Abstract
The most frequent fetal birth defect associated with prenatal Zika virus (ZIKV) infection is brain calcification, which in turn may potentially affect neurological development in infants. Understanding the mechanism could inform the development of potential therapies against prenatal ZIKV brain calcification. In perivascular cells, bone morphogenetic protein (BMP) is an osteogenic factor that undergoes maturation to activate osteogenesis and calcification. Here, we show that ZIKV infection of cultivated primary human brain pericytes triggers BMP2 maturation, leading to osteogenic gene expression and calcification. We observed extensive calcification near ZIKV+ pericytes of fetal human brain specimens and in vertically transmitted ZIKV+ human signal transducer and activator of transcription 2-knockin mouse pup brains. ZIKV infection of primary pericytes stimulated BMP2 maturation, inducing osteogenic gene expression and calcification that were completely blocked by anti-BMP2/4 neutralizing antibody. Not only did ZIKV NS3 expression alone induce BMP2 maturation, osteogenic gene expression and calcification, but purified NS3 protease also effectively cleaved pro-BMP2 in vitro to generate biologically active mature BMP2. These findings highlight ZIKV-induced calcification where the NS3 protease subverts the BMP2-mediated osteogenic signalling pathway to trigger brain calcification.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The gene expression data presented in Extended Data Fig. 2h of this manuscript were reanalysed from RNA-seq data reported by Oh et al.25 and deposited with the Gene Expression Omnibus under accession no. GSE87750. Data supporting the findings of this study are available from the corresponding author upon reasonable request. All RT–qPCR primers and probe sequences used in this study are listed in Supplementary Table 1. Source data are provided with this paper.
References
Bonaldo, M. C. et al. Isolation of infective Zika virus from urine and saliva of patients in Brazil. PLoS Negl. Trop. Dis. 10, e0004816 (2016).
Brasil, P. et al. Zika virus infection in pregnant women in Rio de Janeiro. N. Engl. J. Med. 375, 2321–2334 (2016).
Cao-Lormeau, V.-M. et al. Guillain-Barré syndrome outbreak associated with Zika virus infection in French Polynesia: a case-control study. Lancet 387, 1531–1539 (2016).
Duffy, M. R. et al. Zika virus outbreak on Yap Island, Federated States of Micronesia. N. Engl. J. Med. 360, 2536–2543 (2009).
Wang, L. et al. From mosquitos to humans: genetic evolution of Zika virus. Cell Host Microbe 19, 561–565 (2016).
Phoo, W. W. et al. Structure of the NS2B-NS3 protease from Zika virus after self-cleavage. Nat. Commun. 7, 13410 (2016).
Wu, Y. et al. Zika virus evades interferon-mediated antiviral response through the co-operation of multiple nonstructural proteins in vitro. Cell Discov. 3, 17006 (2017).
Ding, Q. et al. Species-specific disruption of STING-dependent antiviral cellular defenses by the Zika virus NS2B3 protease. Proc. Natl Acad. Sci. USA 115, E6310–E6318 (2018).
Smith, D. R. et al. Neuropathogenesis of Zika virus in a highly susceptible immunocompetent mouse model after antibody blockade of type I interferon. PLoS Negl. Trop. Dis. 11, e0005296 (2017).
Gorman, M. J. et al. An immunocompetent mouse model of Zika virus infection. Cell Host Microbe 23, 672–685 (2018).
Soares de Oliveira-Szejnfeld, P. et al. Congenital brain abnormalities and Zika virus: what the radiologist can expect to see prenatally and postnatally. Radiology 281, 203–218 (2016).
Li, H., Saucedo-Cuevas, L., Shresta, S. & Gleeson, J. G. The neurobiology of Zika virus. Neuron 92, 949–958 (2016).
Pool, K.-L. et al. Association between neonatal neuroimaging and clinical outcomes in Zika-exposed infants from Rio de Janeiro, Brazil. JAMA Netw. Open 2, e198124 (2019).
Mulkey, S. B. et al. Sequential neuroimaging of the fetus and newborn with in utero Zika virus exposure. JAMA Pediatr. 173, 52–59 (2019).
