Abstract
Pattern recognition receptors (PRRs) expressed in antigen-presenting cells are thought to shape pathogen-specific immunity by inducing secretion of costimulatory cytokines during T-cell activation, yet data to support this notion in vivo are scarce. Here, we show that the cytosolic PRR Nod-like Receptor CARD 4 (NLRC4) suppresses, rather than facilitates, effector and memory CD4+ T-cell responses against Salmonella in mice. NLRC4 negatively regulates immunological memory by preventing delayed activation of the cytosolic PRR NLR pyrin domain 3 (NLRP3) that would otherwise amplify the production of cytokines important for the generation of Th1 immunity such as intereukin-18. Consistent with a role for NLRC4 in memory immunity, primary challenge with Salmonella expressing flagellin modified to largely evade NLRC4 recognition notably increases protection against lethal rechallenge. This finding suggests flagellin modification to reduce NLRC4 activation enhances protective immunity, which could have important implications for vaccine development against flagellated microbial pathogens.
Similar content being viewed by others
Main
Antigen-presenting cells (APCs) recognize pathogens via pattern recognition receptors (PRRs), such as the transmembrane Toll-like receptors (TLRs) and the cytosolic nucleotide oligomerization domain (NOD)-like receptors (NLRs)1,2. PRR activation induces APCs to secrete proinflammatory cytokines during presentation of antigens to naïve T cells leading to their differentiation into the most appropriate subset of activated T cells to eliminate the pathogen3. The bridge between the innate and adaptive immune systems is, therefore, formed by PRR activation in APCs. Central to this is the concept that the repertoire of PRRs expressed by APCs is crucial to the specificity of adaptive immune response generated4. The increasing prevalence of antimicrobial resistance means the development of alternative ways to combat infectious diseases is a matter of urgency. Understanding the interactions between pathogens, PRRs and pathogen-specific immunity is, therefore, important to facilitate the development of new vaccines with improved efficacy.
Microbial pathogens produce a number of pathogen-associated molecular patterns that are recognized by different PRRs. Lipopolysaccharide (LPS) from Gram-negative bacteria, for example, is recognized by TLR4 and flagellin by TLR5. Both flagellin and detoxified LPS molecules are used as vaccine adjuvants with flagellin being an antigenic target of antibody and CD4+ T-cell responses5. The NOD-like receptor CARD 4 (NLRC4) also recognizes microbial flagellin6,7, but, in addition, senses the type III secretion system-associated needle and rod proteins8 via the NLR family, apoptosis inhibitory protein (NAIP) 5 or NAIP6 and NAIP1 or NAIP2 in mice9,10,11. NLR pyrin domain 3 (NLRP3) is activated by a diverse range of triggers, including pathogen-associated molecular patterns and danger-associated molecular patterns. NLR activation drives the formation of an inflammasome signalling complex12, which activates caspase-1 to process pro-IL-1β and pro-IL-18 to their mature forms and cleaves gasdermin D to drive an inflammatory form of cell death called pyroptosis13,14. NLRC4 and NLRP3 can be recruited to the same inflammasome to optimize IL-1β and IL-18 release15,16.
IL-1β and IL-18 have well-documented effects on T-cell survival, activation and differentiation17. Little is known, however, about whether and how specific inflammasome components influence the development of adaptive immunity against pathogenic bacteria and, specifically, that of immunological memory, which is critical for vaccine development4. It has been shown, for example, that NLRC4 and NLRP3 activation in innate immune cells optimizes interferon-γ (IFN-γ) release by CD8+ and CD4+ T cells in mice challenged with Salmonella enterica serovar Typhimurium (S. Typhimurium), but this was not an antigen-specific effect18,19. Here, using the murine model of sublethal salmonellosis, we show that NLRC4 suppresses Th1 CD4+ T-cell effector and memory responses against S. Typhimurium without affecting antibody production. Primary challenge with S. Typhimurium engineered to reduce NLRC4 inflammasome activation resulted in increased levels of serum IL-18 and IFN-γ, enhanced Th1 CD4+ T-cell memory responses and improved protection against oral rechallenge with fully virulent Salmonella. Our data indicate that attenuation of NLRC4 activation by flagellin could improve the efficacy of live vaccines against flagellated pathogens such as Salmonella, and potentially that of flagellin-based adjuvants.
Results
NLRC4 restricts bacterial load in vivo and Th1 memory responses ex vivo
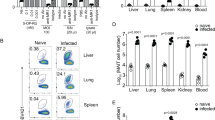
To investigate whether NLRC4 affects the development of pathogen-specific immunity, we first challenged C57BL/6J wild-type and congenic Nlrc4−/− mice with S. Typhimurium M525P. In this model of sublethal salmonellosis, effector Th1 CD4+ T cells drive microbial clearance and establish long-term protection against rechallenge20,21. Nlrc4−/− mice harboured higher microbial numbers in the liver and spleen irrespective of the route of bacterial administration (intravenous (i.v.) in Fig. 1a or oral in Extended Data Fig. 1a). Bacterial burden, as expected22,23, was not significantly different between Nlrc4−/− and wild-type mice lethally challenged with fully virulent S. Typhimurium SL1344 (Extended Data Fig. 1b). The slower kinetics of the sublethal infection reveal a clear role for NLRC4 in regulating bacterial load in vivo.
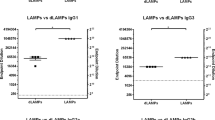
a, Wild-type and Nlrc4−/− mice were challenged i.v. with 9.5 × 103 c.f.u. S. Typhimurium M525P and microbial burden was determined in the spleen and liver at day 1 (n = 5 for wild-type and Nlrc4−/−), 7 (n = 7 for wild-type and n = 6 for Nlrc4−/−), 14, 21 and 35 (n = 6 for wild-type and Nlrc4−/−) post-challenge. b, CD4+ T cells from naïve wild-type and Nlrc4−/− mice were stimulated ex vivo with medium only (without (w/o) stim.), antimouse CD3e (anti-CD3e) or whole bacterial cell extract (extract). Levels of IFN-γ and IL-2 were measured by ELISA in the cell culture supernatant after 72 and 24 h, respectively. c, CD4+ T cells from wild-type and Nlrc4−/− mice 90 d after challenge with 7 × 103 − 1 × 104 c.f.u. of S. Typhimurium M525P were stimulated and levels of IFN-γ and IL-2 were measured as in b. d, CD4+ T cells from wild-type and Nlrc4−/− mice 168 d after challenge with 1.15 × 103 c.f.u. of S. Typhimurium M525P were stimulated as in b, with the exception that concanavalin A (conA) was used instead of antimouse CD3e, and levels of IFN-γ and IL-2 were measured as in b. e, CD4+ T cells from wild-type and Nlrc4−/− mice 90 d after challenge with 7 × 103 c.f.u. of S. Typhimurium M525P were stimulated with whole bacterial cell extract and levels of IL-6, TNF-α and GM-CSF were measured by fluorescent bead immunoassay in the cell culture supernatant after 24 h. f, Same experiment as e, the number of CD4+ T cells secreting IFN-γ in response to whole bacterial cell extract was measured via ELISpot. SFCs, spot-forming cells. g, Wild-type and Nlrc4−/− mice were challenged i.v. with 1.72 × 104 c.f.u. and 112 d later, the percentage of CD4+ T cells secreting IFN-γ in response to whole bacterial cell extract was measured via a cytokine secretion assay. Each symbol represents one mouse with horizontal lines delineating the mean in a. Data are shown as mean ± s.e.m. in b–g. Data are pooled from two and representative of at least four independent experiments for c and representative of three for a and two for b,d–g independent experiments. Statistical significance was calculated by a two-way ANOVA followed by Sidak’s multiple comparisons tests for a–d and two-tailed Mann–Whitney test for e–g.
