Abstract
The microbiota has been shown to promote intestinal tumourigenesis, but a possible anti-tumourigenic effect has also been postulated. Here, we demonstrate that changes in the microbiota and mucus composition are concomitant with tumourigenesis. We identified two anti-tumourigenic strains of the microbiota—Faecalibaculum rodentium and its human homologue, Holdemanella biformis—that are strongly under-represented during tumourigenesis. Reconstitution of ApcMin/+ or azoxymethane- and dextran sulfate sodium-treated mice with an isolate of F. rodentium (F. PB1) or its metabolic products reduced tumour growth. Both F. PB1 and H. biformis produced short-chain fatty acids that contributed to control protein acetylation and tumour cell proliferation by inhibiting calcineurin and NFATc3 activation in mouse and human settings. We have thus identified endogenous anti-tumourigenic bacterial strains with strong diagnostic, therapeutic and translational potential.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Source Data for the figures and Extended Data figures are provided in the online version of the paper. Raw sequencing data and metadata associated to samples are available online at the NCBI under the accession number PRJNA564752.
References
Sears, C. L. & Garrett, W. S. Microbes, microbiota, and colon cancer. Cell Host Microbe 15, 317–328 (2014).
Kostic, A. D. et al. Genomic analysis identifies association of Fusobacterium with colorectal carcinoma. Genome Res. 22, 292–298 (2012).
Mima, K. et al. Fusobacterium nucleatum in colorectal carcinoma tissue according to tumor location. Clin. Transl. Gastroenterol. 7, e200 (2016).
Yu, T. et al. Fusobacterium nucleatum promotes chemoresistance to colorectal cancer by modulating autophagy. Cell 170, 548–563 (2017).
Kinzler, K. W. & Vogelstein, B. Lessons from hereditary colorectal cancer. Cell 87, 159–170 (1996).
Li, Y. et al. Gut microbiota accelerate tumor growth via c-jun and STAT3 phosphorylation in APCMin/+ mice. Carcinogenesis 33, 1231–1238 (2012).
Kostic, A. D. et al. Fusobacterium nucleatum potentiates intestinal tumorigenesis and modulates the tumor-immune microenvironment. Cell Host Microbe 14, 207–215 (2013).
Wu, S. et al. A human colonic commensal promotes colon tumorigenesis via activation of T helper type 17 T cell responses. Nat. Med. 15, 1016–1022 (2009).
Arthur, J. C. et al. Intestinal inflammation targets cancer-inducing activity of the microbiota. Science 338, 120–123 (2012).
Zhan, Y. et al. Gut microbiota protects against gastrointestinal tumorigenesis caused by epithelial injury. Cancer Res. 73, 7199–7210 (2013).
Wong, S. H. et al. Gavage of fecal samples from patients with colorectal cancer promotes intestinal carcinogenesis in germ-free and conventional mice. Gastroenterology 153, 1621–1633 (2017).
Donohoe, D. R. et al. A gnotobiotic mouse model demonstrates that dietary fiber protects against colorectal tumorigenesis in a microbiota- and butyrate-dependent manner. Cancer Discov. 4, 1387–1397 (2014).
Chang, D. H. et al. Faecalibaculum rodentium gen. nov., sp. nov., isolated from the faeces of a laboratory mouse. Antonie Van Leeuwenhoek 108, 1309–1318 (2015).
Johansson, M. E. et al. The inner of the two Muc2 mucin-dependent mucus layers in colon is devoid of bacteria. Proc. Natl Acad. Sci. USA 105, 15064–15069 (2008).
Xiao, X. et al. Role of MUC20 overexpression as a predictor of recurrence and poor outcome in colorectal cancer. J. Transl. Med. 11, 151 (2013).
Betge, J. et al. MUC1, MUC2, MUC5AC, and MUC6 in colorectal cancer: expression profiles and clinical significance. Virchows Arch. 469, 255–265 (2016).
Guglietta, S. et al. Coagulation induced by C3aR-dependent NETosis drives protumorigenic neutrophils during small intestinal tumorigenesis. Nat. Commun. 7, 11037 (2016).
