol. 69, 381–403 (2015).
Ackermann, M. A functional perspective on phenotypic heterogeneity in microorganisms. Nat. Rev. Microbiol. 13, 497–508 (2015).
Slutsky, B. et al. ‘White–opaque transition’: a second high-frequency switching system in Candida albicans. J. Bacteriol. 169, 189–197 (1987).
Kvaal, C. et al. Misexpression of the opaque-phase-specific gene PEP1 (SAP1) in the white phase of Candida albicans confers increased virulence in a mouse model of cutaneous infection. Infect. Immun. 67, 6652–6662 (1999).
Kvaal, C. A., Srikantha, T. & Soll, D. R. Misexpression of the white-phase-specific gene WH11 in the opaque phase of Candida albicans affects switching and virulence. Infect. Immun. 65, 4468–4475 (1997).
Mallick, E. M. et al. Phenotypic plasticity regulates Candida albicans interactions and virulence in the vertebrate host. Front. Microbiol. 7, 780 (2016).
Hernday, A. D. et al. Structure of the transcriptional network controlling white–opaque switching in Candida albicans. Mol. Microbiol. 90, 22–35 (2013).
Hernday, A. D. et al. Ssn6 defines a new level of regulation of white–opaque switching in Candida albicans and is required for the stochasticity of the switch. mBio 7, e01565–15 (2016).
Huang, G. et al. Bistable expression of WOR1, a master regulator of white–opaque switching in Candida albicans. Proc. Natl Acad. Sci. USA 103, 12813–12818 (2006).
Lohse, M. B. et al. Identification and characterization of a previously undescribed family of sequence-specific DNA-binding domains. Proc. Natl Acad. Sci. USA 110, 7660–7665 (2013).
Lohse, M. B. & Johnson, A. D. Identification and characterization of Wor4, a new transcriptional regulator of white-opaque switching. G3 (Bethesda). 6, 721–729 (2016).
Srikantha, T. et al. TOS9 regulates white-opaque switching in Candida albicans. Eukaryot. Cell 5, 1674–1687 (2006).
Srikantha, T., Tsai, L. K., Daniels, K. & Soll, D. R. EFG1 null mutants of Candida albicans switch but cannot express the complete phenotype of white-phase budding cells. J. Bacteriol. 182, 1580–1591 (2000).
Wang, H. et al. Candida albicans Zcf37, a zinc finger protein, is required for stabilization of the white state. FEBS Lett. 585, 797–802 (2011).
Zordan, R. E., Galgoczy, D. J. & Johnson, A. D. Epigenetic properties of white-opaque switching in Candida albicans are based on a self-sustaining transcriptional feedback loop. Proc. Natl Acad. Sci. USA 103, 12807–12812 (2006).
Zordan, R. E., Miller, M. G., Galgoczy, D. J., Tuch, B. B. & Johnson, A. D. Interlocking transcriptional feedback loops control white-opaque switching in Candida albicans. PLoS Biol. 5, e256 (2007).
Frazer, C., Hernday, A. D. & Bennett, R. J. Monitoring phenotypic switching in Candida albicans and the use of next-gen fluorescence reporters. Curr. Protoc. Microbiol. 53, e76 (2019).
Morrow, B., Srikantha, T., Anderson, J. & Soll, D. R. Coordinate regulation of two opaque-phase-specific genes during white-opaque switching in Candida albicans. Infect. Immun. 61, 1823–1828 (1993).
Srikantha, T. & Soll, D. R. A white-specific gene in the white–opaque switching system of Candida albicans. Gene 131, 53–60 (1993).
Jenull, S. et al. The Candida albicans HIR histone chaperone regulates the yeast-to-hyphae transition by controlling the sensitivity to morphogenesis signals. Sci. Rep. 7, 8308 (2017).
Lancaster, A. K., Nutter-Upham, A., Lindquist, S. & King, O. D. PLAAC: a web and command-line application to identify proteins with prion-like amino acid composition. Bioinformatics 30, 2501–2502 (2014).
Franzmann, T. & Alberti, S. Prion-like low-complexity sequences: key regulators of protein solubility and phase behavior. J. Biol. Chem. 294, 7128–7136 (2018).
Wang, J. et al. A molecular grammar governing the driving forces for phase separation of prion-like RNA binding proteins. Cell 174, 688–699 (2018).
Ribbeck, K. & Gorlich, D. The permeability barrier of nuclear pore complexes appears to operate via hydrophobic exclusion. EMBO J. 21, 2664–2671 (2002).
Kroschwald, S., Maharana, S. & Simon, A. Hexanediol: a chemical probe to investigate the material properties of membrane-less compartments. Matters 3, e201702000010 (2017).
