Abstract
A major form of transcriptional regulation in bacteria occurs through the exchange of the primary σ factor of RNA polymerase (RNAP) with an alternative extracytoplasmic function (ECF) σ factor1. ECF σ factors are generally intrinsically active and are retained in an inactive state via the sequestration into σ factor–anti-σ factor complexes until their action is warranted2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20. Here, we report a previously uncharacterized mechanism of transcriptional regulation that relies on intrinsically inactive ECF σ factors, the activation of which and interaction with the β′-subunit of RNAP depends on σ factor phosphorylation. In Vibrio parahaemolyticus, the threonine kinase PknT phosphorylates the σ factor EcfP, which results in EcfP activation and expression of an essential polymyxin-resistant regulon. EcfP phosphorylation occurs at a highly conserved threonine residue, Thr63, positioned within a divergent region in the σ2.2 helix. Our data indicate that EcfP is intrinsically inactive and unable to bind the β′-subunit of RNAP due to the absence of a negatively charged DAED motif in this region. Furthermore, our results indicate that phosphorylation at residue Thr63 mimics this negative charge and licenses EcfP to interact with the β′-subunit in the formation of the RNAP holoenzyme, which in turn results in target gene expression. This regulatory mechanism is a previously unrecognized paradigm in bacterial signal transduction and transcriptional regulation, and our data suggest that it is widespread in bacteria.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All transcriptomics data are listed in Supplementary Tables 4–7 (file: Supplementary Table 4 (WT/ΔecfP chromosome I); Supplementary Table 5 (WT/ΔecfP chromosome II); Supplementary Table 6 (wt/ΔpknT chromosome I); Supplementary Table 7 (WT/ΔpknT chromosome II). Raw proteomics data are presented in Supplementary Tables 8 and 9. Raw data for bioinformatics analysis are provided in Supplementary Data 1–4. Source data for Figs. 1, 2 and 4 can be found in Source Data Fig. 1, Source Data Fig. 2 and Source Data Fig. 4, respectively. Raw source MS label-free quantification peptide intensity data regarding proteomics data are provided in Supplementary Tables 8–11. Supplementary Table 8 relates to Fig. 2g; Supplementary Table 9 to Fig. 2n; Supplementary Table 10 to Extended Data Figs. 1d and 4c; and Supplementary Table 11 to Extended Data Fig. 3e. Additional raw and analysed data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Mascher, T. Signaling diversity and evolution of extracytoplasmic function (ECF) σ factors. Curr. Opin. Microbiol. 16, 148–155 (2013).
Heinrich, J. & Wiegert, T. Regulated intramembrane proteolysis in the control of extracytoplasmic function sigma factors. Res. Microbiol. 160, 696–703 (2009).
Chaba, R., Grigorova, I. L., Flynn, J. M., Baker, T. A. & Gross, C. A. Design principles of the proteolytic cascade governing the SigmaE-mediated envelope stress response in Escherichia coli: keys to graded, buffered, and rapid signal transduction. Genes Dev. 21, 124–136 (2007).
Campagne, S. et al. Structural basis for sigma factor mimicry in the general stress response of Alphaproteobacteria. Proc. Natl Acad. Sci USA. 109, E1405–E1414 (2012).
Herrou, J., Rotskoff, G., Luo, Y., Roux, B. & Crosson, S. Structural basis of a protein partner switch that regulates the general stress response of α-proteobacteria. Proc. Natl Acad. Sci. USA 109, E1415–E1423 (2012).
Wecke, T. et al. Extracytoplasmic function σ factors of the widely distributed group ECF41 contain a fused regulatory domain. Microbiologyopen 1, 194–213 (2012).
Gómez-Santos, N., Pérez, J., Sánchez-Sutil, M. C., Moraleda-Muñoz, A. & Muñoz-Dorado, J. CorE from Myxococcus xanthus is a copper-dependent RNA polymerase sigma factor. PLoS Genet. 7, e1002106 (2011).
Wu, H. et al. The role of C-terminal extensions in controlling ECF σ factor activity in the widely conserved groups ECF 41 and ECF 42. Mol. Microbiol. 12, 498–514 (2019).
Mettrick, K. A. & Lamont, I. L. Different roles for anti-sigma factors in siderophore signalling pathways of Pseudomonas aeruginosa. Mol. Microbiol. 74, 1257–1271 (2009).
Ochs, M. et al. Regulation of citrate-dependent iron transport of Escherichia coli: FecR is required for transcription activation by Feel. Mol. Microbiol. 15, 119–132 (1995).
Paget, M. S. B., Chamberlin, L., Atrih, A., Foster, S. J. & Buttner, M. J. Evidence that the extracytoplasmic function sigma factor σE is required for normal cell wall structure in Streptomyces coelicolor A3(2). J. Bacteriol. 181, 204–211 (1999).
Hong, H. J., Paget, M. S. B. & Buttner, M. J. A signal transduction system in Streptomyces coelicolor that activates expression of a putative cell wall glycan operon in response to vancomycin and other cell wall-specific antibiotics. Mol. Microbiol. 44, 1199–1211 (2002).
Sohn, J., Grant, R. A. & Sauer, R. T. Allosteric activation of DegS, a stress sensor PDZ protease. Cell 131, 572–583 (2007).
Wilken, C., Kitzing, K., Kurzbauer, R., Ehrmann, M. & Clausen, T. Crystal structure of the DegS stress sensor. Cell 117, 483–494 (2004).
Chaba, R. et al. Signal integration by DegS and RseB governs the σE-mediated envelope stress response in Escherichia coli. Proc. Natl Acad. Sci. USA 108, 2106–2111 (2011).