Sakamoto, M., Tanaka, Y., Benno, Y. & Ohkuma, M. Butyricimonas faecihominis sp. nov. and Butyricimonas paravirosa sp. nov., isolated from human faeces, and emended description of the genus Butyricimonas. Int. J. Syst. Evol. Microbiol. 64, 2992–2997 (2014).
Singh, N. et al. Activation of gpr109a, receptor for niacin and the commensal metabolite butyrate, suppresses colonic inflammation and carcinogenesis. Immunity 40, 128–139 (2014).
Boffa, L. C., Vidali, G., Mann, R. S. & Allfrey, V. G. Suppression of histone deacetylation in vivo and in vitro by sodium butyrate. J. Biol. Chem. 253, 3364–3366 (1978).
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Donohoe, D. R. et al. The Warburg effect dictates the mechanism of butyrate-mediated histone acetylation and cell proliferation. Mol. Cell 48, 612–626 (2012).
Peuker, K. et al. Epithelial calcineurin controls microbiota-dependent intestinal tumor development. Nat. Med. 22, 506–515 (2016).
Imai, Y. et al. Histone deacetylase inhibitor panobinostat induces calcineurin degradation in multiple myeloma. JCI Insight 1, e85061 (2016).
Han, K. J., Lee, N. K., Park, H. & Paik, H. D. Anticancer and anti-inflammatory activity of probiotic Lactococcus lactis NK34. J. Microbiol. Biotechnol. 25, 1697–1701 (2015).
Zeller, G. et al. Potential of fecal microbiota for early-stage detection of colorectal cancer. Mol. Syst. Biol. 10, 766 (2014).
Tsilingiri, K., Sonzogni, A., Caprioli, F. & Rescigno, M. A novel method for the culture and polarized stimulation of human intestinal mucosa explants. J. Vis. Exp. 75, e4368 (2013).
Kesari, M. V. et al. Immunohistochemical study of MUC1, MUC2 and MUC5AC in colorectal carcinoma and review of literature. Indian J. Gastroenterol. 34, 63–67 (2015).
Yan, X., Liu, L., Li, H., Qin, H. & Sun, Z. Clinical significance of Fusobacterium nucleatum, epithelial–mesenchymal transition, and cancer stem cell markers in stage III/IV colorectal cancer patients. Onco Targets Ther. 10, 5031–5046 (2017).
Bullman, S. et al. Analysis of Fusobacterium persistence and antibiotic response in colorectal cancer. Science 358, 1443–1448 (2017).
Mendes, R. T. et al. Endothelial cell response to Fusobacterium nucleatum. Infect. Immun. 84, 2141–2148 (2016).
Gur, C. et al. Binding of the Fap2 protein of Fusobacterium nucleatum to human inhibitory receptor TIGIT protects tumors from immune cell attack. Immunity 42, 344–355 (2015).
Chen, Y. et al. Invasive Fusobacterium nucleatum activates beta-catenin signaling in colorectal cancer via a TLR4/P-PAK1 cascade. Oncotarget 8, 31802–31814 (2017).
Chung, L. et al. Bacteroides fragilis toxin coordinates a pro-carcinogenic inflammatory cascade via targeting of colonic epithelial cells. Cell Host Microbe 23, 203–214 (2018).
Dejea, C. M. et al. Patients with familial adenomatous polyposis harbor colonic biofilms containing tumorigenic bacteria. Science 359, 592–597 (2018).
Yan, F. et al. Interleukin-13-induced MUC5AC expression is regulated by a PI3K–NFAT3 pathway in mouse tracheal epithelial cells. Biochem. Biophys. Res. Commun. 446, 49–53 (2014).
San-Millan, I. & Brooks, G. A. Reexamining cancer metabolism: lactate production for carcinogenesis could be the purpose and explanation of the Warburg effect. Carcinogenesis 38, 119–133 (2017).
Faubert, B. et al. Lactate metabolism in human lung tumors. Cell 171, 358–371 (2017).
