Abstract
Recent population-based1,2,3,4 and clinical studies5 have identified a range of factors associated with human gut microbiome variation. Murine quantitative trait loci6, human twin studies7 and microbiome genome-wide association studies1,3,8,9,10,11,12 have provided evidence for genetic contributions to microbiome composition. Despite this, there is still poor overlap in genetic association across human studies. Using appropriate taxon-specific models, along with support from independent cohorts, we show an association between human host genotype and gut microbiome variation. We also suggest that interpretation of applied analyses using genetic associations is complicated by the probable overlap between genetic contributions and heritable components of host environment. Using faecal 16S ribosomal RNA gene sequences and host genotype data from the Flemish Gut Flora Project (n = 2,223) and two German cohorts (FoCus, n = 950; PopGen, n = 717), we identify genetic associations involving multiple microbial traits. Two of these associations achieved a study-level threshold of P = 1.57 × 10−10; an association between Ruminococcus and rs150018970 near RAPGEF1 on chromosome 9, and between Coprococcus and rs561177583 within LINC01787 on chromosome 1. Exploratory analyses were undertaken using 11 other genome-wide associations with strong evidence for association (P < 2.5 × 10−8) and a previously reported signal of association between rs4988235 (MCM6/LCT) and Bifidobacterium. Across these 14 single-nucleotide polymorphisms there was evidence of signal overlap with other genome-wide association studies, including those for age at menarche and cardiometabolic traits. Mendelian randomization analysis was able to estimate associations between microbial traits and disease (including Bifidobacterium and body composition); however, in the absence of clear microbiome-driven effects, caution is needed in interpretation. Overall, this work marks a growing catalogue of genetic associations that will provide insight into the contribution of host genotype to gut microbiome. Despite this, the uncertain origin of association signals will likely complicate future work looking to dissect function or use associations for causal inference analysis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
All microbiome GWAS summary statistics are available online at the University of Bristol data repository, data.bris, at https://doi.org/10.5523/bris.22bqn399f9i432q56gt3wfhzlc. FGFP rarefaction count and transformed microbial trait data can be found in Supplementary Table 2. FGFP genotype data and host metadata from this study are not open access, but are available in accordance and in consent with ethical permission through managed access subject to a data use agreement with the FGFP and organized via principal investigator J.R. The process of enquiry for data access is outlined as follows: following data request by email to jeroen.raes@kuleuven.be, the FGFP data access committee will evaluate access permission, which will be granted upon signature of a data use agreement between the governing legal entities. This is outlined on the study website http://www.raeslab.org/companion/fgfp-gwas/. Raw 16S data are available at the European Genome/Phenome Archive under accession no. EGAS00001004420. The datasets from Universitätsklinikums Schleswig-Holstein are available by application through their biobank (https://www.uksh.de/p2n/).
Code availability
The full analysis pipeline is available at https://github.com/kul-fgfpgwas/rep-cookbook and includes four parts: (1) microbiome processing, (2) genotype quality control and imputation, (3) genome-wide association analysis and (4) phylogenetic analysis.
References
Rothschild, D. et al. Environment dominates over host genetics in shaping human gut microbiota. Nature 555, 210–215 (2018).
McDonald, D. et al. American Gut: an open platform for citizen science microbiome research. mSystems 3, e00031-18 (2018).
Zhernakova, A. et al. Population-based metagenomics analysis reveals markers for gut microbiome composition and diversity. Science 352, 565–569 (2016).
Falony, G. et al. Population-level analysis of gut microbiome variation. Science 352, 560–564 (2016).
Gilbert, J. A. et al. Current understanding of the human microbiome. Nat. Med. 24, 392–400 (2018).
McKnite, A. M. et al. Murine gut microbiota is defined by host genetics and modulates variation of metabolic traits. PLoS ONE 7, e39191 (2012).
Goodrich, J. K. et al. Genetic determinants of the gut microbiome in UK twins. Cell Host Microbe 19, 731–743 (2016).
Blekhman, R. et al. Host genetic variation impacts microbiome composition across human body sites. Genome Biol. 16, 191 (2015).
