Abstract
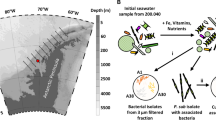
Niche theory is a foundational ecological concept that explains the distribution of species in natural environments. Identifying the dimensions of any organism’s niche is challenging because numerous environmental factors can affect organism viability. We used serial invasion experiments to introduce Ruegeria pomeroyi DSS-3, a heterotrophic marine bacterium, into a coastal phytoplankton bloom on 14 dates. RNA-sequencing analysis of R. pomeroyi was conducted after 90 min to assess its niche dimensions in this dynamic ecosystem. We identified ~100 external conditions eliciting transcriptional responses, which included substrates, nutrients, metals and biotic interactions such as antagonism, resistance and cofactor synthesis. The peak bloom was characterized by favourable states for most of the substrate dimensions, but low inferred growth rates of R. pomeroyi at this stage indicated that its niche was narrowed by factors other than substrate availability, most probably negative biotic interactions with the bloom dinoflagellate. Our findings indicate chemical and biological features of the ocean environment that can constrain where heterotrophic bacteria survive.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Data that support the findings of the present study have been deposited in the National Center for Biotechnology Information‘s Sequence Read Archive with BioProject nos. PRJNA641119 (RNA-seq) and PRJNA511156–PRJNA511331 (16S and 18S rRNA data), and the Biological and Chemical Oceanography Data Management Office under https://doi.org/10.1575/1912/bco-dmo.756413.2 at https://www.bco-dmo.org/dataset/756413/data (environmental data). Source data are provided with this paper.
References
Carlson, C. A. et al. Seasonal dynamics of SAR11 populations in the euphotic and mesopelagic zones of the northwestern Sargasso Sea. ISME J. 3, 283–295 (2009).
Palovaara, J. et al. Stimulation of growth by proteorhodopsin phototrophy involves regulation of central metabolic pathways in marine planktonic bacteria. Proc. Natl Acad. Sci. USA 111, E3650–E3658 (2014).
Poretsky, R. S., Sun, S., Mou, X. & Moran, M. A. Transporter genes expressed by coastal bacterioplankton in response to dissolved organic carbon. Environ. Microbiol. 12, 616–627 (2010).
Church, M. J., Hutchins, D. A. & Ducklow, H. W. Limitation of bacterial growth by dissolved organic matter and iron in the Southern Ocean. Appl. Environ. Microbiol. 66, 455–466 (2000).
Persson, O. P. et al. High abundance of virulence gene homologues in marine bacteria. Environ. Microbiol. 11, 1348–1357 (2009).
Yeung, L. Y. et al. Impact of diatom–diazotroph associations on carbon export in the Amazon River plume. Geophys. Res. Lett. 39, L18609 (2012).
Colwell, R. K. & Fuentes, E. R. Experimental studies of the niche. Annu. Rev. Ecol. Syst. 6, 281–310 (1975).
Hutchinson, G. E. Concluding remarks. Cold Spring Harb. Symp. Quant. Biol. 22, 415–427 (1957).
Cohan, F. M. What are bacterial species? Annu. Rev. Microbiol. 56, 457–487 (2002).
Erguder, T. H., Boon, N., Wittebolle, L., Marzorati, M. & Verstraete, W. Environmental factors shaping the ecological niches of ammonia-oxidizing archaea. FEMS Microbiol. Rev. 33, 855–869 (2009).
Meier, D. V. et al. Niche partitioning of diverse sulfur-oxidizing bacteria at hydrothermal vents. ISME J. 11, 1545–1558 (2017).
Martens-Habbena, W., Berube, P. M., Urakawa, H., José, R. & Stahl, D. A. Ammonia oxidation kinetics determine niche separation of nitrifying Archaea and Bacteria. Nature 461, 976–979 (2009).
Gifford, S. M., Sharma, S., Booth, M. & Moran, M. A. Expression patterns reveal niche diversification in a marine microbial assemblage. ISME J. 7, 281–298 (2013).
Landa, M., Burns, A. S., Roth, S. J. & Moran, M. A. Bacterial transcriptome remodeling during sequential co-culture with a marine dinoflagellate and diatom. ISME J. 11, 2677–2690 (2017).
Ottesen, E. A. et al. Pattern and synchrony of gene expression among sympatric marine microbial populations. Proc. Natl Acad. Sci. USA 110, E488–E497 (2013).