Kato, M. & McKnight, S. L. A solid-state conceptualization of information transfer from gene to message to protein. Annu. Rev. Biochem. 87, 351–390 (2018).
Doedt, T. et al. APSES proteins regulate morphogenesis and metabolism in Candida albicans. Mol. Biol. Cell. 15, 3167–3180 (2004).
Zhao, Y. et al. The APSES family proteins in fungi: characterizations, evolution and functions. Fungal Genet. Biol. 81, 271–280 (2015).
Larson, A. G. et al. Liquid droplet formation by HP1α suggests a role for phase separation in heterochromatin. Nature 547, 236–240 (2017).
Soniat, M. M. et al. Next-generation DNA curtains for single-molecule studies of homologous recombination. Methods Enzymol. 592, 259–281 (2017).
Brown, M. W. et al. Dynamic DNA binding licenses a repair factor to bypass roadblocks in search of DNA lesions. Nat. Commun. 7, 10607 (2016).
Myler, L. R. et al. Single-molecule imaging reveals the mechanism of Exo1 regulation by single-stranded DNA binding proteins. Proc. Natl Acad. Sci. USA 113, E1170–E1179 (2016).
Hyman, A. A., Weber, C. A. & Julicher, F. Liquid–liquid phase separation in biology. Annu. Rev. Cell Dev. Biol. 30, 39–58 (2014).
Greig, J. A. et al. Arginine-enriched mixed-charge domains provide cohesion for nuclear speckle condensation. Mol. Cell 77, 1237–1250 (2020).
Nott, T. J. et al. Phase transition of a disordered nuage protein generates environmentally responsive membraneless organelles. Mol. Cell 57, 936–947 (2015).
Pak, C. W. et al. Sequence determinants of intracellular phase separation by complex coacervation of a disordered protein. Mol. Cell 63, 72–85 (2016).
Murthy, A. C. et al. Molecular interactions underlying liquid–liquid phase separation of the FUS low-complexity domain. Nat. Struct. Mol. Biol. 26, 637–648 (2019).
Boija, A. et al. Transcription factors activate genes through the phase-separation capacity of their activation domains. Cell 175, 1842–1855 (2018).
Janicki, S. M. et al. From silencing to gene expression: real-time analysis in single cells. Cell 116, 683–698 (2004).
Owen, I. & Shewmaker, F. The role of post-translational modifications in the phase transitions of intrinsically disordered proteins. Int. J. Mol. Sci. 20, 5501 (2019).
Alby, K. & Bennett, R. J. Stress-induced phenotypic switching in Candida albicans. Mol. Biol. Cell 20, 3178–3191 (2009).
Shrinivas, K. et al. Enhancer features that drive formation of transcriptional condensates. Mol. Cell 75, 549–561 (2019).
Boehning, M. et al. RNA polymerase II clustering through carboxy-terminal domain phase separation. Nat. Struct. Mol. Biol. 25, 833–840 (2018).
Kwon, I. et al. Phosphorylation-regulated binding of RNA polymerase II to fibrous polymers of low-complexity domains. Cell 155, 1049–1060 (2013).
McSwiggen, D. T., Mir, M., Darzacq, X. & Tjian, R. Evaluating phase separation in live cells: diagnosis, caveats, and functional consequences. Genes Dev. 33, 1619–1634 (2019).
Nobile, C. J. et al. A recently evolved transcriptional network controls biofilm development in Candida albicans. Cell 148, 126–138 (2012).
Fox, E. P. et al. An expanded regulatory network temporally controls Candida albicans biofilm formation. Mol. Microbiol. 96, 1226–1239 (2015).
Homann, O. R. & Johnson, A. D. MochiView: versatile software for genome browsing and DNA motif analysis. BMC Biol. 8, 49 (2010).
Peti, W. & Page, R. Strategies to maximize heterologous protein expression in Escherichia coli with minimal cost. Protein Expr. Purif. 51, 1–10 (2007).
Horton, R. M., Hunt, H. D., Ho, S. N., Pullen, J. K. & Pease, L. R. Engineering hybrid genes without the use of restriction enzymes: gene splicing by overlap extension. Gene 77, 61–68 (1989).
Reuss, O., Vik, A., Kolter, R. & Morschhauser, J. The SAT1 flipper, an optimized tool for gene disruption in Candida albicans. Gene 341, 119–127 (2004).
Gerami-Nejad, M., Dulmage, K. & Berman, J. Additional cassettes for epitope and fluorescent fusion proteins in Candida albicans. Yeast 26, 399–406 (2009).
Care, R. S., Trevethick, J., Binley, K. M. & Sudbery, P. E. The MET3 promoter: a new tool for Candida albicans molecular genetics. Mol. Microbiol. 34, 792–798 (1999).