Speer, M. Y. et al. Smooth muscle cells give rise to osteochondrogenic precursors and chondrocytes in calcifying arteries. Circ. Res. 104, 733–741 (2009).
Felin, J. E. et al. Nuclear variants of bone morphogenetic proteins. BMC Cell Biol. 11, 20 (2010).
Pillai, I. C. L. et al. Cardiac fibroblasts adopt osteogenic fates and can be targeted to attenuate pathological heart calcification. Cell Stem Cell 20, 218–232 (2017).
Fu, H. et al. A tau homeostasis signature is linked with the cellular and regional vulnerability of excitatory neurons to tau pathology. Nat. Neurosci. 22, 47–56 (2019).
Dubrac, A. et al. NCK-dependent pericyte migration promotes pathological neovascularization in ischemic retinopathy. Nat. Commun. 9, 3463 (2018).
Montagne, A. et al. Pericyte degeneration causes white matter dysfunction in the mouse central nervous system. Nat. Med. 24, 326–337 (2018).
Goswami, R. et al. Expression of osteogenic molecules in the caudate nucleus and gray matter and their potential relevance for basal ganglia calcification in hypoparathyroidism. J. Clin. Endocrinol. Metab. 99, 1741–1748 (2014).
Boström, K. et al. Bone morphogenetic protein expression in human atherosclerotic lesions. J. Clin. Invest. 91, 1800–1809 (1993).
Derwall, M. et al. Inhibition of bone morphogenetic protein signaling reduces vascular calcification and atherosclerosis. Arterioscler. Thromb. Vasc. Biol. 32, 613–622 (2012).
Liu, L. et al. Inhibition of vascular calcification. Arterioscler. Thromb. Vasc. Biol. 38, 2382–2395 (2018).
Oh, Y. et al. Zika virus directly infects peripheral neurons and induces cell death. Nat. Neurosci. 20, 1209–1212 (2017).
Corry, J., Arora, N., Good, C. A., Sadovsky, Y. & Coyne, C. B. Organotypic models of type III interferon-mediated protection from Zika virus infections at the maternal–fetal interface. Proc. Natl Acad. Sci. USA 114, 9433–9438 (2017).
Rausch, K. et al. Screening bioactives reveals nanchangmycin as a broad spectrum antiviral active against Zika virus. Cell Rep. 18, 804–815 (2017).
Lee, S. J. et al. Pyruvate dehydrogenase kinase 4 promotes vascular calcification via SMAD1/5/8 phosphorylation. Sci. Rep. 5, 16577 (2015).
Liang, Q. Zika virus NS4A and NS4B proteins deregulate Akt-mTOR signaling in human fetal neural stem cells to inhibit neurogenesis and induce autophagy. Cell Stem Cell 19, 663–671 (2016).
Cui, Y., Jean, F., Thomas, G. & Christian, J. L. BMP-4 is proteolytically activated by furin and/or PC6 during vertebrate embryonic development. EMBO J. 17, 4735–4743 (1998).
Leighton, M. & Kadler, K. E. Paired basic/furin-like proprotein convertase cleavage of pro-BMP-1 in the trans-Golgi network. J. Biol. Chem. 278, 18478–18484 (2003).
Künnapuu, J., Björkgren, I. & Shimmi, O. The Drosophila DPP signal is produced by cleavage of its proprotein at evolutionary diversified furin-recognition sites. Proc. Natl Acad. Sci. USA 106, 8501–8506 (2009).
Zhang, Z. et al. Crystal structure of unlinked NS2B-NS3 protease from Zika virus. Science 354, 1597–1600 (2016).
Dhore, C. R. et al. Differential expression of bone matrix regulatory proteins in human atherosclerotic plaques. Arterioscler. Thromb. Vasc. Biol. 21, 1998–2003 (2001).
Melo, A. S. et al. Congenital Zika virus infection: beyond neonatal microcephaly. JAMA Neurol. 73, 1407–1416 (2016).
Moore, C. A. et al. Characterizing the pattern of anomalies in congenital Zika syndrome for pediatric clinicians. JAMA Pediatr. 171, 288–295 (2017).