Galambos, D., Anderson, R. E., Reveillaud, J. & Huber, J. A. Genome-resolved metagenomics and metatranscriptomics reveal niche differentiation in functionally redundant microbial communities at deep-sea hydrothermal vents. Environ. Microbiol. 21, 4395–4410 (2019).
Nuccio, E. E. et al. Niche differentiation is spatially and temporally regulated in the rhizosphere. ISME J. 14, 999–1014 (2020).
Shaiber, A. & Eren, A. M. Composite metagenome-assembled genomes reduce the quality of public genome repositories. mBio 10, e00725-19 (2019).
Cottrell, M. T. & Kirchman, D. L. Transcriptional control in marine copiotrophic and oligotrophic bacteria with streamlined genomes. Appl. Environ. Microbiol. 82, 6010–6018 (2016).
Bell, T. Next-generation experiments linking community structure and ecosystem functioning. Environ. Microbiol. Rep. 11, 20–22 (2019).
Mallon, C. A., Van Elsas, J. D. & Salles, J. F. Microbial invasions: the process, patterns, and mechanisms. Trends Microbiol. 23, 719–729 (2015).
Kiene, R. P. et al. Unprecedented DMSP concentrations in a massive dinoflagellate bloom in Monterey Bay, CA. Geophys. Res. Lett. 46, 12279–12288 (2019).
Anderson, S. R., Diou-Cass, Q. P. & Harvey, E. L. Short-term estimates of phytoplankton growth and mortality in a tidal estuary. Limnol. Oceanogr. 63, 2411–2422 (2018).
Anderson, S. R. & Harvey, E. L. Seasonal variability and drivers of microzooplankton grazing and phytoplankton growth in a subtropical estuary. Front Mar. Sci. 6, 174–174 (2019).
González, J. M. et al. Silicibacter pomeroyi sp. nov. and Roseovarius nubinhibens sp. nov., dimethylsulfoniopropionate-demethylating bacteria from marine environments. Int. J. Syst. Evol. Microbiol. 53, 1261–1269 (2003).
Luo, H. & Moran, M. A. Evolutionary ecology of the marine Roseobacter clade. Microbiol. Mol. Biol. Rev. 78, 573–587 (2014).
Colwell, R. K. & Rangel, T. F. Hutchinson’s duality: the once and future niche. Proc. Natl Acad. Sci. USA 106, 19651–19658 (2009).
Holt, R. D. Bringing the Hutchinsonian niche into the 21st century: ecological and evolutionary perspectives. Proc. Natl Acad. Sci. USA 106, 19659–19665 (2009).
Alneberg, J. et al. Ecosystem-wide metagenomic binning enables prediction of ecological niches from genomes. Commun. Biol. 3, 119 (2020).
Baltar, F. et al. Towards integrating evolution, metabolism, and climate change studies of marine ecosystems. Trends Ecol. Evol. 34, 1022–1033 (2019).
Muller, E. E. Determining microbial niche breadth in the environment for better ecosystem fate predictions. mSystems 4, e00080-19 (2019).
Chan, L.-K. et al. Transcriptional changes underlying elemental stoichiometry shifts in a marine heterotrophic bacterium. Front. Microbiol. 3, 159 (2012).
Kudela, R. M., Seeyave, S. & Cochlan, W. P. The role of nutrients in regulation and promotion of harmful algal blooms in upwelling systems. Prog. Oceanogr. 85, 122–135 (2010).
Moran, M. A. et al. Genome sequence of Silicibacter pomeroyi reveals adaptations to the marine environment. Nature 432, 910–913 (2004).
Amin, S. A. et al. Interaction and signalling between a cosmopolitan phytoplankton and associated bacteria. Nature 522, 98–101 (2015).
Sharpe, G. C., Gifford, S. M. & Septer, A. N. A model Roseobacter employs a diffusible killing mechanism to eliminate competitors. mSystems 5, e00443-20 (2020).
Gil-Turnes, M. S., Hay, M. E. & Fenical, W. Symbiotic marine bacteria chemically defend crustacean embryos from a pathogenic fungus. Science 246, 116–118 (1989).
Lopanik, N., Lindquist, N. & Targett, N. Potent cytotoxins produced by a microbial symbiont protect host larvae from predation. Oecologia 139, 131–139 (2004).