We determined whether NLRC4 would play any role in the development of memory CD4+ T-cell responses. Wild-type and Nlrc4−/− mice were challenged with PBS (naïve) or 104 colony-forming units (c.f.u.) of S. Typhimurium M525P, an inoculum size that was equally immunogenic to a tenfold higher dose in wild-type mice (Extended Data Fig. 2) and well tolerated by the Nlrc4−/− mice. To elicit a response from memory cells only, CD4+ T cells were isolated from mice that had cleared the primary infection after a minimum of 90 d. On stimulation with bacterial extract (antigen-specific), CD4+ T cells from naïve mice, irrespective of genotype, secreted negligible amounts of the Th1 cytokines IFN-γ and IL-2 (Fig. 1b), confirming the specificity of the response generated by CD4+ T cells from infected mice (Fig. 1c). Similarly, the amount of IFN-γ secreted by CD4+ T cells when stimulated with antimouse CD3e (non-antigen-specific) increased more than 50-fold in response to infection (Fig. 1c).
We compared the magnitude of the ex vivo memory responses between wild-type and Nlrc4−/− mice. T cells from Nlrc4−/− mice secreted more IFN-γ and IL-2 than cells from wild-type mice in response to both Salmonella antigen and antimouse CD3e (Fig. 1c). This amplified response persisted up to 170 d after challenge (Fig. 1d, concanavalin A was used in early experiments as a polyclonal stimulus but later replaced by antimouse CD3e). The Th1-related cytokines TNF-α, IL-6 and GM-CSF in the CD4+ T-cell culture supernatant from Nlrc4−/− mice were also elevated (Fig. 1e). These data indicate that NLRC4 activation by Salmonella specifically restricts CD4+ T-cell-mediated memory immunity.
We then quantified the number of CD4+ T cells secreting IFN-γ in response to bacterial extract using ELISpot and cytokine secretion assays. Both assays showed a larger population of IFN-γ-secreting cells in Nlrc4−/− mice (Fig. 1f,g). We also analysed the surface expression of CD62L and CD44 in IFN-γ+cells and found that over 90% in both wild-type and Nlrc4−/− mice had an effector memory T-cell profile (CD44high CD62L−) with central memory cells (CD44high CD62L+) representing less than 5% of the population (Extended Data Fig. 3). These data indicate that NLRC4 restricts T-cell memory immunity against Salmonella by reducing the size of antigen-specific memory pool.
To test whether NLRC4 could also affect protection against lethal rechallenge in vivo, wild-type and Nlrc4−/− mice were systemically challenged with S. Typhimurium M525P and, after clearing the primary infection, rechallenged orally with fully virulent Salmonella. Nlrc4−/− mice were only marginally better protected than their wild-type counterparts (Extended Data Fig. 4b, mean survival time of 9 versus 8 d, respectively), an outcome that may have been affected by the different routes of infection between primary challenge (i.v.) and rechallenge (orally). In the same trial, however, naïve Nlrc4−/− mice lethally challenged with Salmonella had shorter survival times than naïve wild-type mice (Extended Data Fig. 4a, 6 versus 8 d, respectively). These data indicate that, in the absence of NLRC4, the compromised innate immune control of microbial dissemination presumably counteracts any benefits arising from augmented T-cell memory immunity.
NLRC4 restricts Th1 effector responses during microbial clearance of a primary infection
To determine whether NLRC4 also affects the development of Th1 effector immunity, we isolated splenic CD4+ T cells from infected wild-type and Nlrc4−/− mice during an active primary infection and measured the amount of IFN-γ released ex vivo. Production of IFN-γ in response to bacterial extract was comparable between the two genotypes when CD4+ T cells were isolated early during infection (Figs. 2a, 1 and 7 d). Notably, as the infection progressed into the phase of CD4+ T-cell-mediated bacterial clearance, T cells from Nlrc4−/− mice secreted significantly more IFN-γ than cells from wild-type mice (Fig. 2a, 14 and 21 d). By the end of microbial clearance, however, there were no difference in Th1 responses between Nlrc4−/− and wild-type mice (Fig. 2a, 35 d). Salmonella-induced Th1 CD4+ T-cell responses can also affect the development of B-cell-dependent immunity24, so we measured the levels of serum antibodies against S. Typhimurium LPS at 90 d after primary infection. We found no difference in the levels of IgG2c, IgG2b and IgM between wild-type and Nlrc4−/− mice (Fig. 2b). These data collectively show that NLRC4 restricts both Th1 CD4+ T-cell-mediated effector and memory immune responses against Salmonella, but has no effect on B-cell-mediated immunity.
a, Wild-type and Nlrc4−/− mice were challenged i.v. with 7.2 × 103 − 1 × 104 c.f.u. of S. Typhimurium M525P and T-cell-mediated responses were analysed over time. CD4+ T cells were cocultured ex vivo with feeder cells in the presence of medium only (w/o stim.), antimouse CD3e (anti-CD3e) or whole bacterial cell extract (extract) and IFN-γ was measured by ELISA in the cell culture supernatant after 72 h. b, Wild-type and Nlrc4−/− mice were challenged i.v. with 7 × 103 c.f.u. S. Typhimurium M525P and levels of IgG2b, IgG2c and IgM antibodies against S. Typhimurium LPS were measured in the serum at 90 d post-challenge via ELISA. Data are shown as mean ± s.e.m. in a. Each symbol represents one mouse with horizontal lines delineating the mean in b. Statistical significance was assessed by a two-tailed Mann–Whitney test. Data are pooled from two independent experiments for a, day 7 and generated from one experiment in a, days 1, 14, 21 and 35. Data are representative of two independent experiments for b.
Negative regulation of CD4+ T-cell-mediated memory responses by NLRC4 is NLRP3-dependent
NLRC4 is not expressed in naïve or activated CD4+ T cells25,26. We proposed, therefore, that any effect on CD4+ T-cell memory immunity is most likely to occur in response to inflammasome signalling by APCs. NLRC4 and NLRP3 are both activated in murine macrophages infected with Salmonella22. We, therefore, assessed ex vivo the memory T-cell responses in Nlrp3−/− and Nlrp3−/−Nlrc4−/− mice. Secretion of both IFN-γ and IL-2 was comparable between wild-type, Nlrp3−/− or Nlrp3−/−Nlrc4−/− mice (Fig. 3a,b) and, as expected, we saw no difference in the numbers of CD4+ T cells secreting IFN-γ in response to bacterial extract (Fig. 3c,d). To confirm that our independent comparisons between wild-type and Nlrc4−/− (Fig. 1c), wild-type and Nlrp3−/− (Fig. 3a) and wild-type and Nlrp3−/−Nlrc4−/− mice (Fig. 3b) do not suffer from interexperimental variation, we compared IFN-γ production between the different groups of wild-type mice used in these experiments and found no significant difference (Extended Data Fig. 5, P = 0.25 via one-way analysis of variance (ANOVA)). These data indicate that the amplification of memory CD4+ T-cell responses seen the absence of Nlrc4−/− is driven by NLRP3 activation, while NLRP3 does not have an effect when NLRC4 is active.