Driggers, R. W. et al. Zika virus infection with prolonged maternal viremia and fetal brain abnormalities. N. Engl. J. Med. 374, 2142–2151 (2016).
Yamada, M. et al. High frequency of calcification in basal ganglia on brain computed tomography images in Japanese older adults. Geriatr. Gerontol. Int. 13, 706–710 (2013).
Koyama, S. et al. Clinical and radiological diversity in genetically confirmed primary familial brain calcification. Sci. Rep. 7, 12046 (2017).
Livingston, J. H., Stivaros, S., Warren, D. & Crow, Y. J. Intracranial calcification in childhood: a review of aetiologies and recognizable phenotypes. Dev. Med. Child. Neurol. 56, 612–626 (2014).
Nielsen-Saines, K. et al. Delayed childhood neurodevelopment and neurosensory alterations in the second year of life in a prospective cohort of ZIKV-exposed children. Nat. Med. 25, 1213–1217 (2019).
Adachi, K. et al. Early clinical infancy outcomes for microcephaly and/or small for gestational age Zika-exposed infants. Clin. Infect. Dis. 70, 2663–2672 (2020).
Einspieler, C. et al. Association of infants exposed to prenatal Zika virus infection with their clinical, neurologic, and developmental status evaluated via the general movement assessment tool. JAMA Netw. Open 2, e187235 (2019).
Lopes Moreira, M. E. et al. Neurodevelopment in infants exposed to Zika virus in utero. N. Engl. J. Med. 379, 2377–2379 (2018).
Zin, A. A. et al. Screening criteria for ophthalmic manifestations of congenital Zika virus infection. JAMA Pediatr. 171, 847–854 (2017).
Petribu, N. C. L. et al. Follow-up brain imaging of 37 children with congenital Zika syndrome: case series study. BMJ 359, j4188 (2017).
Bayless, N. L., Greenberg, R. S., Swigut, T., Wysocka, J. & Blish, C. A. Zika virus infection induces cranial neural crest cells to produce cytokines at levels detrimental for neurogenesis. Cell Host Microbe 20, 423–428 (2016).
Tian, X., Brookes, O. & Battaglia, G. Pericytes from mesenchymal stem cells as a model for the blood–brain barrier. Sci. Rep. 7, 39676 (2017).
Crisan, M. et al. A perivascular origin for mesenchymal stem cells in multiple human organs. Cell Stem Cell 3, 301–313 (2008).
Zarb, Y. et al. Ossified blood vessels in primary familial brain calcification elicit a neurotoxic astrocyte response. Brain 142, 885–902 (2019).
Vanlandewijck, M. et al. A molecular atlas of cell types and zonation in the brain vasculature. Nature 554, 475–480 (2018).
Crisan, M. et al. Perivascular multipotent progenitor cells in human organs. Ann. N. Y. Acad. Sci. 1176, 118–123 (2009).
Xu, J. et al. Human perivascular stem cell-derived extracellular vesicles mediate bone repair. eLife 8, e48191 (2019).
Cataneo, A. H. D. et al. The citrus flavonoid naringenin impairs the in vitro infection of human cells by Zika virus. Sci. Rep. 9, 16348 (2019).
Goodfellow, F. T. et al. Strain-dependent consequences of Zika virus infection and differential impact on neural development. Viruses 10, 550 (2018).
Simonin, Y., van Riel, D., Ven de Perre, P., Rockx, B. & Salinas, S. Differential virulence between Asian and African lineages of Zika virus. PLoS Negl. Trop. Dis. 11, e0005821 (2017).
Foo, S.-S. Asian Zika virus strains target CD14+ blood monocytes and induce M2-skewed immunosuppression during pregnancy. Nat. Microbiol. 2, 1558–1570 (2017).
Tripathi, S. et al. A novel Zika virus mouse model reveals strain specific differences in virus pathogenesis and host inflammatory immune responses. PLoS Pathog. 13, e1006258 (2017).
Hillger, F., Herr, G., Rudolph, R. & Schwarz, E. Biophysical comparison of BMP-2, proBMP-2, and the free pro-peptide reveals stabilization of the pro-peptide by the mature growth factor. J. Biol. Chem. 280, 14974–14980 (2005).