Croft, M. T., Lawrence, A. D., Raux-Deery, E., Warren, M. J. & Smith, A. G. Algae acquire vitamin B12 through a symbiotic relationship with bacteria. Nature 438, 90–93 (2005).
Sañudo-Wilhelmy, S. A., Gómez-Consarnau, L., Suffridge, C. & Webb, E. A. The role of B vitamins in marine biogeochemistry. Annu. Rev. Mar. Sci. 6, 339–367 (2014).
Biers, E. J. et al. Occurrence and expression of gene transfer agent genes in marine bacterioplankton. Appl. Environ. Microbiol. 74, 2933–2939 (2008).
Gravel, D. et al. Experimental niche evolution alters the strength of the diversity–productivity relationship. Nature 469, 89–94 (2011).
Vergin, K. L. et al. High intraspecific recombination rate in a native population of Candidatus Pelagibacter ubique (SAR11). Environ. Microbiol. 9, 2430–2440 (2007).
McDaniel, L. D. et al. High frequency of horizontal gene transfer in the oceans. Science 330, 50–50 (2010).
Nuss, A. M., Glaeser, J., Berghoff, B. A. & Klug, G. Overlapping alternative sigma factor regulons in the response to singlet oxygen in Rhodobacter sphaeroides. J. Bacteriol. 192, 2613–2623 (2010).
Berghoff, B. A. et al. Anoxygenic photosynthesis and photooxidative stress: a particular challenge for Roseobacter. Environ. Microbiol. 13, 775–791 (2011).
Zhao, K., Liu, M. & Burgess, R. R. The global transcriptional response of Escherichia coli to induced σ32 protein involves σ32 regulon activation followed by inactivation and degradation of σ32 in vivo. J. Biol. Chem. 280, 17758–17768 (2005).
Diaz, J. M. et al. Widespread production of extracellular superoxide by heterotrophic bacteria. Science 340, 1223–1226 (2013).
Wietz, M., Duncan, K., Patin, N. V. & Jensen, P. R. Antagonistic interactions mediated by marine bacteria: the role of small molecules. J. Chem. Ecol. 39, 879–891 (2013).
Maguire, B. A. Inhibition of bacterial ribosome assembly: a suitable drug target? Microbiol. Mol. Biol. Rev. 73, 22–35 (2009).
Wei, Y. et al. High-density microarray-mediated gene expression profiling of Escherichia coli. J. Bacteriol. 183, 545–556 (2001).
Wilson, D. N. & Nierhaus, K. H. The weird and wonderful world of bacterial ribosome regulation. Crit. Rev. Biochem. Mol. Biol. 42, 187–219 (2007).
Vinas, N. Relationships between Growth Rate and Gene Expression in Ruegeria pomeroyi DSS-3, a Model Marine Alphaproteobacterium. MSc thesis, Clemson Univ. (2015).
Ishihama, A. Functional modulation of Escherichia coli RNA polymerase. Annu. Rev. Microbiol. 54, 499–518 (2000).
González, J. M., Kiene, R. P. & Moran, M. A. Transformation of sulfur compounds by an abundant lineage of marine bacteria in the α-subclass of the class Proteobacteria. Appl. Environ. Microbiol. 65, 3810–3819 (1999).
Denger, K., Lehmann, S. & Cook, A. M. Molecular genetics and biochemistry of N-acetyltaurine degradation by Cupriavidus necator H16. Microbiology 157, 2983–2991 (2011).
Lidbury, I., Murrell, J. C. & Chen, Y. Trimethylamine N-oxide metabolism by abundant marine heterotrophic bacteria. Proc. Natl Acad. Sci. USA 111, 2710–2715 (2014).
Mou, X., Sun, S., Edwards, R. A., Hodson, R. E. & Moran, M. A. Bacterial carbon processing by generalist species in the coastal ocean. Nature 451, 708–711 (2008).
Schulz, A. et al. Feeding on compatible solutes: a substrate‐induced pathway for uptake and catabolism of ectoines and its genetic control by EnuR. Environ. Microbiol. 19, 926–946 (2017).
Weinitschke, S., Sharma, P. I., Stingl, U., Cook, A. M. & Smits, T. H. Gene clusters involved in isethionate degradation by terrestrial and marine bacteria. Appl. Environ. Microbiol. 76, 618–621 (2010).
Jessup, D. A., Miller, M. A., Ryan, J. P., Nevins, H. M. & Kerkering, H. A. Mass stranding of marine birds caused by a surfactant-producing red tide. PLoS ONE 4, 4550 (2009).