Davenport, E. R. et al. Genome-wide association studies of the human gut microbiota. PLoS ONE 10, e0140301 (2015).
Bonder, M. J. et al. The effect of host genetics on the gut microbiome. Nat. Genet. 48, 1407–1412 (2016).
Wang, J. et al. Genome-wide association analysis identifies variation in vitamin D receptor and other host factors influencing the gut microbiota. Nat. Genet. 48, 1396–1406 (2016).
Turpin, W. et al. Association of host genome with intestinal microbial composition in a large healthy cohort. Nat. Genet. 48, 1413–1417 (2016).
Wang, J. et al. Meta-analysis of human genome-microbiome association studies: the MiBioGen consortium initiative. Microbiome 6, 101 (2018).
Vandeputte, D., Tito, R. Y., Vanleeuwen, R., Falony, G. & Raes, J. Practical considerations for large-scale gut microbiome studies. FEMS Microbiol. Rev. 41, S154–S167 (2017).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Krawczak, M. et al. PopGen: population-based recruitment of patients and controls for the analysis of complex genotype–phenotype relationships. Public Health Genomics 9, 55–61 (2006).
Ferreira-Halder, C. V., Faria, A. V., de, S. & Andrade, S. S. Action and function of Faecalibacterium prausnitzii in health and disease. Best Pract. Res. Clin. Gastroenterol. 31, 643–648 (2017).
Cohen, L. J. et al. Commensal bacteria make GPCR ligands that mimic human signalling molecules. Nature 549, 48–53 (2017).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Coady, M. J., Wallendorff, B., Gagnon, D. G. & Lapointe, J.-Y. Identification of a novel Na +/myo -inositol cotransporter. J. Biol. Chem. 277, 35219–35224 (2002).
Raffler, J. et al. Genome-wide association study with targeted and non-targeted NMR metabolomics identifies 15 novel loci of urinary human metabolic individuality. PLOS Genet. 11, e1005487 (2015).
Ugrankar, R., Theodoropoulos, P., Akdemir, F., Henne, W. M. & Graff, J. M. Circulating glucose levels inversely correlate with Drosophila larval feeding through insulin signaling and SLC5A11. Commun. Biol. 1, 110 (2018).
Puddu, A., Sanguineti, R., Montecucco, F. & Viviani, G. L. Evidence for the gut microbiota short-chain fatty acids as key pathophysiological molecules improving diabetes. Mediators Inflamm. 2014, 162021 (2014).
Gao, Z. et al. Butyrate improves insulin sensitivity and increases energy expenditure in mice. Diabetes 58, 1509–1517 (2009).
Zambell, K. L., Fitch, M. D. & Fleming, S. E. Acetate and butyrate are the major substrates for de novo lipogenesis in rat colonic epithelial cells. J. Nutr. 133, 3509–3515 (2003).
Nishina, P. M. & Freedland, R. A. Effects of propionate on lipid biosynthesis in isolated rat hepatocytes. J. Nutr. 120, 668–673 (1990).
Machiela, M. J. & Chanock, S. J. LDlink: a web-based application for exploring population-specific haplotype structure and linking correlated alleles of possible functional variants. Bioinformatics 31, 3555–3557 (2015).
Kamat, M. A. et al. PhenoScanner V2: an expanded tool for searching human genotype–phenotype associations. Bioinformatics 35, 4851–4853 (2019).
Watanabe, K., Taskesen, E., van Bochoven, A. & Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 8, 1826 (2017).
Davey Smith, G. & Hemani, G. Mendelian randomization: genetic anchors for causal inference in epidemiological studies. Hum. Mol. Genet. 23, R89–R98 (2014).
Wade, K. H. & Hall, L. J. Improving causality in microbiome research: can human genetic epidemiology help? Wellcome Open Res. 4, 199 (2020).
Tito, R. Y. et al. Population-level analysis of Blastocystis subtype prevalence and variation in the human gut microbiota. Gut 68, 1180–1189 (2018).