Gallardo, I. F. et al. High-throughput universal DNA curtain arrays for single-molecule fluorescence imaging. Langmuir 31, 10310–10317 (2015).
Hammer, O., Harper, D. A. & Ryan, P. D. PAST: paleontological statistics software package for education and data analysis. Palaeontol. Electron 4, 1–9 (2001).
Acknowledgements
We thank R. Tjian for the gift of reporter cell lines, S. Sandler and J. Morschhauser for plasmids, L. Brossay for help with tissue culture, G. Williams for help with confocal microscopy, and members of the Bennett laboratory for helpful discussions. This work is supported by the National Institute of Allergy and Infectious Disease (grant nos. AI081704, AI135228 and AI141893 to R.J.B. and AI137975 to A.D.H.), the Burroughs Wellcome Fund (PATH award to R.J.B.), the National Heart, Lung and Blood Institute (grant no. T32HL134625 to M.I.S.), the National Institute of Dental and Craniofacial Research (grant no. F31DE02968001 to M.I.S.), the National Institute of Mental Health (grant no. T32MH020068 to V.H.R.), the National Institute of Neurological Disorders and Stroke (grant no. F31NS110301 to V.H.R.), a Howard Hughes Medical Institute International Student Fellowship (to Y.K.), the National Institute of General Medical Sciences (grant nos. GM120554 to I.J.F. and GM118530 to N.L.F.), the Welch Foundation (grant no. F-1808 to I.J.F.) and the National Science Foundation (grant nos. 1453358 to I.J.F. and 1845734 to N.L.F.).
Author information
Authors and Affiliations
Contributions
C.F. and R.J.B. conceived the study. C.F., M.I.S., Y.K., M.H., M.A.D., N.V.J., A.D.H., V.H.R. and R.J.B. investigated the study. C.F., M.I.S., Y.K. and A.H. formally analysed the study. N.V.J., A.H., N.L.F., I.J.F. and R.J.B. provided the resources. C.F., M.I.S. and R.J.B. wrote the original draft of the manuscript. C.F., M.I.S., A.D.H., N.L.F., I.J.F. and R.J.B. reviewed and edited the manuscript. C.F., M.I.S., Y.K. and A.D.H. visualized the study. N.L.F., I.J.F. and R.J.B. supervised it. A.D.H., N.L.F., I.J.F. and R.J.B. acquired the funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
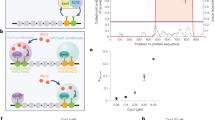
Extended Data Fig. 1 ChIP-chip data for master white-opaque TFs at select C. albicans genes.
Top, ChIP-chip enrichment peaks shown for Wor1 (orange), Wor2 (pink), Wor3 (blue), Czf1 (green), Efg1 (purple) and Ahr1 (red). Solid lines indicate TF binding and dotted lines indicate controls. ORFs are represented by purple boxes and lighter purple boxes represent untranslated regions. Bottom, Positions of consensus DNA binding sites for each TF. The large circles represent motif hits with >75% of the maximum score, medium circles represent motif hits that have 50–75% of the maximum score, and small circles represent motif hits that have 25–50% of the maximum score. ChIP enrichment plot generated from data in refs. 27,30,36 and motif analysis performed using data from refs. 27,30.
Extended Data Fig. 2 Purified C. albicans white-opaque TFs used in this study.
a, Schematic of TF expression constructs, including 6x histidine tag, MBP, and TEV protease site. b, Purified proteins used in this study. SDS-PAGE gels of C. albicans Wor1, Efg1, Czf1 and Wor4 HIS6-MBP-TF fusion proteins, as well as proteins with different PrLD deletions and those where the DBD has been replaced with GFP. c, Image of a HIS6-MBP-Efg1 protein solution (30 μM) without (left) and with (right) the addition of TEV protease for 30 min at 22 °C. Cloudiness indicates formation of phase-separated condensates, as confirmed by microscopy. Protein droplets formed in 10 mM Tris-HCl, pH 7.4, 150 mM NaCl at 22 °C. Scale bar; 5 μm. Representative data for an experiment repeated more than three times with similar results.