Duncan, S. H., Louis, P. & Flint, H. J. Lactate-utilizing bacteria, isolated from human feces, that produce butyrate as a major fermentation product. Appl. Environ. Microbiol. 70, 5810–5817 (2004).
Zhernakova, A. et al. Population-based metagenomics analysis reveals markers for gut microbiome composition and diversity. Science 352, 565–569 (2016).
De Maesschalck, C. et al. Faecalicoccus acidiformans gen. nov., sp. nov., isolated from the chicken caecum, and reclassification of Streptococcus pleomorphus (Barnes et al. 1977), Eubacterium biforme (Eggerth 1935) and Eubacterium cylindroides (Cato et al. 1974) as Faecalicoccus pleomorphus comb. nov., Holdemanella biformis gen. nov., comb. nov. and Faecalitalea cylindroides gen. nov., comb. nov., respectively, within the family Erysipelotrichaceae. Int. J. Syst. Evol. Microbiol. 64, 3877–3884 (2014).
Atarashi, K. et al. Treg induction by a rationally selected mixture of clostridia strains from the human microbiota. Nature 500, 232–236 (2013).
Moser, A. R., Pitot, H. C. & Dove, W. F. A dominant mutation that predisposes to multiple intestinal neoplasia in the mouse. Science 247, 322–324 (1990).
Furet, J. P. et al. Comparative assessment of human and farm animal faecal microbiota using real-time quantitative PCR. FEMS Microbiol. Ecol. 68, 351–362 (2009).
Manzari, C. et al. Draft genome sequence of Sphingobium sp. strain ba1, resistant to kanamycin and nickel ions. FEMS Microbiol. Lett. 361, 8–9 (2014).
Fosso, B. et al. BioMaS: a modular pipeline for bioinformatic analysis of metagenomic amplicons. BMC Bioinform. 16, 203 (2015).
Zhang, J, Kobert, K, Flouri, T. & Stamatakis, A. PEAR: a fast and accurate Illumina paired-end read merger. Bioinformatics 30, 614–620 (2014).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Cole, J. R. et al. The ribosomal database project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res. 37, D141–D145 (2009).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Alonso-Alemany, D. et al. Further steps in TANGO: improved taxonomic assignment in metagenomics. Bioinformatics 30, 17–23 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60 (2011).
Salzman, N. H. et al. Enteric salmonella infection inhibits Paneth cell antimicrobial peptide expression. Infect. Immun. 71, 1109–1115 (2003).
Oksanen, J. et al. Vegan: community ecology package. R package vegan version 2.2-1 (2015); https://cran.r-project.org/web/packages/vegan/index.html
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2009).
Cooper, H. S., Murthy, S. N., Shah, R. S. & Sedergran, D. J. Clinicopathologic study of dextran sulfate sodium experimental murine colitis. Lab. Invest. 69, 238–249 (1993).
Boivin, G. P. et al. Pathology of mouse models of intestinal cancer: consensus report and recommendations. Gastroenterology 124, 762–777 (2003).
Truong, D. T. et al. MetaPhlAn2 for enhanced metagenomic taxonomic profiling. Nat. Methods 12, 902–903 (2015).
Pasolli, E. et al. Accessible, curated metagenomic data through ExperimentHub. Nat. Methods 14, 1023–1024 (2017).
Segata, N., Bornigen, D., Morgan, X. C. & Huttenhower, C. PhyloPhlAn is a new method for improved phylogenetic and taxonomic placement of microbes. Nat. Commun. 4, 2304 (2013).
Gargari, G. et al. Consumption of a Bifidobacterium bifidum strain for 4 weeks modulates dominant intestinal bacterial taxa and fecal butyrate in healthy adults. Appl. Environ. Microbiol. 82, 5850–5859 (2016).
Tuominen, V. J., Ruotoistenmaki, S., Viitanen, A., Jumppanen, M. & Isola, J. ImmunoRatio: a publicly available web application for quantitative image analysis of estrogen receptor (ER), progesterone receptor (PR), and Ki-67. Breast Cancer Res. 12, R56 (2010).