Jones, T. et al. Mass mortality of marine birds in the Northeast Pacific caused by Akashiwo sanguinea. Mar. Ecol. Prog. Ser. 579, 111–127 (2017).
Xu, N. et al. Acute toxicity of the cosmopolitan bloom-forming dinoflagellate Akashiwo sanguinea to finfish, shellfish, and zooplankton. Aquat. Microb. Ecol. 80, 209–222 (2017).
Kiene, R. P. & Linn, L. J. Distribution and turnover of dissolved DMSP and its relationship with bacterial production and dimethylsulfide in the Gulf of Mexico. Limnol. Oceanogr. 45, 849–861 (2000).
Motard-Côté, J., Kieber, D. J., Rellinger, A. & Kiene, R. P. Influence of the Mississippi River plume and non-bioavailable DMSP on dissolved DMSP turnover in the northern Gulf of Mexico. Environ. Chem. 13, 280–280 (2016).
Lally, E. T., Hill, R. B., Kieba, I. R. & Korostoff, J. The interaction between RTX toxins and target cells. Trends Microbiol. 7, 356–361 (1999).
Billen, G. & Fontigny, A. Dynamics of a Phaeocystis-dominated spring bloom in Belgian coastal waters. II. Bacterioplankton dynamics. Mar. Ecol. Prog. Ser. 37, 249–257 (1987).
Buchan, A., LeCleir, G. R., Gulvik, C. A. & González, J. M. Master recyclers: features and functions of bacteria associated with phytoplankton blooms. Nat. Rev. Microbiol. 12, 686–698 (2014).
Bunse, C. et al. Spatio-temporal interdependence of bacteria and phytoplankton during a Baltic Sea spring bloom. Front. Microbiol. 7, 517–517 (2016).
Pinhassi, J. et al. Changes in bacterioplankton composition under different phytoplankton regimens. Appl. Environ. Microbiol. 70, 6753–6766 (2004).
Teeling, H. et al. Substrate-controlled succession of marine bacterioplankton populations induced by a phytoplankton bloom. Science 336, 608–611 (2012).
Morris, J. J., Johnson, Z. I., Szul, M. J., Keller, M. & Zinser, E. R. Dependence of the cyanobacterium Prochlorococcus on hydrogen peroxide scavenging microbes for growth at the ocean’s surface. PLoS ONE 6, e16805 (2011).
Stock, F. et al. N-acyl homoserine lactone derived tetramic acids impair photosynthesis in Phaeodactylum tricornutum. ACS Chem. Biol. 14, 198–203 (2019).
Bruno, J. F., Stachowicz, J. J. & Bertness, M. D. Inclusion of facilitation into ecological theory. Trends Ecol. Evol. 18, 119–125 (2003).
Morris, J. J., Lenski, R. E. & Zinser, E. R. The black queen hypothesis: evolution of dependencies through adaptive gene loss. mBio 3, e00036-12 (2012).
Pacheco, A. R., Moel, M. & Segrè, D. Costless metabolic secretions as drivers of interspecies interactions in microbial ecosystems. Nat. Commun. 10, 103 (2019).
Saupe, E. E. et al. Reconstructing ecological niche evolution when niches are incompletely characterized. Syst. Biol. 67, 428–438 (2018).
Fu, H., Uchimiya, M., Gore, J. & Moran, M. A. Ecological drivers of bacterial community assembly in synthetic phycospheres. Proc. Natl Acad. Sci. USA 117, 3656–3662 (2020).
González, J. M., Mayer, F., Moran, M. A., Hodson, R. E. & Whitman, W. B. Microbulbifer hydrolyticus gen. nov., sp. nov., and Marinobacterium georgiense gen. nov., sp. nov., two marine bacteria from a lignin-rich pulp mill waste enrichment community. Int. J. Syst. Evol. Microbiol. 47, 369–376 (1997).
Nowinski, B. et al. Microbial metagenomes and metatranscriptomes during a coastal phytoplankton bloom. Sci. Data 6, 129 (2019).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Anders, S., Pyl, P. T. & Huber, W. Genome analysis HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. 9, 559 (2008).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550–550 (2014).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2012).
Guillou, L. et al. The Protist Ribosomal Reference database (PR2): a catalog of unicellular eukaryote small sub-unit rRNA sequences with curated taxonomy. Nucleic Acids Res. 41, 597–604 (2013).