The Th1 memory response was assessed by stimulating CD4+ T cells ex vivo with medium only (w/o stim.), antimouse CD3e (anti-CD3e), or whole bacterial cell extract (extract). a, CD4+ T cells from wild-type and Nlrp3−/− mice 100 d after challenge with 2.2 × 104 c.f.u. S. Typhimurium M525P and IFN-γ and IL-2 were measured by ELISA in the cell culture supernatant after 72 and 24 h, respectively. b, CD4+ T cells from wild-type and Nlrp3−/−Nlrc4−/− mice 90 d after challenge with 1.72 × 104 c.f.u. S. Typhimurium M525P and IFN-γ was measured as in a. c, Same experiment as a, the percentage of CD4+ T cells secreting IFN-γ in response to bacterial extract was determined via a cytokine secretion assay. d, Same experiment as b, the percentage of CD4+ T cells secreting IFN-γ was determined via a cytokine secretion assay. Data are shown as mean ± s.e.m. Statistical significance was calculated using a two-tailed, unpaired Student’s t-test assuming equal variances for a and c and a two-tailed Mann–Whitney test for b and d. All data are representative of two independent experiments.
NLRC4 restricts NLRP3-dependent production of IL-18 and IFN-γ
In macrophages infected with Salmonella SL1344, NLRC4 is activated rapidly to induce caspase-1-mediated pyroptosis and IL-1β and IL-18 release27, while NLRP3 activation is delayed to optimize IL-1β and IL-18 production without inducing pyroptosis15,22. Infection of wild-type and Nlrp3−/− macrophages with S. Typhimurium M525P resulted in pyroptotic cell death within 2 h of infection, while Nlrc4−/− and Nlrc4−/−Nlrp3−/− cells lysed between 6 and 12 h (Extended Data Fig. 6a). In the absence of NLRC4, cells produced IL-1β via NLRP3 and because these cells do not rapidly die, they secrete large amounts of IL-1β between 6 and 24 h (Extended Data Fig. 6b).
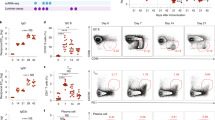
Both IL-1β and IL-18 are involved in initiation and propagation of antigen-specific immunity against bacterial pathogens, therefore, we examined whether NLRC4 and NLRP3 affect the production of these cytokines in vivo during infection with S. Typhimurium M525P. Serum IL-1β levels were below the detection limit, as expected22, throughout the course of infection. In Nlrc4−/− mice, levels of IL-18 were lower 1 d after challenge, but as the sublethal infection progressed, they markedly increased when compared to those detected in wild-type mice (Fig. 4b). This increase coincided with the onset of effector CD4+ T-cell-driven microbial clearance with IL-18 remaining elevated in Nlrc4−/− mice for a total of 3 weeks (Fig. 4b, days 7 to 28). This corresponds to the time that Nlrc4−/− mice have heightened effector CD4+ T-cell responses (Fig. 2a) suggesting a link between the increased levels of IL-18 and increased Th1 immunity.
Wild-type, Nlrp3−/−, Nlrc4−/− and Nlrp3−/−Nlrc4−/− mice were challenged i.v. with 1.86 × 104 c.f.u. S. Typhimurium M525P. a, Bacteria were counted in the spleen and the liver at day 7 (n = 5 for wild-type, Nlrp3−/− and Nlrp3−/−Nlrc4−/− and n = 4 for Nlrc4−/−), 21 (n = 6 for all genotypes) and 35 (n = 6 for wild-type, Nlrp3−/− and Nlrp3−/−Nlrc4−/− and n = 7 for Nlrc4−/−) post-challenge. b, IL-18 was measured in the serum at day 1 pre-challenge (n = 5 for wild-type and Nlrp3−/−Nlrc4−/−, n = 4 for Nlrp3−/− and n = 6 for Nlrc4−/−) and day 1 (n = 5 for wild-type and n = 6 for all other genotypes), 7 (n = 5 for wild-type, Nlrp3−/− and Nlrp3−/−Nlrc4−/− and n = 4 for Nlrc4−/−), 14 (n = 6 for wild-type and n = 5 for all other genotypes), 21 (n = 6 for all genotypes), 28 (n = 6 for wild-type, Nlrp3−/− and Nlrp3−/−Nlrc4−/− and n = 5 for Nlrc4−/−) and 35 (n = 6 for wild-type, Nlrp3−/− and Nlrp3−/−Nlrc4−/− and n = 7 for Nlrc4−/−) postchallenge by ELISA. c, IFN-γ was measured by fluorescent bead immunoassay in the serum over time with the same number of mice as in b apart from day 1 pre-challenge (n = 5 for wild-type and n = 4 for all other genotypes). Each symbol represents one mouse with horizontal lines delineating the mean in a. Data are shown as mean ± s.e.m. in b,c. Overall statistical significance has been determined by a one-way ANOVA for each interval separately. For wild-type versus Nlrc4−/− mice, data are representative of three independent experiments. For wild-type versus Nlrp3−/− and Nlrp3−/−Nlrc4−/− mice, data are representative of two independent experiments.
We tested whether this rise in serum IL-18 seen in Nlrc4−/− mice was NLRP3-dependent. Nlrp3−/− and Nlrp3−/−Nlrc4−/− mice, such as Nlrc4−/− mice, had decreased serum IL-18 levels very early during infection (Fig. 4b, day 1), but, unlike Nlrc4−/− mice, did not have increased IL-18 response as the infection progressed further (Fig. 4b, days 7–28). Nlrc4−/− and Nlrp3−/−Nlrc4−/− mice had similar bacterial burdens (Fig. 4a), but different IL-18 profiles (Fig. 4b), suggesting that this surge in serum IL-18 in the absence of NLRC4 depends on NLRP3 activation. IL-18 is a potent inducer of IFN-γ28, which is indispensable for protection against Salmonella20,29. IFN-γ levels in Nlrc4−/− mice remained elevated for 3 weeks compared to those in the Nlrp3−/−Nlrc4−/− mice (Fig. 4c, days 7–28) matching that of the serum IL-18 production. During the early stages of infection, cells such as natural killer cells are the main source of serum IFN-γ30 explaining why at day 7 Nlrc4−/− mice have higher levels of IFN-γ in the serum (Fig. 4c), but not when their CD4+ T cells are stimulated with bacterial extract ex vivo (Fig. 2a). Our data collectively indicate that, consistent with its role in downregulating CD4+ T-cell memory responses, NLRC4 activation suppresses NLRP3-driven amplification of serum IL-18 and IFN-γ.
S. Typhimurium expressing flagellin from non-pathogenic E. coli evades NLRC4 recognition
Flagellin activates both TLR5 and NAIP/NLRC4 with the latter also recognizing Salmonella type III secretory proteins such as PrgJ. TLR5 is important for generating T-cell immunity against S. Typhimurium31, but our data indicate that NLRC4 downregulates CD4+ T-cell-mediated memory, while, at the same time, enhancing host survival during the early stages of a lethal infection (Extended Data Fig. 4a). We considered that primary challenge with Salmonella genetically engineered to partially evade recognition by NLRC4, by mutating rather than deleting fliC, should retain TLR5 activation, enhance memory immunity and, therefore, potentially increase resistance to lethal rechallenge.
We genetically modified the wild-type M525P strain, which naturally lacks fljB, by replacing its native fliC with that from non-pathogenic E. coli K-12 substrain MG1655, to generate S. Typhimurium M525P ΔfliC::fliCMG1655 (M525P fliCMG1655). Comparison of the predicted FliC proteins from S. Typhimurium and E. coli K-12 MG1655 showed high sequence homology at the N- and C-terminal regions of the protein, but with a variable central domain (Extended Data Fig. 7). Flagellin from this E. coli strain activates TLR5 as efficiently as flagellin from S. Typhi but is impaired in activating NLRC4 (ref. 32). M525P fliCMG1655 also retains bacterial motility (Extended Data Fig. 8a), which may be important to facilitate infection and spread into the tissues. When tested in vitro, this mutant induced less cell death and IL-1β production than the wild-type strain in wild-type but not in Nlrc4−/− macrophages, but could still activate NAIP1/2-NLRC4 signalling presumably via its type III secretion system (Extended Data Fig. 8b,c). TNF-α production by wild-type macrophages infected with M525P fliCMG1655 or M525P were comparable (Extended Data Fig. 8d) suggesting no deficit in TLR activation. These data indicate that M525P fliCMG1655 shows reduced, but not obliterated, NLRC4 inflammasome recognition.