Acknowledgements
We thank M. Bugatti and S. Licini (supported by Fondazione Beretta, Brescia, Italy) for their technical support in the histological analyses; A. Thomas and E. Pasolli for performing the bioinformatics analyses; C. Faccani for technical support for the in vitro experiments; and E. Mileti, C. Burrello and M. R. Giuffré for their technical support for the in vivo experiments. This work has been supported by grants from the Italian Association for Cancer Research (grant no. AIRC IG 17628) and the European Research council (grant no. 615735, HOMEOGUT ERC) to M.R. E.Z., I.S. and A.B. were/are recipients of a FIRC fellowship. T.S. is a recipient of a fellowship from Fondazione Veronesi. G.F. is a recipient of a grant from the Italian Ministry of Health (grant no. GR-2013-02359806).
Author information
Authors and Affiliations
Contributions
E.Z. and C.P. ideated, performed and analysed all of the experiments. T.S. and F.S. helped with the execution of experiments. A.B., I.S. and S.Guglietta helped with the execution of the mouse experiments. B.F., M.M. and G.Pesole performed the 16S rRNA metagenomic analysis. L.M. and W.V. designed and carried out the histological analyses. G.N. performed the ex vivo stimulation of human colonic mucosa experiments. A.B. performed the confocal analyses. J.T. executed the metabolomic analyses. B.O. assisted in the execution of the in vitro experiments. K.A. and K.H. isolated F. PB1 and carried out the germ-free experiments. S.A. and S.Guglielmetti set up F. PB1 growth and supernatant production. S.C. set up H. biformis and L. lactis growth and supernatant production. G.F. performed the FACS analyses. F.A. and N.S. performed the phylogenetic analysis and human CRC dataset interrogation. G.Penna contributed with ideas and interpretation of results. M.R. ideated the study, coordinated the work and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Faecal microbiota diversity upon tumour progression in ApcMin/+ mice.
a, Chao1 and Simpson indexes of the faecal microbiota of WT and ApcMin/+ mice; n = 8 mice/group. Box plots show the interquartile range, median value and whiskers min to max. Data from two independent experiments. b, Tumour multiplicity in the small intestine of ApcMin/+ mice assessed at different time points; weeks 6, 28 n = 5; weeks 8, 26 n = 8; weeks 10 n = 9; weeks 12, 16, 24 n = 3; weeks 14 n = 7; weeks 18 n = 10; weeks 20 n = 33; weeks 22, 30 n = 4 mice/group. c, Normalized read counts for differentially represented species in WT and ApcMin/+ mice shown as heatmaps. The counts were log transformed and used to define the colour gradient of the heatmap; n = 8 mice/group. Data from two independent experiments. a-c, Data are represented as means ± s.e.m.
Extended Data Fig. 2 Mucin production is altered in the ileum of ApcMin/+ mice.
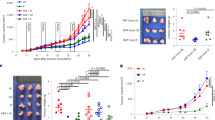
a, Representative Western blots of two independent experiments with consistent results showing the abundance of Muc1, Muc13 and Muc20 in ileal extracts from WT (healthy tissues, H) and healthy (H) and tumour (T) tissues of ApcMin/+ mice analysed at 11 weeks of age; Right: densitometric quantification (WT n = 3; ApcMin/+ n = 4 biologically independent samples). b, Representative confocal images of WT and ApcMin/+ mice ileum intestinal tissues stained with Muc1 (red) and DAPI (blue); scale bars, 50 μm. Mean fluorescence intensity of MUC1 was measured to quantify the expression of the protein in the intestinal of ApcMin/+ and WT mice (WT n = 7, ApcMin/+ n = 5 biologically independent samples). c, ApcMin/+ mice were treated with vehicle (n = 10 mice/group) or F. PB1 (n = 11 mice/group) from week 8 to 10. Number of microscopic lesions, shown as % relative to vehicle, was evaluated on H&E slides by an expert pathologist. a-c, Data from two independent experiments and represented as means ± s.e.m. P values were determined by one-way ANOVA using Tukey post-test (a), two-tailed unpaired Mann Whitney U test (b), or two-tailed unpaired Student’s t-test (c).