Bolyen, E. et al. Reproducible, interactive, scalable, and extensible microbiome data science using QIIME2. Nat. Biotechnol. 37, 852–857 (2019).
Lee, M. D. GToTree: a user-friendly workflow for phylogenomics. Bioinformatics 35, 4162–4164 (2019).
Acknowledgements
We thank C. Preston, J. Birch, C. Sholin and the MBARI ESP team for providing field sampling infrastructure and expertise; S. Sharma for providing bioinformatic assistance; C. Smith, C. Thomas and K. Esson for assisting with field and laboratory techniques; R. Michisaki for providing expertise on microbial biomass estimates; and the University of Georgia Genomics and Bioinformatics Core for supplying sequencing services. This work was supported by the Simons Foundation (grant no. 542391 to M.A.M.) within the Principles of Microbial Ecosystems (PriME) Collaborative and by NSF (IOS-1656311). The rRNA amplicon sequencing was provided through the DOE Joint Genome Institute Community Sequencing Program.
Author information
Authors and Affiliations
Contributions
B.N. and M.A.M. conceived of the study. B.N. collected the data. B.N. and M.A.M. analysed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Microbiology thanks Virginia Armbrust and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Protist and bacterial community composition during the 2016 Monterey Bay autumn bloom based on rRNA gene sequencing.
Each bar represents 1 replicate sample. a) 18S rRNA, order level. b) 18S rRNA, genus level. c) 16S rRNA, family level.
Extended Data Fig. 2
Salinity and temperature of seawater sampled by CTD at Monterey Bay Station MO in Fall, 2016.
Extended Data Fig. 3 Correlations of Z-score normalized R. pomeroyi gene expression module eigengenes with environmental data.
Expression was measured by RNAseq after incubation in seawater collected on 14 dates at Monterey Bay Station M0 in Fall, 2016. Dinoflagellate parasites are members of the Syndinales clade. Cells are colored by Pearson’s R parameter. Two stars indicate correlations at p < 0.01; one star indicates correlations at p < 0.05. d.f. = 12.
Extended Data Fig. 4 Correlations of representative R. pomeroyi genes for transport of ammonium (amtB) and urea (urtB) with Akashiwo biomass.
The extreme value of Akashiwo biomass is removed in the inset. **, significant Pearson’s R correlation, with p < 0.001 for amtB and urtB in the main figure, d.f. = 12; and p = 0.010 for amtB and p = 0.331 for umtB in the inset figure, d.f. = 11.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2.
Source data
Source Data Fig. 1
Microbial abundance and Chl a, gene expression data for glucose catabolism genes.
Source Data Fig. 2
Gene expression data for PCA, 18S rRNA amplicon relative abundance and mean z-scores for gene modules.
Source Data Fig. 3
Gene expression data by date and replicate.
Source Data Fig. 4
Gene expression data for ribosomal proteins, rpoD and dmdA, genome accession nos. and DMSP concentrations.
Source Data Extended Data Fig. 1
The 18S rRNA ASV abundance at the order level, 18S rRNA ASV abundance at the genus level and 16S rRNA ASV abundance at the family level.
Source Data Extended Data Fig. 2
Temperature and salinity data.
Source Data Extended Data Fig. 3
Environmental data and module eigengenes.
Source Data Extended Data Fig. 4
Nitrogen gene expression data and A. sanguinea biomass data.
Rights and permissions
About this article
Cite this article
Nowinski, B., Moran, M.A. Niche dimensions of a marine bacterium are identified using invasion studies in coastal seawater. Nat Microbiol 6, 524–532 (2021). https://doi.org/10.1038/s41564-020-00851-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-020-00851-2
This article is cited by
-
Progress and challenges in exploring aquatic microbial communities using non-targeted metabolomics
The ISME Journal (2023)
-
Functional annotation and importance of marine bacterial transporters of plankton exometabolites
ISME Communications (2023)
-
Growth-stage-related shifts in diatom endometabolome composition set the stage for bacterial heterotrophy
ISME Communications (2022)
-
Diel investments in metabolite production and consumption in a model microbial system
The ISME Journal (2022)
-
Rapid bacterioplankton transcription cascades regulate organic matter utilization during phytoplankton bloom progression in a coastal upwelling system
The ISME Journal (2022)