Wild-type mice infected with M525P fliCMG1655 had a higher bacterial burden than those infected with M525P (Fig. 5a), comparable to that seen in Nlrc4−/− mice challenged with M525P (Fig. 1a). Bacterial loads of M525P and M525P fliCMG1655 were similar in Nlrc4−/− mice confirming that the increased spread of M525P fliCMG1655 into the tissues was NLRC4-dependent (Fig. 5b). Serum levels of IL-18 and IFN-γ were significantly increased for 2 weeks (Fig. 5c,d, day 7 to 21) in mice infected with M525P fliCMG1655 when compared to mice infected with M525P, again comparable to the differences seen between wild-type and Nlrc4−/− mice infected with M525P (compare Fig. 4b,c with Fig. 5c,d). Infection of wild-type mice with M525P fliCMG1655 did not, however, induce as high levels, or as sustained production, of IL-18 as seen in Nlrc4−/− mice infected with M525P (compare Fig. 4b with Fig. 5c). This is consistent with M525P fliCMG1655 retaining some residual capacity to activate NLRC4.
Wild-type and Nlrc4−/− mice were challenged i.v. with 1.5 × 104 c.f.u. S. Typhimurium M525P or M525P fliCMG1655. a, Bacteria were counted in the spleen and the liver of wild-type mice over time (n = 6 for both groups for all days apart from day 35; n = 5 for M525P and n = 6 for M525P fliCMG1655). b, Bacteria were counted in the spleen and the liver of Nlrc4−/− mice over time (day 4 and 7; n = 4 for both groups, day 14 and 28; n = 3 for M525P and n = 4 for M525P fliCMG1655). c, IL-18 was measured in the serum of wild-type mice over time by ELISA with the same number of mice as in a. d, IFN-γ was measured in the serum of wild-type mice over time by a fluorescent bead immunoassay with the same number of mice as in a. Each symbol represents one mouse with horizontal lines delineate the mean in a,b. Data are shown as mean ± s.e.m. in c,d. Statistical significance was calculated by a two-way ANOVA followed by Sidak’s multiple comparisons tests in a and b and a two-tailed Mann–Whitney test for each time point separately in c and d. All data are representative of two independent experiments.
Primary challenge with S. Typhimurium M525P fliCMG1655 enhances CD4+ T-cell memory responses and improves protection against lethal rechallenge
We compared the CD4+ T-cell memory responses between wild-type mice challenged with M525P or M525P fliCMG1655 and found them to be higher in the M525P fliCMG1655 group (Fig. 6a), comparable to those seen in Nlrc4−/− mice (Fig. 1c). To determine whether primary challenge with S. Typhimurium M525P fliCMG1655 would confer any resistance against lethal rechallenge, wild-type mice were systemically infected with M525P or M525P fliCMG1655 and allowed to clear the infection before rechallenge with fully virulent SL1344. Mice initially infected with M525P survived for 8.5 d while the survival rate of mice infected with M525P fliCMG1655 was markedly increased with approximately 75% of them surviving up to 21 d postlethal rechallenge (Fig. 6b). Mice in the M525P fliCMG1655 group also lost less weight from days 4 to 6 after rechallenge when compared to mice in the M525P group (Fig. 6c). These data indicate that reduced NLRC4 activation during primary challenge promotes protection against rechallenge, although the mechanism by which this occurs is unclear.
a, CD4+ T cells from wild-type mice 122 d after challenge with 1.5 × 104 c.f.u. S. Typhimurium M525P or M525P fliCMG1655 were stimulated with medium only (w/o stim.), antimouse CD3e (anti-CD3e) or whole bacterial cell extract (extract). IFN-γ was measured by ELISA in the cell culture supernatant after 24 h. b, Wild-type mice were challenged with 1.6 × 104 c.f.u. S. Typhimurium M525P or 1.5 × 104 c.f.u. M525P fliCMG1655 and allowed to clear the primary infection for 128 d. They were then rechallenged orally with 4.8 × 106 c.f.u. S. Typhimurium SL1344 and euthanized on detection of adverse signs. NA, not applicable. c, In the same experiment as in b, mouse weight was recorded at least once daily and shown for the first 7 d of infection. d, CD4+ T cells obtained from wild-type and IL-18−/− mice 104 d after primary challenge with 1.6 × 104 c.f.u. S. Typhimurium M525P were stimulated and IFN-γ was measured as in a. e, CD4+ T cells obtained from wild-type and IL-18−/− mice 110 d after primary challenge with 3.3 × 104 c.f.u. S. Typhimurium M525P fliCMG1655 were stimulated and IFN-γ was measured as in a. Data are shown as mean ± s.e.m. in a,c–e and as a survival curve in b. Statistical significance was calculated by a two-tailed Mann–Whitney test in a, d and e and a two-sided log-rank in b. Data are representative of two independent experiments for a and were generated from one experiment for b–e.
Increased levels of serum IL-18 correlated with increased effector immunity (Figs. 2a and 4b), we, therefore, tested whether IL-18 contributes to the amplification of CD4+ T-cell immunity seen when NLRC4 activation is impaired. We compared the ex vivo memory responses between wild-type and IL-18−/−mice challenged with M525P and found no difference suggesting that, when NLRC4 is fully activated, IL-18 does not affect the potency of the response (Fig. 6d). When mice were challenged with M525P fliCMG1655, however, the IFN-γ response was reduced in IL-18−/− mice and, although this did not meet statistical significance (Fig. 6e, P = 0.0513) is, nevertheless, suggestive of a link between NLRC4, IL-18 and CD4+ T-cell-mediated Th1 memory.
Discussion
Pathogen-mediated activation of PRRs should determine the specificity of, and enhance the production of, antigen-specific immunity, yet here we show that activation of the PRR NLRC4 supresses memory CD4+ T-cell-mediated responses against Salmonella. The CD4+ T-cell-mediated immunity is critical for host control and long-term protection against intraphagosomal pathogens such as Salmonella. Equally important is the fact that mice initially challenged with a Salmonella strain that evades detection by NLRC4 were protected against an otherwise lethal rechallenge. This work, therefore, has important implications for vaccine development against pathogens that efficiently activate the NLRC4 inflammasome, such as Legionella33, Pseudomonas34 and Shigella35.
We used inflammasome-deficient mice to show that activation of NLRC4 by S. Typhimurium curtails antigen-specific CD4+ T-cell-mediated memory responses and confirmed this phenotype by using a Salmonella mutant defective in activating NLRC4. In the wild-type strain we replaced native flagellin with flagellin from E. coli that is impaired in activating NLRC4, but can still activate TLR5 and keep the mutant motile, both of which could potentially affect the development of immunity against Salmonella. This mutant enhanced antigen-specific immunity, most probably by its clear ability to evade NLRC4 activation, but it would be interesting to test if a flagellin-deficient mutant would induce similar effects in vivo.
Relatively little is known about whether NLRC4 can affect the development of antigen-specific immunity against pathogens. Studies investigating the effect of NLRC4 on CD8+ T-cell immunity have shown contrasting results. Listeria monocytogenes constitutively expressing flagellin from Legionella pneumophila suppressed CD8+ T-cell immunity in mice and failed to protect them against rechallenge36. Flagellin recognition by NLRC4 during Salmonella infection, in contrast, induces IFN-γ release by memory CD8+ T cells19. A similar phenomenon has been described in Salmonella-induced CD4+ Th1 effector cells that require both NLRC4 and NLRP3 to optimally secrete IFN-γ in response to LPS and flagellin18. Unlike our work, however, both these studies describe a role for NLRC4 in non-antigen-specific immune responses and suggest that NLRC4 activation drives the induction, rather than downregulation, of T-cell responses.