Hildebrand, F., Tadeo, R., Voigt, A., Bork, P. & Raes, J. LotuS: an efficient and user-friendly OTU processing pipeline. Microbiome 2, 37 (2014).
Holmes, I., Harris, K. & Quince, C. Dirichlet multinomial mixtures: generative models for microbial metagenomics. PLoS ONE 7, e30126 (2012).
Karssen, L. C., van Duijn, C. M. & Aulchenko, Y. S. The GenABEL Project for statistical genomics. F1000Res. 5, 914 (2016).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2016).
Wang, K. et al. PennCNV: an integrated hidden Markov model designed for high-resolution copy number variation detection in whole-genome SNP genotyping data. Genome Res. 17, 1665–1674 (2007).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
O’Connell, J. et al. Haplotype estimation for biobank-scale data sets. Nat. Genet. 48, 817–820 (2016).
Howie, B. N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
The H. R. Consortium. A reference panel of 64,976 haplotypes for genotype imputation. Nat. Genet. 48, 1279–1283 (2016).
Deelen, P. et al. Improved imputation quality of low-frequency and rare variants in European samples using the ‘Genome of The Netherlands’. Eur. J. Hum. Genet. 22, 1321–1326 (2014).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Graw, S., Henn, R., Thompson, J. A. & Koestler, D. C. PwrEWAS: a user-friendly tool for comprehensive power estimation for epigenome wide association studies (EWAS). BMC Bioinformatics 20, 218 (2019).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Liu, J. Z. et al. Meta-analysis and imputation refines the association of 15q25 with smoking quantity. Nat. Genet. 42, 436–440 (2010).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2 – approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 44, W242–W245 (2016).
Durinck, S., Spellman, P. T., Birney, E. & Huber, W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat. Protoc. 4, 1184–1191 (2009).
Pers, T. H. et al. Biological interpretation of genome-wide association studies using predicted gene functions. Nat. Commun. 6, 5890 (2015).
Iotchkova, V. et al. GARFIELD classifies disease-relevant genomic features through integration of functional annotations with association signals. Nat Genet. 51, 343–353 (2019).
Bowden, J., Davey Smith, G., Haycock, P. C. & Burgess, S. Consistent estimation in Mendelian randomization with some invalid instruments using a weighted median estimator. Genet. Epidemiol. 40, 304–314 (2016).
Hartwig, F. P., Davey Smith, G. & Bowden, J. Robust inference in summary data Mendelian randomization via the zero modal pleiotropy assumption. Int. J. Epidemiol. 46, 1985–1998 (2017).
Bowden, J., Davey Smith, G. & Burgess, S. Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int. J. Epidemiol. 44, 512–525 (2015).
Shungin, D. et al. New genetic loci link adipose and insulin biology to body fat distribution. Nature 518, 187–196 (2015).
Locke, A. E. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Morris, A. et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat. Genet. 44, 981–990 (2012).
Liu, J. Z. et al. Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat. Genet. 47, 979–986 (2015).
Okada, Y. et al. Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature 506, 376–381 (2014).
Lambert, J.-C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 45, 1452–1458 (2013).
Simón-Sánchez, J. et al. Genome-wide association study reveals genetic risk underlying Parkinson’s disease. Nat. Genet. 41, 1308–1312 (2009).
Ripke, S. et al. A mega-analysis of genome-wide association studies for major depressive disorder. Mol. Psychiatry 18, 497–511 (2013).