Tang, H. et al. Zika virus infects human cortical neural progenitors and attenuates their growth. Cell Stem Cell 18, 587–590 (2016).
Yao, M. et al. BMP2-SMAD signaling represses the proliferation of embryonic neural stem cells through YAP. J. Neurosci. 34, 12039–12048 (2014).
Nakashima, K. et al. BMP2-mediated alteration in the developmental pathway of fetal mouse brain cells from neurogenesis to astrocytogenesis. Proc. Natl Acad. Sci. USA 98, 5868–5873 (2001).
Cheng, H. et al. Osteogenic activity of the fourteen types of human bone morphogenetic proteins (BMPs). J. Bone Joint Surg. Am. 85, 1544–1552 (2003).
Zhang, M., Ngo, J., Pirozzi, F., Sun, Y.-P. & Wynshaw-Boris, A. Highly efficient methods to obtain homogeneous dorsal neural progenitor cells from human and mouse embryonic stem cells and induced pluripotent stem cells. Stem Cell Res. Ther. 9, 67 (2018).
Corselli, M. et al. Perivascular support of human hematopoietic stem/progenitor cells. Blood 121, 2891–2901 (2013).
Acknowledgements
This work was partly supported by: grant no. K99DE028573 to W.C.; grant nos. CA200422, CA251275, AI140705, AI140705S, AI140718, AI152190, AI116585, AI116585S, DE023926, DE027888, DE028521 and KGM9942011 to J.U.J.; grant nos. AI069120, AI056154, AI078389 and AI28697 to G.C.; grant nos. AI129534-01, AI298847-01 to K.N.S.; grant nos. AI140718 to J.U.J., G.C. and K.N.S.; the Departamento de Ciência e Tecnologia (DECIT/25000.072811/2016-17) do Ministério da Saúde do Brasil and Coordenacao de Aperfeicoamento de Pessoal de Nivel Superior CAPES/ 88887.116627/2016-01 (P.B.); grant nos. R01AR071734 and R01AG058624 to D.E.; and National Institutes of Health/National Institute of Allergy and Infectious Diseases grant nos. U19AI118610 and R21AI129486 to A.G.‐S.
Author information
Authors and Affiliations
Contributions
W.C. and J.U.J. designed the experiments. W.C. and S.-S.F. performed the experiments and analysed the data. W.C. prepared the figures and wrote the manuscript. E.H., C.W., S.-A.L. and W.-S.L. assisted with the experiments. D.E., M.E.L.M., A.G.-S., K.N.-S., P.B., G.C. and E.A.-P. provided expertise and tools for the study. J.U.J. edited the manuscript and oversaw all study design and data analysis. All other authors read the manuscript and provided comments.
Corresponding author
Ethics declarations
Competing interests
J.U.J. is a scientific adviser of the Vaccine Stabilization Institute, a California corporation. The remaining authors declare no competing interests.
Additional information
Peer review information Nature Microbiology thanks Laurent Nguyen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Perinatal ZIKV infection in brain tissues of deceased fetuses exhibits pronounced calcifications.
a, Paraffin-embedded brain tissue sections (10μm) from ZIKV-positive deceased human fetuses were stained with Von Kossa (red arrow). b, Immunohistochemical staining for NeuN (neurons), PDGFRβ (pericytes) or GFAP (astrocytes) were performed using sequential brain sections of ZIKV-positive human fetal brain tissue. Arrows indicate NeuN-, PDGFRβ- or GFAP-positive cells (c) RNAScope duplex in-situ hybridization with ZIKV RNA and PDGFRβ mRNA of healthy human fetal brain tissues. Black arrows indicate PDGFRβ-positive cells lining the blood vessels. Neither non-specific staining ZIKV RNA nor calcium deposit was detected in healthy tissues. Data in a-c are examined from biologically independent human specimens (healthy n = 3; ZIKV n = 5) and are representative of two independent repeats.
Extended Data Fig. 2 Asian ZIKV infection, but not African ZIKV elicits in vitro calcification in pericytes.