Our data show that NLRC4 activation negatively regulates CD4+ T-cell-mediated memory immunity against Salmonella via an NLRP3-dependent mechanism. Salmonella engages first NLRC4 and subsequently NLRP3, which maximizes IL-1β/IL-18 production in vitro15,22 and is particularly important when NLRC4 activation is impaired. Our work shows that mice lacking NLRC4 have increased serum IL-18 levels only when they have functional NLRP3, while deficiency in both PRRs prevents IL-18 amplification in the serum. We propose that during S. Typhimurium infection, NLRC4 prevents NLRP3 from amplifying cytokine production by APCs and this, in turn, restricts the potency of pathogen-specific immune responses. Defective NLRC4 activation by increasing the levels of Th1-related cytokines in the serum enhances the expansion of antigen-specific CD4+ T cells and, consequently, this would give rise to a larger pool of memory cells. These cytokines could either be produced by the APCs during their interaction with naive T cells or by cells other than APCs at a later stage to act on activated effector T cells4.
The NLRP3-dependent mechanism by which NLRC4 suppresses T-cell memory responses is unclear but our data indicate that IL-18 could be important. IL-18 enhances Th1 antigen-specific immunity by increasing IFNγ production from Th1 CD4+ T cells and natural killer cells37. It is also important for the maintenance of potent Th1 responses against Listeria38 and promotes antigen-specific clonal expansion and survival of effector CD4+ T cells in response to flagellin39. In our model, elevated serum IL-18 coincides with increased CD4+ T-cell effector responses and persists for two to three weeks during the microbial clearance phase when most effector cells die and only a small fraction survive to become quiescent, long-lived memory cells40. We saw that when NLRC4 activation is defective, IL-18 deficiency partially prevents the amplification of memory responses. Other mechanisms may be important, however, such as the production of IL-1β that can enhance Th1 cell expansion41 or cytokine-independent mechanisms, such as phagosome acidification, which can affect the development of adaptive immunity against Gram-positive bacteria42.
In conclusion, here we describe an unknown mechanism by which reduced activation of a PRR, NLRC4, enhances antigen-specific memory responses and promotes long-term protection against rechallenge. This reveals a clear link between NLRC4 activation and restriction of pathogen-specific immunity within a physiological setting. Whether this phenomenon is induced by the pathogen as a strategy to impede the development of optimal memory responses or by the host to avoid immunopathology by tightly regulating levels of proinflammatory cytokines remains to be determined.
Methods
Mice
Wild-type C57BL/6J mice were obtained from Charles River. Nlrc4−/− and Nlrp3−/− mice on a C57BL/6J background (after at least eight back crosses onto the C57BL/6J background) were kindly provided by K. Fitzgerald (University of Massachusetts Medical School). The Nlrc4−/−Nlrp3−/− mice were generated in-house by crossing Nlrc4−/− with Nlrp3−/− mice. All mouse colonies were bred independently. Mice were genotyped by PCR using standard protocols. All mice were maintained in a specific pathogen-free facility according to the Animals Scientific Procedures outlined by the UK Home Office regulations. All work involving live animals complied with the University of Cambridge Ethics Committee regulations and was performed under the Home Office Project Licence numbers 80/2572 and P48B8DA35. Both male and female mice between 8 and 24 weeks of age were used in infection trials and as donors for primary bone marrow-derived macrophages (BMDMs).
Bacterial strains and animal infections
S. Typhimurium strains SL1344 and M525P43, strains of high and intermediate virulence, respectively, were used in this study. For in vitro studies, S. Typhimurium was grown from single colonies to exponential phase, by inoculating a 17.5 h overnight culture 1 in 10 into LB medium and incubating at 37 °C, 200 r.p.m. for 2 h. For in vivo infections, stationary phase S. Typhimurium cultures were washed and resuspended in Dulbecco’s PBS (D-PBS, Sigma). For i.v. challenge experiments, 0.2 ml of the inoculum were administered systemically via the lateral tail vein. For oral challenge experiments, mice were lightly anaesthetized with isoflurane and then inoculated with 0.2 ml of the inoculum via oral gavage. Control mice were inoculated with sterile PBS only. The exact dose of Salmonella administered was determined by serial dilution and plating the inoculum on LB plates before and after infection. Primary challenge in all experiments was performed by i.v. administration of approximately 104 c.f.u. S. Typhimurium 525P wild-type or mutant per mouse unless mentioned otherwise. Exact doses for each experiment are given in the legends of the respective graphs. All experiments were performed after approval from the University of Cambridge ethical review committee and under the UK Government Home Office regulations licence numbers 80/2135, 80/11763, PF86EABB1 and P48B8DA35. In this work, no statistical methods were used to predetermine the sample size. The experiments were not randomized, and the investigators were not blinded to allocation during the experiments and outcome assessments.
Humane endpoint curves
In animal trials that involved rechallenge with fully virulent S. Typhimurium, we constructed humane endpoint curves as an alternative to survival curves. To do this, mice were weighed once and assessed twice daily for manifestation of clinical signs associated with generalized infection. These include weight loss of over 15% of maximum body weight, marked piloerection, subdued behavioural pattern even when provoked and reduced exploration, isolation from peers, intermittent hunched posture and persistent oculo-nasal discharge. Mice would be euthanized without delay when any of these signs persisted for more than 12 h.
Bacterial isolation and enumeration
Mice were euthanized at specific intervals following infection and their spleens and livers removed aseptically. Organs were homogenized in 10 ml sterile water using a Colworth stomacher. Organ homogenates were serially diluted and plated on LB agar plates. LB plates were incubated overnight at 37 °C followed by enumeration of c.f.u. In some experiments, spleens were first passed through a cell strainer to produce single cell suspensions in RPMI medium, a fraction of which was then used for bacterial enumeration.
Construction of S. Typhimurium M525PΔfliC::fliCE.coli and bacterial motility assays
To engineer S. Typhimurium that expresses the FliC protein of the non-pathogenic E. coli K-12 strain MG1655, a DNA fragment was synthesized by GeneArt Gene Synthesis (Life Technologies) encoding E. coli K-12 MG1655 fliC adjacent to a chloramphenicol resistance cassette (CmR) and flanked by 60 bp arms homologous to the sequence immediately upstream and downstream of S. Typhimurium M525P fliC. This fliCMG1655 cassette was transformed into electrocompetent S. Typhimurium M525P cells carrying the pBADλRed plasmid44,45. Transformants were selected on LB agar supplemented with 25 μg ml−1 of chloramphenicol and positive clones were confirmed by sequencing. The growth rate of the mutant strain was assessed by performing viable bacterial counts and found to be similar to that of the wild-type strain (Supplementary Fig. 1).
To assess bacterial motility, overnight cultures grown in LB medium were diluted into fresh LB medium to an optical density (OD600) of 0.05 and grown at 37 °C until cultures reached an OD600 of 1.0. Two microlitres of culture were inoculated into 0.25% tryptone agar (2.5 g l−1 of agar, 10 g l−1 of tryptone, 5 g l−1 of NaCl). Plates were incubated at 37 °C for between 4 and 6 h and swarm diameter was measured.
Cell culture and infections
BMDMs were prepared as previously described15. They were infected with S. Typhimurium grown to exponential phase as previously described46. Cell viability was assessed using the Cytotox 96 Non-Radioactive Cytotoxicity assay (Promega).