Extended Data Fig. 3 F. PB1 does not have a major impact on immune cells.
a-c, Flow cytometry analyses of the small intestinal and colonic lamina propria of germ-free ICR mice monocolonized with F. PB1 (GF + F. PB1 n = 10 mice/group) for the presence of T reg (a), Th17 (b) and Th1 (c) cells. Germ-free (GF, n = 9 mice/group) and SPF (n = 7 mice/group) mice were used as controls. d-g WT and ApcMin/+ mice treated with vehicle (Veh) or F. PB1 from week 8 to 12. d,e, Flow cytometric analysis of T reg, Th1 and Th17 cell populations in the small intestinal lamina propria. FoxP3+CD25+ are gated on the live CD45+ CD3+ CD4+ cells; Helios+ is gated on the FoxP3+ CD25+ cells (WT Veh, ApcMin/+ F. PB1 n = 12; WT F. PB1 n = 14; ApcMin/+ Veh n = 11 mice/group). IL17+, IFNγ+ and IL17+ IFNγ+ cells are gated on the live CD45+ CD3+ CD4+ cells (WT Veh, ApcMin/+ Veh n = 9; WT F. PB1 n = 11; ApcMin/+ F. PB1 n = 10 mice/group). Data shown as % of CD45+ cells (d) or as absolute number / whole tissue (e). f,g, Flow cytometric analysis of mononuclear phagocytes (CD11b+F4/80+ macrophages, CD11c+CD11b+ dendritic cells and Ly6ChiCD11b+ inflammatory monocytes) and neutrophils (Ly6G+CD11b+) in the small intestinal lamina propria (WT Veh n = 5; WT F. PB1, ApcMin/+ F. PB1 n = 6; ApcMin/+ Veh n = 4). Data shown as percentages relative to the CD45+ CD3- population (f) or as absolute number / whole tissue (g). h, Flow cytometric analysis of peripheral blood cells. Data shown as absolute number / ml blood (WT Veh, ApcMin/+ F. PB1 n = 13; WT F. PB1 n = 15; ApcMin/+ Veh n = 12 mice/group) P values were determined by one-way ANOVA with Bonferroni post-test (a-d), Kruskal-Wallis with Dunn post test (e), two-tailed unpaired Mann-Whitney U test (f,g,h Ly6G+CD11b+) or two-tailed unpaired Student’s t-test (h, Ly6ChiCD11b+). a-h, Data are represented as means ± s.e.m.
Extended Data Fig. 4 F. PB1 administration alters microbiota and its metabolic profile.
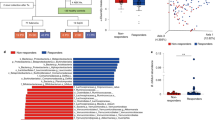
a, LEfSe comparison results between the microbiota of Vehicle and F. PB1 treated mice at 12 weeks with the highest linear discriminant analysis LDA score (log10 ≥ 2.0); Veh, n = 8; F. PB1 n = 14 mice/group. b, Faecal concentrations of succinate, isovalerate and valerate in WT and ApcMin/+ mice treated with vehicle (Veh) or F. PB1 as detected by UPLC–HR–MS; WT Veh, WT F. PB1 n = 6; ApcMin/+ Veh n = 11; ApcMin/+ F. PB1 n = 10 mice/group. Data from two independent experiments and represented as means ± s.e.m. P values were determined by two-way ANOVA with Bonferroni post-test.
Extended Data Fig. 5 SCFAs and F. PB1 spent medium (SUP) increase histone H3 acetylation and reduce NFATc3-calcineurin pathway in vitro on mouse intestinal tumour cell lines.
a, Representative WB showing H3K27 acetylation, PP2B-A and NFATc3 in cell lines treated (+) or not (-) with a mix of SCFAs. Three independent experiments were performed with consistent results. b, Densitometric quantification of WB in Fig. 4d showing NFATc3 and PP2B-A (normalized to actin) and H3K27 acetylation (normalized to vinculin). Two or three independent experiments were performed with consistent results (n = 2 or n = 3 biologically independent experiments). Data are represented as means ± s.e.m. and P values were determined by two-tailed unpaired Student’s t-test. c, Densitometric quantification of WB in Fig. 4e showing NFATc3 (normalized to vinculin) and H3K27 acetylation (normalized to H3 tot). To calculate the protein expression induced by SUP evap as a percentage, the densitometric value of SUP was assumed to be 100%. Data from two independent WB (n = 2 biologically independent experiments).