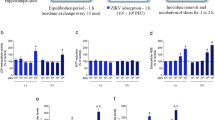
Human fetal pericytes were infected with different ZIKV strains at MOI of 0.5 or mock (PBS) as control (n = 4 biologically independent cells per group). a, At 3 dpi, cell viability was quantified. See Supplementary Data Fig. 1 for gating strategy. b, BMP4, BMP6, BMP7, BMP9 and NOG expression between ZIKV MR766- and ZIKV H/PF/2013-infected pericytes at 1, 3 and 4 dpi (n = 6 biologically independent cells per group). Data in a and b are presented as mean ± SEM in box plots showing the upper (75%) and lower (25%) quartiles, with the horizontal line as the median and the whiskers as the maximum and minimum values observed. c, Osteogenic gene expression between ZIKV MR766- and ZIKV IbH30656-infected pericytes was normalized to GAPDH and expressed as fold change relative to mock controls (n = 3 biologically independent cells per group). Data are presented as mean ± SEM. d, At 8 and 14 dpi, mock- or ZIKV-infected human fetal pericytes were subjected to Alizarin red staining for calcium deposition. Data are representative of two independent experiments. e-f, Alizarin red staining were performed on mock- or ZIKV-infected SK-N-SH and U251 at 14 and 21 dpi. Data are representative of four independent experiments. g, Human primary fetal pericytes (n = 5) and astrocytes (n = 6) were infected with PBS (mock) or ZIKV PRVABC59 at MOI of 0.5. At 3 dpi, osteogenic gene expressions were normalized to GAPDH and expressed as fold changes relative to mock controls. Data are presented as mean ± SEM and are representative of two independent experiments. h, RNAseq data generated from Nature Neuroscience; 2017; 20(9); p1209-1212 10.1038/nn.4612 (GEO ID: GSE87750) was reanalyzed for osteogenic gene expression. Human primary peripheral neurons were infected with ZIKV PRVABC59 of MOI 0.4 and harvested at 3 dpi. Statistical analyses were performed using one-way ANOVA with Tukey’s posttest (a and c). ***P < 0.001 and ****P < 0.0001. Exact P values in a compared between ZIKV MR766 and ZIKV H/PF/2013 group (P = 0.0008), ZIKV MR766 and ZIKV PRVABC59 group (P = 0.0005), ZIKV IBH30656 and ZIKV H/PF/2013 group (P = < 0.0001) and ZIKV IBH30656 and ZIKV PRVABC59 group (P = < 0.0001).
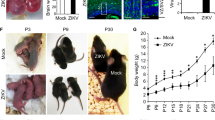
Extended Data Fig. 3 ZIKV infection of 3-weeks-old human STAT2 knockin mice resulted in brain calcification.
Three-week-old hSTAT2KI were s.c. infected with ZIKV. a, At 8 dpi, ZIKV RNA was quantified in the brain (MR766 n = 7; PRVABC59 n = 5). b, Osteogenic gene expression in mock- or ZIKV-infected hSTAT2KI brains (PBS n = 5; MR766 n = 6; PRVABC59 n = 6) were normalized to GAPDH and presented as fold changes relative to mock controls. c, Brain sections were stained with H&E or Alizarin red for calcium deposit. Black dotted line in the magnified insert indicates vasculature and blue dotted line in the magnified insert indicates prominent calcification sites. Data are presented as mean ± SEM, using Mann-Whitney U-test (a) or two-way ANOVA followed by Tukey’s multiple comparisons test (b). *P < 0.05, **P < 0.01 and ***P < 0.001. Exact P values in a compared between ZIKV MR766 and ZIKV PRVABC59 group (P = 0.0051). Data in a-c are representative of three independent experiments.