CD4+ T-cell ex vivo stimulation assays
Mice were euthanized, their spleens aseptically removed and, after mechanical disruption, passed through a 70-μm cell strainer (BD Biosciences) to obtain single cell suspensions in prewarmed RPMI containing 2% (vol/vol) FCS (Hyclone). Cell suspensions were washed once and resuspended in Red Blood Cell Lysis buffer (Sigma), mixed and incubated for 10 min on ice. Total splenocytes were washed twice and passed through a 30-μm filter (Partec). CD4+ T cells were purified using CD4 microbeads and magnetic positive selection according to the manufacturer’s guidelines (Miltenyi Biotec). This routinely resulted in cell populations containing over 95% CD4+ cells. CD4+ T cells were resuspended in complete RPMI medium containing 10% FCS (Hyclone), 5 mM l-glutamine, 100 U ml−1 of penicillin, 100 μg ml−1 of streptomycin, 1 mM HEPES and 0.02 mM β-mercaptoethanol. Cells were seeded at 2 × 105 cells per well in round bottom 96-well plates (Greiner Bio-One) together with wild-type naïve total splenocytes previously treated (1 h at 37 °C and 5% CO2) with Mitomycin C 25 μg ml−1 (Sigma) at a 1:1 ratio (total cell number of 4 × 105 cells per well). Cocultures from each mouse were left unstimulated, stimulated with 2 μg ml−1 antimouse CD3e (clone 145-2C11, eBioscience) or 20 μg ml−1 whole Salmonella cell extract, prepared as described previously47 and incubated at 37 °C and 5% CO2. In some experiments, the polyclonal stimulus was provided by concanavalin A (rather than antimouse CD3e), which was used at a final concentration of 5 μg ml−1. Cell culture supernatants were collected after 24 and 72 h incubation for cytokine quantification.
When CD4+ T Cells were stimulated ex vivo to assess memory immunity, it was important to confirm that the primary infection has cleared and the effector immune response subsided. For this reason, any mice whose microbial load was above detection limit (20 c.f.u. for the spleen and 10 c.f.u. for the liver) were not included in the study.
ELISpot and cytokine secretion assay
The numbers of CD4+ T cells secreting IFN-γ were assessed by ELISpot and cytokine secretion assays. For ELISpot, capture and detection antibodies supplied by the mouse IFN-γ ELISpot Development Module (R&D Systems) were used according to the manufacturer’s guidelines. Briefly, cocultures of purified CD4+ T cells with syngeneic feeder cells were incubated in duplicate wells in the presence or absence of whole Salmonella cell extract for 12 h. Cell suspensions were then transferred in twofold serial dilutions to a capture antibody-coated 96-well filtration plate (Millipore) and incubated for a further 12 h. ELISpot Blue Colour Module (R&D Systems) analysis of the cell supernatant was performed according to the manufacturer’s guidelines. Data were expressed as IFN-γ spot-forming cells per 106 cells. The number of CD4+ T cells secreting IFN-γ was also determined using a flow cytometry-based cytokine secretion assay (Miltenyi Biotec) according to the manufacturer’s guidelines. To reduce non-specific background staining and increase sensitivity, cells were simultaneously stained with a fixable viability dye and antimouse CD4 antibody (clone GK1.5) (both from Invitrogen) while the sequential gating strategy used is shown in Supplementary Fig. 2. Data from 100,000 viable cells were acquired from each mouse using a DxP Multi-color updated FACSCan (BD) analysed with FlowJo Software (TreeStar). In subsequent experiments, cells were also stained with antimouse CD44 (clone IM7) and CD62L (clone MEL-14) during cytokine secretion assay to better characterize the class of memory CD4+ T cells secreting IFN-γ (effector or central memory). The sequential gating strategy for these analyses is shown in Extended Data Fig. 3. In this case, data from 100,000 viable cells were acquired from each mouse using an Attune NxT Flow Cytometer (Invitrogen) and analysed with FlowJo Software (TreeStar).
Enzyme-linked immunosorbent assay (ELISA) and multiplex immunoassays for cytokine quantification
Cytokine levels were measured in the serum and cell culture supernatant by bead-based, multiple cytokine detection immunoassay and/or ELISA analysis. Mouse serum was analysed via ELISA for levels of IL-1β (BD OptEIA set) and IL-18 (MBL International) and via multiplex immunoassays (ProcartaPlex, Invitrogen) for IFN-γ and TNF-α. Cell culture supernatant collected from the CD4+ T-cell stimulation assays was analysed by ELISA for levels of IFN-γ and IL-2 (DuoSet Development kit, R&D Systems) and a flow cytometric, bead-based immunoassay (FlowCytomix, ebioscience) for simultaneous detection of mouse GM-CSF, IL-6 and TNF-α. IL-1β production from BMDMs infected with Salmonella was also analysed by ELISA. ELISAs and multiplex immunoassays were performed according to the manufacturer’s guidelines.
Detection of serum anti-LPS antibodies
Antibody (Ab) titres against S. Typhimurium LPS were measured in the serum of infected and control mice by ELISA as previously described48. Serum Ab titres were calculated on the basis a standard curve constructed by twofold serial dilutions of the same positive serum sample run on every plate. The dilution at which OD450–OD570 of the standard sample approximated the value of 1 was arbitrarily appointed as 1,000 U.
Statistical analyses
All statistical analyses were performed with GraphPad Prism v.8 with each specific analysis described in the appropriate figure legend. Exact P values are shown on each graph separately with P < 0.05 considered statistically significant.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
All data used to generate the figures presented in this work are available in Cambridge Research Repository Apollo with the identifier https://doi.org/10.17863/CAM.41575. Source data are provided with this paper.
References
Takeda, K. & Akira, S. Toll-like receptors. Curr. Protoc. Immunol. 109, 14.12.1–14.12.10 (2015).
Gross, O., Thomas, C. J., Guarda, G. & Tschopp, J. The inflammasome: an integrated view. Immunol. Rev. 243, 136–151 (2011).
Iwasaki, A. & Medzhitov, R. Control of adaptive immunity by the innate immune system. Nat. Immunol. 16, 343–353 (2015).
Evavold, C. L. & Kagan, J. C. How inflammasomes inform adaptive immunity. J. Mol. Biol. 430, 217–237 (2018).
McSorley, S. J., Cookson, B. T. & Jenkins, M. K. Characterization of CD4+ T cell responses during natural infection with Salmonella typhimurium. J. Immunol. 164, 986–993 (2000).
Miao, E. A. et al. Cytoplasmic flagellin activates caspase-1 and secretion of interleukin 1 beta via Ipaf. Nat. Immunol. 7, 569–575 (2006).
Franchi, L. et al. Cytosolic flagellin requires Ipaf for activation of caspase-1 and interleukin 1 beta in salmonella-infected macrophages. Nat. Immunol. 7, 576–582 (2006).
Miao, E. A. et al. Innate immune detection of the type III secretion apparatus through the NLRC4 inflammasome. Proc. Natl Acad. Sci. USA 107, 3076–3080 (2010).
Lightfield, K. L. et al. Critical function for Naip5 in inflammasome activation by a conserved carboxy-terminal domain of flagellin. Nat. Immunol. 9, 1171–1178 (2008).
Zhao, Y. et al. The NLRC4 inflammasome receptors for bacterial flagellin and type III secretion apparatus. Nature 477, 596–600 (2011).
Kofoed, E. M. & Vance, R. E. Innate immune recognition of bacterial ligands by NAIPs determines inflammasome specificity. Nature 477, 592–595 (2011).
Martinon, F., Burns, K. & Tschopp, J. The inflammasome: a molecular platform triggering activation of inflammatory caspases and processing of proIL-beta. Mol. Cell 10, 417–426 (2002).