Extended Data Fig. 6 Effect of F. PB1 spent medium (SUP) in tumourigenesis.
a-c, Representative histologies from ileal and colon sections of ApcMin/+mice (a,Veh n = 13; SUP n =11 mice/group; b,Veh n = 6; SUP n =7 mice/group) and AOM and DSS treated WT mice (c,Veh n = 5; SUP n = 6 mice/group) treated with vehicle or F. PB1 SUP. Sections stained for H&E from FFPE blocks of swiss roll of small (a, 5X magnification, scale bars 500 μm; b, 3X magnification, scale bars 750 μm) and large (c, 2X magnification, scale bars 900 μm) bowel and treated as indicated by labels. Slides were scanned by Aperio Scanscope and digital images were obtained. Dysplastic lesions were selected and measured. In the lower right high power insert, a detail of dysplastic glands (200X magnification, blue scale bars 100 μm).
Extended Data Fig. 7 L. lactis does not colonize the mouse gut but its spent medium (SUP) has anti-tumourigenic effect in vivo.
a, Quantification of SCFAs in SUP of F. PB1 and L. lactis by UPLC–HR–MS. Data from 2 or 6 independent experiments (L.lactis SUP n = 2; F. PB1 SUP n = 6 biologically independent experiments). b,c, ApcMin/+ mice received vehicle (Veh, n = 7 mice/group), F. PB1 or L. lactis (n = 8 mice/group) from week 8 to 10. b, Tumour multiplicity in the small intestine normalized to vehicle-treated ApcMin/+ mice. c, Area and maximum diameter (axis lenght) of ileal dysplastic lesions normalized to the total number of lesions per mouse. Box plots show the interquartile range, median value and whiskers min to max. d, qPCR of L. lactis abundance normalized to panbacterial primers targeting the 16S rRNA gene (UNI 16S) in bacterial DNA extracted from faeces (both at time 0 and 48h after last gavage) and mucus from the ileum and colon of WT and ApcMin/+ mice pretreated or not with antibiotics (ABX) and then monocolonized with L. lactis (n = 4 mice/group). e,f, 11 weeks old ApcMin/+ mice treated with broths not fermented (Veh) or fermented by L. lactis (SUP) in the presence of ABX (n = 6 mice/group). e, tumour multiplicity in the small intestine normalized to vehicle-treated ApcMin/+ mice. f, Area and maximum diameter of ileal dysplastic lesions normalized to the total number lesions per mouse. Box plots show the interquartile range, median value and whiskers min to max. g, Cell proliferation assay on mouse CRC cell lines treated or not with sodium lactate at different concentrations for 48h. t0 is the signal from cells at the time of stimulation. Two independent experiments were performed with consistent results. Data presented as means ± s.d. of a representative experiment (n = 6 biologically independent samples). P values were determined by two-tailed unpaired Mann-Whitney U test (e,f), one-way ANOVA with Bonferroni post-test (b,g) or Kruskal-Wallis test with Dunn post-test (c,d). Data are presented as means ± s.e.m. in a-f.