Acknowledgements
FGFP procedures were approved by the Medical Ethics Committee of the University of Brussels–Brussels University Hospital (approval no. 143201215505, 5 December 2012). A declaration concerning the FGFP privacy policy was submitted to the Belgian Commission for the Protection of Privacy. Written informed consent was obtained from all participants. The FGFP was funded with the support of the Flemish government (no. IWT130359), the Research Fund–Flanders (FWO) Odysseus programme (no. G.0924.09), the King Baudouin Foundation (no. 2012-J80000-004), FP7 METACARDIS HEALTH-F4-2012-305312, VIB, the Rega Institute for Medical Research and KU Leuven. We acknowledge the contribution of Flemish general practitioners and pharmacists to data and sample collection. Finally, we thank all FGFP volunteers for their participation in the project. R.B. is funded by FWO through a postdoctoral fellowship (no. 1221620N). S.V.-S. is supported by Marie Curie Actions FP7 People COFUND–Proposal 267139 (acronym OMICS@VIB). N.J.T. is a Wellcome Trust Investigator (no. 202802/Z/16/Z), is supported by the University of Bristol NIHR Biomedical Research Centre (no. BRC-1215-20011) and works within the CRUK Integrative Cancer Epidemiology Programme (no. C18281/A19169). S.R. and N.J.T. work within a unit that receives funding from the MRC and University of Bristol (nos. MC_UU_12013/3 and MC_UU_00011/6). D.A.H. is supported by the Wellcome Investigator Award (no. 202802/Z/16/Z). K.H.W. is supported by the Elizabeth Blackwell Institute for Health Research, University of Bristol and the Wellcome Trust Institutional Strategic Support Fund (no. 204813/Z/16/Z). M.C.R. was funded by the German Research Foundation Collaborative Research Centre 1182 (no. CRC1182; Origin and Function of Metaorganisms).
Author information
Authors and Affiliations
Contributions
J.R. and N.J.T. conceived and designed the study. J.W., R.Y.T., G.F., M.J., S.V.-S. and L.H. performed sampling and metadata collection. J.W., R.Y.T., L.R. and C.V. extracted and sequenced DNA. D.A.H., R.B., K.H.W., J.W. and M.C.R. performed microbiome GWAS and MR analysis. D.A.H., R.B., J.W., K.H.W., N.T. and J.R. wrote the manuscript. All authors revised and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Variability in heritability.
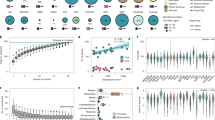
Scatter plots of heritability estimates in the FGFP data set derived from rank normal, box-cox and log2 transformed abundance data (a, b); and estimates between the FGFP data and those previously published (c, d). (a) The plot compares rank normal transformed (x-axis) data to box-cox transformed data (y-axis). (b) The plot compares rank normal transformed (x-axis) data to log2 transformed data (y-axis). (c) The plot compares box-cox transformed data from FGFP (x-axis) and Goodrich et al. (y-axis). (d) The plot compares FGFP rank normal transformed data (x-axis) to the quantile-quantile normalized data of Davenport et al. (y-axis). The color of each point indicates a measure of average abundance in FGFP, and the size of each dot illustrates the prevalence for each taxon. The red line is a loess curve fit through the data. Confidence intervals of the fitted curve is shaded in grey. An estimate of the Spearman’s rho correlation coefficient between the data points is provided in the sub-title of each plot, along with its’ two-sided p-value. Sample size and heritability estimates can be found in Supplementary Table 3.
Extended Data Fig. 2 Effect estimates for top hits.
A forest plot of beta (effect) point estimates (blue dots) and standard errors (horizontal bars) for each of the genome-wide significant SNP-microbial trait (MT) associations. Each of the three cohorts used in the meta-analysis are represented in each MT plot. The vertical bar indicates zero on the effect- or x-axis. Sample sizes for each microbial trait and data used to generate this plot can be found in Supplementary Table 5.
Extended Data Fig. 3 Mendelian randomization framework.
(a) the causal effect of a microbial trait on a disease, whereby any invalidation of the exclusion restriction criteria implies a pleiotropic association between the genetic variant and the disease independent of the microbial trait, (b) the genetic variant seemingly associated with the microbial trait is predominantly associated with the disease, which in turn effects the microbial trait (that is, reverse causation) and (c) the genetic variant is associated with an external variable that is independently associated with the microbial trait and outcome.
Extended Data Fig. 4 FGFP DADA2 data summaries.