Broz, P. & Dixit, V. M. Inflammasomes: mechanism of assembly, regulation and signalling. Nat. Rev. Immunol. 16, 407–420 (2016).
Ding, J. & Shao, F. Growing a gasdermin pore in membranes of pyroptotic cells. EMBO J. 37, e100067 (2018).
Man, S. M. et al. Inflammasome activation causes dual recruitment of NLRC4 and NLRP3 to the same macromolecular complex. Proc. Natl Acad. Sci. USA 111, 7403–7408 (2014).
Qu, Y. et al. NLRP3 recruitment by NLRC4 during Salmonella infection. J. Exp. Med. 213, 877–885 (2016).
Garlanda, C., Dinarello, C. A. & Mantovani, A. The interleukin-1 family: back to the future. Immunity 39, 1003–1018 (2013).
O’donnell, H. et al. Toll-like receptor and inflammasome signals converge to amplify the innate bactericidal capacity of T helper 1 cells. Immunity 40, 213–224 (2014).
Kupz, A. et al. NLRC4 inflammasomes in dendritic cells regulate noncognate effector function by memory CD8(+) T cells. Nat. Immunol. 13, 162–169 (2012).
Hess, J., Ladel, C., Miko, D. & Kaufmann, S. H. E. Salmonella typhimurium aroA– infection in gene-targeted immunodeficient mice - major role of CD4(+) TCR-alpha beta cells and IFN-gamma in bacterial clearance independent of intracellular location. J. Immunol. 156, 3321–3326 (1996).
Nauciel, C. Role of Cd4+ T-cells and T-independent mechanisms in acquired-resistance to Salmonella-Typhimurium infection. J. Immunol. 145, 1265–1269 (1990).
Broz, P. et al. Redundant roles for inflammasome receptors NLRP3 and NLRC4 in host defense against Salmonella. J. Exp. Med. 207, 1745–1755 (2010).
Lara-Tejero, M. et al. Role of the caspase-1 inflammasome in Salmonella typhimurium pathogenesis. J. Exp. Med. 203, 1407–1412 (2006).
Mastroeni, P., Simmons, C., Fowler, R., Hormaeche, C. E. & Dougan, G. Igh-6-/- (B-cell-deficient) mice fail to mount solid acquired resistance to oral challenge with virulent Salmonella enterica serovar Typhimurium and show impaired Th1 T-cell responses to Salmonella antigens. Infect. Immun. 68, 46–53 (2000).
Heng, T. S. P., Painter, M. W. & Project, I. G. The immunological genome project: networks of gene expression in immune cells. Nat. Immunol. 9, 1091–1094 (2008).
Bruchard, M. et al. The receptor NLRP3 is a transcriptional regulator of TH2 differentiation. Nat. Immunol. 16, 859–870 (2015).
Mariathasan, S. et al. Differential activation of the inflammasome by caspase-1 adaptors ASC and Ipaf. Nature 430, 213–218 (2004).
Tsutsui, H., Matsui, K., Okamura, H. & Nakanishi, K. Pathophysiological roles of interleukin-18 in inflammatory liver diseases. Immunol. Rev. 174, 192–209 (2000).
Mastroeni, P., Villarreal-Ramos, B. & Hormaeche, C. E. Role of T cells, TNF alpha and IFN gamma in recall of immunity to oral challenge with virulent salmonellae in mice vaccinated with live attenuated Aro-Salmonella. Vaccines. Micro. Pathogenesis 13, 477–491 (1992).
Kupz, A. et al. Contribution of Thy1+ NK cells to protective IFN-gamma production during Salmonella typhimurium infections. Proc. Natl Acad. Sci. USA 110, 2252–2257 (2013).
Letran, S. E. et al. TLR5-deficient mice lack basal inflammatory and metabolic defects but exhibit impaired CD4 T cell responses to a flagellated pathogen. J. Immunol. 186, 5406–5412 (2011).
Yang, J. et al. Flagellins of Salmonella Typhi and nonpathogenic Escherichia coli are differentially recognized through the NLRC4 pathway in macrophages. J. Innate Immun. 6, 47–57 (2014).
Amer, A. et al. Regulation of Legionella phagosome maturation and infection through flagellin and host Ipaf. J. Biol. Chem. 281, 35217–35223 (2006).
Franchi, L. et al. Critical role for Ipaf in Pseudomonas aeruginosa-induced caspase-1 activation. Eur. J. Immunol. 37, 3030–3039 (2007).
Suzuki, T. et al. Differential regulation of caspase-1 activation, pyroptosis, and autophagy via Ipaf and ASC in Shigella-infected macrophages. PLoS Pathog. 3, e111 (2007).
Sauer, J. D. et al. Listeria monocytogenes engineered to activate the Nlrc4 inflammasome are severely attenuated and are poor inducers of protective immunity. Proc. Natl Acad. Sci. USA 108, 12419–12424 (2011).
Yoshimoto, T. et al. IL-12 up-regulates IL-18 receptor expression on T cells, Th1 cells, and B cells: synergism with IL-18 for IFN-gamma production. J. Immunol. 161, 3400–3407 (1998).
Neighbors, M. et al. A critical role for interleukin 18 in primary and memory effector responses to Listeria monocytogenes that extends beyond its effects on Interferon gamma production. J. Exp. Med. 194, 343–354 (2001).
Maxwell, J. R. et al. IL-18 bridges innate and adaptive immunity through IFN-gamma and the CD134 pathway. J. Immunol. 177, 234–245 (2006).
Pepper, M. & Jenkins, M. K. Origins of CD4+ effector and central memory T cells. Nat. Immunol. 12, 467–471 (2011).
Ben-Sasson, S. Z. et al. IL-1 acts directly on CD4 T cells to enhance their antigen-driven expansion and differentiation. Proc. Natl Acad. Sci. USA 106, 7119–7124 (2009).
Sokolovska, A. et al. Activation of caspase-1 by the NLRP3 inflammasome regulates the NADPH oxidase NOX2 to control phagosome function. Nat. Immunol. 14, 543–553 (2013).
Mastroeni, P. et al. Antimicrobial actions of the NADPH phagocyte oxidase and inducible nitric oxide synthase in experimental salmonellosis. II Effects on microbial proliferation and host survival in vivo. J. Exp. Med. 192, 237–247 (2000).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
Mo, E., Peters, S. E., Willers, C., Maskell, D. J. & Charles, I. G. Single, double and triple mutants of Salmonella enterica serovar Typhimurium degP (htrA), degQ (hhoA) and degS (hhoB) have diverse phenotypes on exposure to elevated temperature and their growth in vivo is attenuated to different extents. Micro. Pathog. 41, 174–182 (2006).
Man, S. M. et al. Salmonella infection induces recruitment of Caspase-8 to the inflammasome to modulate IL-1 beta production. J. Immunol. 191, 5239–5246 (2013).
Harrison, J. A., Villarreal-Ramos, B., Mastroeni, P., DeHormaeche, R. D. & Hormaeche, C. E. Correlates of protection induced by live Aro(-) Salmonella Typhimurium vaccines in the murine typhoid model. Immunology 90, 618–625 (1997).
Dehormaeche, R. D., Jessop, H. & Bundell, C. Antibodies to the C epitope of Neisseria gonorrhoeae are present in patients with gonorrhea and absent in normal sera. J. Gen. Microbiol. 134, 1289–1297 (1988).
Acknowledgements
This work was supported by grants from Biotechnology and Biological Sciences Research Council (BBSRC) (nos. BB/H003916/1 and BB/K006436/1), Zoetis UK (no. BB/K006436/1) and Wellcome Trust (no. 108045/Z/15/Z) to C.E.B. A.S.B was supported by a Wellcome Trust 4-year PhD studentship. We thank K. A. Fitzgerald and D. Golenbock for suppling the gene-deficient mouse strains, S. M. Man for critical review of the manuscript and helpful discussions, and A. Cooke and members of her group, especially P. Zaccone and S. Newland, for sharing their expertise with P.T. and helpful discussions.