Extended Data Fig. 8 H. biformis does not colonize the mouse gut but its spent medium (SUP) increases histone H3 acetylation, reduces NFATc3 activation and proliferation in vitro on human colorectal cancer cell lines.
a, Cell proliferation assay on human CRC cell lines treated or not (NT) with broth fermented by F. PB1 (F. PB1 SUP) or by H. biformis (H. biformis SUP) and with a mix of SCFAs as a control. t0 is the signal from cells at the time of stimulation. Two independent experiments were performed with consistent results. Data are presented as means ± s.d. of a representative experiment (n = 6 biologically independent samples). b, Densitometric quantification of WB in Fig. 6e showing NFATc3 and H3K27 acetylation (normalized to vinculin). Two or three independent experiments were performed with consistent results (n = 2 or n = 3 biologically independent experiments). c-e, ApcMin/+ mice received vehicle (Veh) or H. biformis from week 8 to 10 (n = 5 mice/group). c, tumour multiplicity in the small intestine normalized to vehicle-treated ApcMin/+ mice. d, Area and maximum diameter (axis lenght) of ileal dysplastic lesions normalized to the total number of lesions per mouse. Box plots show the interquartile range, median value and whiskers min to max. e, qPCR of H. biformis abundance normalized to panbacterial primers targeting the 16S rRNA gene (UNI 16S) in bacterial DNA extracted from faeces and mucus from the ileum. P values were determined by one-way ANOVA using Bonferroni (a) or Tukey (b, NFATc3 HT29 and H3K27 Caco2) post-test, by two-tailed unpaired Student’s t-test (b, H3K27 HT29) or by two-tailed Mann-Whitney U test (c-e). Data are represented as means ± s.e.m. in b-e.
Extended Data Fig. 9 Treatments with SCFAs or F. PB1 spent medium (SUP) increase histone H3 acetylation, reduce NFATc3 activation and proliferation in human CRC samples.
a-c, Human colon tumour samples (hCRC) incubated ex vivo with broth fermented with F. PB1 (SUP) or non-fermented (Veh); with medium alone (NT) or with SCFAs MIX. a, Representative immunohistochemistry of one out of three independent experiments of H3K27 acetylation and NFATc3 expression. Scale bars, 50 μm. b,c, Representative images of ileal dysplastic lesions stained with Ki67. Scale bars, 50 μm. Three independent experiments were performed with consistent results. Dot plots show the percentage of Ki67 positive cells relative to vehicle (b) or to NT (c). Data from three independent experiments (b, n = 7; c, n = 5 biologically independent samples), are represented as means ± s.e.m. and P values were determined by two-tailed unpaired Mann-Whitney U test.
Extended Data Fig. 10 Model.
Upon intestinal tumourigenesis selectively some bacterial species do not expand, due to different mucus composition or to competition with other bacteria and their corresponding metabolites. If these bacterial species are reintroduced, they induce the release of SCFAs. F. PB1 or H. biformis metabolic products act as HDAC inhibitors in the adenomas inducing an increase in acetylation and downmodulation of the calcineurin-NFATc3 pathway, which is involved in cell proliferation.
Supplementary information
Supplementary Information
Supplementary Tables 1–3 and flow cytometry gating strategy.
Source data
Source Data Fig. 1
Statistical Source Data.
Source Data Fig. 2
Statistical Source Data.
Source Data Fig. 3
Statistical Source Data.
Source Data Fig. 4
Statistical Source Data.
Source Data Fig. 4
Unprocessed western blot.
Source Data Fig. 5
Statistical Source Data.
Source Data Fig. 6
Statistical Source Data.
Source Data Fig. 6
Unprocessed western blot.
Source Data Extended Data Fig. 1
Statistical Source Data.
Source Data Extended Data Fig. 2
Statistical Source Data.
Source Data Extended Data Fig. 2
Unprocessed western blot.
Source Data Extended Data Fig. 3
Statistical Source Data.
Source Data Extended Data Fig. 4
Statistical Source Data.
Source Data Extended Data Fig. 5
Statistical Source Data.
Source Data Extended Data Fig. 5
Unprocessed western blot.
Source Data Extended Data Fig. 7
Statistical Source Data.
Source Data Extended Data Fig. 8
Statistical Source Data.
Source Data Extended Data Fig. 9
Statistical Source Data.
Rights and permissions
About this article
Cite this article
Zagato, E., Pozzi, C., Bertocchi, A. et al. Endogenous murine microbiota member Faecalibaculum rodentium and its human homologue protect from intestinal tumour growth. Nat Microbiol 5, 511–524 (2020). https://doi.org/10.1038/s41564-019-0649-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0649-5