Rank ordered mean percent abundance (a) and prevalence (b) for each observed phylum, in the FGFP cohort. Prevalence is defined as the proportion of individuals in the cohort containing a rarefaction read assigned to a taxon, in this instance a phylum. Data for this plot can be found in Supplementary Table 3. (c) A scatter plot of the nMDS, β-diversity for the quality-controlled FGFP cohort. Individuals are colored by their defined enterotype, and boxplots illustrating the enterotype structure across the two nMDS (β-diversity) axes are presented. The stress level, of forcing the individual level data into two axes, in the nMDS analysis is presented in the lower right-hand corner of the scatter plot. Dashed lines are equidistant guidelines. Each box plot presents the mean, first and third quantiles, and 95% confidence intervals of the data distribution. The sample sizes for each enterotype class is Bact1 = 793, Rum = 660, Bact2 = 418, Prev = 388, as presented in Supplementary Table 2.
Extended Data Fig. 5 Selection of taxa for mGWAS analysis.
(a) Association between abundance and prevalence in 16S DAD2 data. A scatter plot illustrating the relationship between average abundance and prevalence in the FGFP cohort. Taxon retained for analysis in the mGWAS are colored in blue, while those that did not meet our analysis criteria are colored in red. The size of each taxon dot represents the number of individuals in the FGFP dataset in which that taxon made up at least 5% of the data or had 500 rarefaction reads. Dashed blue lines represent the prevalence floor (horizontal) and average abundance = 40 (vertical). The grey curve is a loess best fitting curve. Data used to generate this plot can be found in Supplementary Table 3. (b) Clustering dendrogram of 139 mGWAS taxa meeting criteria for analysis in mGWAS. The red horizontal line (0.015) represents the level at which we defined clades or clusters of statistically redundant taxa, of which the lowest taxonomic level taxa was retained for analysis. Data used to generate this plot can be found in Supplementary Table 2. (c) Correlation structure among 139 mGWAS taxa. A Spearman’s correlation plot of the 139 mGWAS worthy taxa. The color key scales from −1 to 1 and indicates the estimated Spearman’s rho correlation coefficient. The 139 taxa are clustered into eight squares, denoting the best eight clusters in the data. Eight was chosen for illustration purposes because there are eight phyla in the data. However, the clustered data does not necessarily conform to the eight phyla. Data used to generate this plot can be found in Supplementary Table 2.
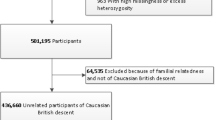
Extended Data Fig. 6 Flowcharts for genotyping and mGWAS.
(a) Flowchart on the genotyping and quality control steps taken to prepare the FGFP data for the mGWAS. (b) Flowchart for 16S DADA2 MT data preparation, FGFP mGWAS and targeted meta-analysis.
Extended Data Fig. 7 GCTA-GREML power estimates.
GCTA-GREML (genome wide complex trait analysis – genomic-relatedness-based restricted maximum-likelihood) genetic (co)variation power estimates. The plot on the left illustrates the relationship between the power (y-axis) to accurately quantify narrow sense heritability (x-axis) for varying sampling sizes (from 300 to 2100). The plot on the right illustrates the same relationship but for presence/absence microbial traits, where the number of presence and absence individuals is balanced.
Supplementary information
Supplementary Information
Supplementary Figs. 1–11, Tables 6, 15 and 16 and discussion.
Supplementary Table
Supplementary Tables 1–5, 7–14 and 17–22.
Rights and permissions
About this article
Cite this article
Hughes, D.A., Bacigalupe, R., Wang, J. et al. Genome-wide associations of human gut microbiome variation and implications for causal inference analyses. Nat Microbiol 5, 1079–1087 (2020). https://doi.org/10.1038/s41564-020-0743-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-020-0743-8
This article is cited by
-
Human genetic associations of the airway microbiome in chronic obstructive pulmonary disease
Respiratory Research (2024)
-
A genome-wide association study reveals the relationship between human genetic variation and the nasal microbiome
Communications Biology (2024)
-
Integration of polygenic and gut metagenomic risk prediction for common diseases
Nature Aging (2024)
-
Host genetic regulation of human gut microbial structural variation
Nature (2024)
-
Statistical modeling of gut microbiota for personalized health status monitoring
Microbiome (2023)