Author information
Authors and Affiliations
Contributions
P.T. designed, performed and analysed all in vivo, ex vivo and in vitro experiments. P.M., J.A.W., L.J.H., A.S.B. and S.J.W performed part of some in vivo experiments. O.J.B. performed the bacterial motility assays. J.A.W supervised all bacterial mutagenesis work. A.S.B. performed all bacterial mutagenesis work. P.M., D.J.M. and C.E.B. conceived the study and secured the funding. C.E.B. and P.T. supervised the study. P.T., J.A.W. and C.E.B. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 NLRC4 is required to restrict microbial spread during an oral sub-lethal but not an oral lethal infection with S. Typhimurium.
a, Wild-type and nlrc4−/− mice were challenged orally with 4.4 × 109 CFU S. Typhimurium M525P to establish a sub-lethal infection and microbial burden was determined in the spleen and liver at 5 days (n = 6 for wild-type and Nlrc4−/−) and 14 days (n = 7 for wild-type and n = 6 for Nlrc4−/−) post-challenge. b, Wild type and Nlrc4−/− mice were challenged orally 8 × 108 CFU S. Typhimurium SL1344 to establish a lethal infection and microbial burden was determined in the spleen and liver at 4 days post-challenge. Each symbol represents one mouse and horizontal lines delineate the mean. Statistical significance was calculated by 2-way ANOVA followed by Sidak’s multiple comparisons test for a and two-tailed Mann Whitney test for b. Data are representative of two independent experiments.
Extended Data Fig. 2 The effect of infection dose on the potency of ex vivo CD4+ T cell-mediated responses.
Wild-type mice were challenged with either 1.58 × 105 CFU, 1.72 × 104 CFU or 1.53 × 103 CFU S. Typhimurium M525P. Mice were euthanised 104-125 days after challenge and their splenic CD4+ T cells were stimulated with medium only (w/o stim.), anti-mouse CD3e (anti-CD3e) or whole bacterial cell extract (extract). a, IFN-γ measured by ELISA in the cell culture supernatant after 72 hours. b, IL-2 measured by ELISA in the cell culture supernatant after 24 hours. Data are shown as mean ± s.e.m. Statistical significance was calculated by a one-way ANOVA. Data were generated from one experiment.
Extended Data Fig. 3 Flow cytometric analysis reveals that CD4+ T cells secreting IFN-γ ex vivo have the profile of effector memory, rather than central memory, CD4+ T cells.
CD4+ T cells isolated from the spleen of wild-type and Nlrc4−/− mice 100 days after primary challenge with 1.24 × 104 CFU S. Typhimurium M525P were stimulated ex vivo with whole bacterial cell extract and phenotypically characterised via flow cytometry. Viable cells were first selected using a fixable viability dye (FVD) and single cells only were further included in the analysis based on their forward scatter area vs. forward scatter height. Single viable cells expressing CD4 were then separated to IFN-γ- and IFN-γ+ with both groups analysed for expression of CD44 and CD62L. For CD4+IFN-γ- cells, the phenotype CD44+CD62−represents effector T cells, while the phenotype CD44−CD62+ represents naïve T cells. For CD4+IFN-γ+ cells, the phenotype CD44+CD62−represents effector memory T cells, while the phenotype CD44+CD62+ represents central memory T cells. Percentages shown in the respective quadrants are mean values ± SD from N = 5 wild-type and N = 6 Nlrc4−/− mice. Data were generated from one experiment.
Extended Data Fig. 4 NLRC4 does not affect protection against lethal re-challenge.
Wild-type and Nlrc4−/− mice were challenged with either sterile PBS (naïve) or 1-1.15 × 104 CFU S. Typhimurium M525P and allowed to clear the primary infection for 90-106 days. They were then re-challenged orally with 1.37-7.2 × 107 CFU S. Typhimurium SL1344 to establish a lethal infection and inspected twice daily. They were euthanized upon detection of adverse signs and a humane end point curve was constructed for a, naïve mice and b, mice ‘immunised’ with Salmonella. Data has been pooled from 2 independent experiments. Statistical significance was calculated by a two-sided log-rank test.
Extended Data Fig. 5 Data variation between independent trials comparing CD4+ T cell-mediated responses in wild-type, Nlrc4-/-, Nlrp3-/- and Nlrp3-/-Nlrc4-/- mice is not statistically significant.
Levels of IFN-γ produced by CD4+ T cells stimulated ex vivo with whole bacterial cell extract were compared between wild-type mice used in three independent trials shown in Fig. 1c, Fig. 3a and Fig. 3b. Each symbol represents one mouse and horizontal lines delineate the mean. The data was tested for statistical significance by one-way ANOVA.
Extended Data Fig. 6 In the absence of NLRC4, Salmonella-infected macrophages do not undergo early cell death enabling them to survive longer and secrete IL−1β via activation of NLRP3.
Unprimed bone marrow derived macrophages (BMDMs) from wild-type, Nlrc4−/−, Nlrp3−/− and Nlrc4−/−Nlrp3−/− mice were infected with S. Typhimurium M525P in logarithmic growth at the MOI=50. a, Cytotoxicity was determined by measuring levels of LDH released in the cell culture supernatant. b, Levels of IL-1β were determined by ELISA in the cell culture supernatant. Data are shown as mean of triplicate wells ± s.e.m. Data are representative of two independent experiments.
Extended Data Fig. 7 Pairwise Sequence Alignment of FliC from E. coli K-12 MG1655 and S. Typhimurium M525P.
Comparison of the predicted FliC proteins from S. Typhimurium M525P and E. coli K-12 MG1655 by Clustal Omega (EMBL-EBI).
Extended Data Fig. 8 Salmonella expressing flagellin from non-pathogenic E. coli (M525P fliCMG1655) is motile but impaired at activating the NLRC4 inflammasome in macrophages.
a, Comparison of the motility diameter between S. Typhimurium M525P, M525P fliCMG1655 and SL1344. Data are expressed as % of the diameter achieved by the SL1344. b–d, Unprimed bone marrow derived macrophages from wild-type and Nlrc4−/− mice were infected for 1 h, 2 h, 6 h and 24 h with S. Typhimurium M525P or M525P fliCMG1655 in logarithmic growth. b, Cytotoxicity was determined by measuring levels of LDH released in the cell culture supernatant at an MOI of 50. c, Levels of IL-1β and d, TNF-α were determined by ELISA in the cell culture supernatant at an MOI of 1. Data has been pooled from 6 independent experiments and shown as mean ± s.e.m. in a. Data are representative of two independent experiments in b–d and shown as mean of triplicate wells ± s.e.m.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Rights and permissions
About this article
Cite this article
Tourlomousis, P., Wright, J.A., Bittante, A.S. et al. Modifying bacterial flagellin to evade Nod-like Receptor CARD 4 recognition enhances protective immunity against Salmonella. Nat Microbiol 5, 1588–1597 (2020). https://doi.org/10.1038/s41564-020-00801-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-020-00801-y
This article is cited by
-
Immunology of gut microbiome and liver in non-alcoholic fatty liver disease (NAFLD): mechanisms, bacteria, and novel therapeutic targets
Archives of Microbiology (2024)
-
Inflammasome activation, NLRP3 engagement and macrophage recruitment to tumor microenvironment are all required for Salmonella antitumor effect
Cancer Immunology, Immunotherapy (2022)
-
Insights into inflammasome regulation: cellular, molecular, and pathogenic control of inflammasome activation
Immunologic Research (2022)