Extended Data Fig. 3 Hexanediol treatment selectively disrupts C. albicans TF condensates even during co-compartmentalization with other TFs.
a, Images of Efg1, Czf1, Wor1 (CaCmWor1), and Wor4 droplets at the indicated concentrations with or without 10% 1,6- or 2,5-hexanediol. For hexanediol treatment, proteins were incubated with TEV for 30 minutes in 10 mM Tris-HCl, pH 7.4, 150 mM NaCl, at 22 °C, and then mixed with 1,6- or 2,5-hexanediol in the same buffer, incubated for 10 minutes, and imaged. Wor1, Wor4, and Czf1 assays also included 5% PEG-8000. Where indicated for Wor4, hexanediol was added for 10 minutes and then TEV/PEG-8000 added and the sample incubated for an additional 30 minutes prior to imaging. Images represent a single experimental replicate with assays repeated at least twice with similar results. Scale bars; 10 μm. b, Representative images of fluorescently labeled Efg1, Wor1 (CaWor1), Wor4, and Czf1 proteins compartmentalized within Efg1 condensates, and treated with 10% 1,6- or 2,5-hexanediol. Unlabeled bulk protein (15 μM) was mixed with each of the fluorescently labeled proteins (37.5 nM) in 10 mM Tris-HCl, pH 7.4, 150 mM NaCl. Proteins were then incubated at 22 °C with TEV for 30 minutes and treated with 1,6- or 2,5-hexanediol in the same buffer for 10 minutes prior to imaging. Dylight NHS-Ester labeling of the 4 proteins used fluors of 405, 488, 550 and 633 nm. Images represent a single experimental replicate with assays performed three times with similar results. Scale bar, 10 μm; images are maximum Z-stack projections. c, Representative images of fluorescently labeled Efg1, Wor1 (CaWor1), Wor4, and Czf1 proteins compartmentalized within Czf1, Wor1(CaCmWor1), or Wor4 condensates. Unlabeled bulk proteins (15 μM) were mixed with each of the fluorescently labeled proteins (37.5 nM) in 10 mM Tris-HCl, pH 7.4, 150 mM NaCl. Proteins were then incubated at 22 °C with TEV for 30 min. Dylight NHS-Ester labeling of the 4 proteins used fluors of 488, 550, 405, and 633 nm. Images represent a single experimental replicate, with assays performed three times with similar results. Scale bars, 10 μm; images are maximum Z-stack projections.
Extended Data Fig. 4 PrLDs enable the co-partitioning of C. albicans white-opaque TFs.
Analysis of the ability of full-length or truncated TFs to co-partition within Efg1 condensates. a, Schematics of the GFP fusion proteins tested in phase separation assays. b, Efg1-GFP, Wor4-GFP, Czf1-GFP or Wor1-GFP variants were evaluated for their ability to co-partition with unlabeled Efg1 droplets. For each protein, the DBD was replaced with GFP. In all assays, proteins were incubated with TEV for 30 min at 22 °C in 10 mM Tris-HCl, pH 7.4, 150 mM NaCl. Bulk (full-length) Efg1 was present at 30 μM with 3 μM TF-GFP fusion proteins included in each reaction. Box and whisker plots show all data points, maximum to minimum, and indicate enrichment ratios for each TF-GFP fusion protein with condensates formed by full-length Efg1. For each plot, data are median (line), mean (‘+’), 25–75th percentiles (box), and 5–95th percentiles (whiskers). Droplets were located in the DIC channel, and the intensity for the GFP signal inside the droplet compared to the signal intensity outside the droplet, following subtraction of fluorescence background. At least five images were used for quantification, with 25 total droplets measured for each construct. Statistical significance was performed using a two-tailed Mann-Whitney U-test; P-values: a, < 0.0001; ns, not significant. Scale bars; 5 μm.
Supplementary information
Supplementary Table 1
List of plasmids, oligos and strains used in the study.
Source data
Source Data Fig. 3
Excel file of quantitative DNA curtain data.
Source Data Fig. 4
Excel file of phenotypical switching in Candida data.
Source Data Fig. 5
Excel file of quantitative analysis of microscopy data.
Source Data Fig. 6
Excel file of quantitative analysis of microscopy data.
Source Data Extended Data Fig. 4
Excel file of quantitative analysis of microscopy data for Extended Data Fig. 4.
Rights and permissions
About this article
Cite this article
Frazer, C., Staples, M.I., Kim, Y. et al. Epigenetic cell fate in Candida albicans is controlled by transcription factor condensates acting at super-enhancer-like elements. Nat Microbiol 5, 1374–1389 (2020). https://doi.org/10.1038/s41564-020-0760-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-020-0760-7
This article is cited by
-
Heterotypic interactions can drive selective co-condensation of prion-like low-complexity domains of FET proteins and mammalian SWI/SNF complex
Nature Communications (2024)
-
Phase separation in fungi
Nature Microbiology (2023)
-
Multifactor transcriptional control of alternative oxidase induction integrates diverse environmental inputs to enable fungal virulence
Nature Communications (2023)
-
Intrinsically disordered CO2 sensors
Nature Cell Biology (2022)
-
Phase separation and cell fate in Candida
Nature Microbiology (2020)