Extended Data Fig. 4 ZIKV infection of osteoblastic-like cells increase osteogenic gene expression.
a-b, Mock- and ZIKV-infected U2OS cells were harvested at 2 and 4 dpi respectively for osteogenic gene expression. c, Band intensity of pSMAD1/5/9 from U2OS whole cell lysate. d, At day 1-4, IgG or nAb-treated and mock or ZIKV-infected U2OS cells (n = 6) were harvested for RNA extraction and viral load against ZIKV NS1 RNA was quantified using qRT-PCR (N.D.; Not detected) (e) Human primary fetal brain pericytes are infected with ZIKV PRVABC59 or PBS (mock control) that were treated with 2 μg/mL of mouse IgG isotype control or human BMP2/4 neutralizing antibody. At 4 dpi, cells were harvested for osteogenic gene expressions, normalized to GAPDH and presented as fold changes relative to mock controls (n = 4/group). Data in a, b, d and e are presented as mean ± SEM in box plots showing the upper (75%) and lower (25%) quartiles, with the horizontal line as the median and the whiskers as the maximum and minimum values observed. Data are analyzed using Kruskal-Wallis test followed by Dunn’s multiple comparisons (a), Mann-Whitney U-test (b), or two-tailed unpaired Student t-test (e). *P < 0.05, **P < 0.01, ***P < 0.001. Exact P values in b compared between Mock and ZIKV H/PF/2013 (BMP2 P = 0.0002; RUNX2 P = 0.0011; OSX P = 0.0002; ALPL P = 0.0002; DMP1 P = 0.0002 and PDPN P = 0.0002). Exact P values in e compared between IgG-treated and Nab-treated group (BMP2 P = 0.0054; RUNX2 P = 0.0005; OSX P = 0.0085; DMP1 P = 0.0099 and PDPN P = 0.0005). Data are representative of two independent experiments.
Extended Data Fig. 5 Overexpression of ZIKV NS3 protease induced osteogenic gene expression in osteoblastic-like cells.
U2OS cells (n = 6) were transiently transfected with plasmids encoding individual ZIKV genes for 3 days. Transfected cells were harvested for qRT-PCR analysis of osteogenic gene expressions. Gene expressions were normalized to GAPDH and expressed as fold changes relative to vector control. Data are presented as mean ± SEM in box plots showing the upper (75%) and lower (25%) quartiles, with the horizontal line as the median and the whiskers as the maximum and minimum values observed. Data are representative of three independent experiments.
Extended Data Fig. 6 ZIKV NS3 protease is highly conserved across different lineages.
a, Sequence alignment comparison between BMP2 gene in human (homo sapiens; amino acid residues 1-396) and hamster (Cricetulus; amino acid residues 1-399). b, Purity of BL21 strain-derived recombinant ZIKV NS3 protease, CHO-FD11 cells-derived full-length BMP2 and BMP4 were determined by coomassie blue stain. MW: molecular weight. c, Sequence alignment comparison between human BMP2 (amino acid residues 1-396) and BMP4 (amino acid residues 1-408). d, Sequence alignment of NS3 protease domain (amino acid residues 1-177) was compared across two African ZIKV (MR766, IbH30656) and two Asian ZIKV (PRVABC59 and H/PF/2103). The red-colored letters indicate protease catalytic triad. e, In vitro cleavage assay of carboxyl terminal HA-tagged purified human BMP4 and purified ZIKV NS3 protease were performed at 37 °C for 3 h, followed by immunoblot analysis. Data in b and e are representative of three independent repeats.
Supplementary information
Source data
Source Data Fig. 2
Numerical source data.
Source Data Fig. 3
Numerical source data.
Source Data Fig. 4
Numerical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Numerical source data.
Source Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 2
Numerical source data.
Source Data Extended Data Fig. 3
Numerical source data.
Source Data Extended Data Fig. 4
Numerical source data.
Source Data Extended Data Fig. 5
Numerical source data.
Source Data Extended Data Fig. 6
Unprocessed western blots.
Rights and permissions
About this article
Cite this article
Chen, W., Foo, SS., Hong, E. et al. Zika virus NS3 protease induces bone morphogenetic protein-dependent brain calcification in human fetuses. Nat Microbiol 6, 455–466 (2021). https://doi.org/10.1038/s41564-020-00850-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-020-00850-3
This article is cited by
-
Mechanistic insights into bone remodelling dysregulation by human viral pathogens
Nature Microbiology (2024)
-
ZIKV actively induces calcification in the fetal brain
Nature Microbiology (2021)