Campbell, E. A. et al. A conserved structural module regulates transcriptional responses to diverse stress signals in bacteria. Mol. Cell 27, 793–805 (2007).
Zdanowski, K. et al. Assignment of the zinc ligands in RsrA, a redox-sensing ZAS protein from Streptomyces coelicolor. Biochemistry 45, 8294–8300 (2006).
Trepreau, J. et al. Spectroscopic characterization of the metal-binding sites in the periplasmic metal-sensor domain of CnrX from Cupriavidus metallidurans CH34. Biochemistry 50, 9036–9045 (2011).
Trepreau, J. et al. Structural basis for metal sensing by CnrX. J. Mol. Biol. 408, 766–779 (2011).
Francez-Charlot, A. et al. Sigma factor mimicry involved in regulation of general stress response. Proc. Natl Acad. Sci. USA 106, 3467–3472 (2009).
Staroń, A. et al. The third pillar of bacterial signal transduction: classification of the extracytoplasmic function (ECF) σ factor protein family. Mol. Microbiol. 74, 557–581 (2009).
Jogler, C. et al. Identification of proteins likely to be involved in morphogenesis, cell division, and signal transduction in Planctomycetes by comparative genomics. J. Bacteriol. 194, 6419–6430 (2012).
Bayer-Santos, E. et al. Xanthomonas citri T6SS mediates resistance to Dictyostelium predation and is regulated by an ECF σ factor and cognate Ser/Thr kinase. Environ. Microbiol. 20, 1562–1575 (2018).
McCarter, L. The multiple identities of Vibrio parahaemolyticus. J. Mol. Microbiol. Biotech. 1, 51–57 (1999).
Muraleedharan, S., Freitas, C., Mann, P., Glatter, T. & Ringgaard, S. A cell length-dependent transition in MinD-dynamics promotes a switch in division-site placement and preservation of proliferating elongated Vibrio parahaemolyticus swarmer cells. Mol. Microbiol. 109, 365–384 (2018).
Freitas, C., Glatter, T. & Ringgaard, S. The release of a distinct cell type from swarm colonies facilitates dissemination of Vibrio parahaemolyticus in the environment. ISME J. 14, 230–244 (2020).
Letchumanan, V., Chan, K. G. & Lee, L. H. Vibrio parahaemolyticus: a review on the pathogenesis, prevalence, and advance molecular identification techniques. Front. Microbiol. 5, 705 (2014).
Lane, W. J. & Darst, S. A. Molecular evolution of multisubunit RNA polymerases: structural analysis. J. Mol. Biol. 395, 686–704 (2010).
Joo, D. M., Ng, N. & Calendar, R. A σ32 mutant with a single amino acid change in the highly conserved region 2.2 exhibits reduced core RNA polymerase affinity. Proc. Natl Acad. Sci. USA 94, 4907–4912 (1997).
Sharp, M. M. et al. The interface of σ with core RNA polymerase is extensive, conserved, and functionally specialized. Genes Dev. 13, 3015–3026 (1999).
Wilson, M. J. & Lamont, I. L. Mutational analysis of an extracytoplasmic-function sigma factor to investigate its interactions with RNA polymerase and DNA. J. Bacteriol. 188, 1935–1942 (2006).
Li, L., Fang, C., Zhuang, N., Wang, T. & Zhang, Y. Structural basis for transcription initiation by bacterial ECF σ factors. Nat. Commun. 10, 1153 (2019).
Heering, J., Alvarado, A. & Ringgaard, S. Induction of cellular differentiation and single cell imaging of Vibrio parahaemolyticus swimmer and swarmer cells. J. Vis. Exp. 123, e55842 (2017).
Ringgaard, S., Schirner, K., Davis, B. M. & Waldor, M. K. A family of ParA-like ATPases promotes cell pole maturation by facilitating polar localization of chemotaxis proteins. Genes Dev. 25, 1544–1555 (2011).
Ringgaard, S. et al. ParP prevents dissociation of CheA from chemotactic signaling arrays and tethers them to a polar anchor. Proc. Natl Acad. Sci. USA 111, E255–E264 (2013).
Alvarado, A. et al. Coupling chemosensory array formation and localization. eLife 6, e31058 (2017).
Finn, R. D., Clements, J. & Eddy, S. R. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 39, W29–W37 (2011).
Sievers, F. & Higgins, D. G. in Multiple Sequence Alignment Methods (ed. Russell, D. J.) Ch. 6 (Humana Press, 2014).
Altschul, S. F., Gertz, E. M., Agarwala, R., Schäffer, A. A. & Yu, Y. K. PSI-BLAST pseudocounts and the minimum description length principle. Nucleic Acids Res. 37, 815–824 (2009).
Nguyen, L. T., Schmidt, H. A., Von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Barik, S., Sureka, K., Mukherjee, P., Basu, J. & Kundu, M. RseA, the SigE specific anti-sigma factor of Mycobacterium tuberculosis, is inactivated by phosphorylation-dependent ClpC1P2 proteolysis. Mol. Microbiol. 75, 592–606 (2010).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 44, W242–W245 (2016).
Huang, X., Pinto, D., Fritz, G. & Mascher, T. Environmental sensing in Actinobacteria: a comprehensive survey on the signaling capacity of this phylum. J. Bacteriol. 197, 2517–2535 (2015).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Murakami, K. S. Structural biology of bacterial RNA polymerase. Biomolecules 5, 848–864 (2015).
Helmann, J. D. & Chamberlin, M. J. Structure and function of bacterial sigma factors. Annu. Rev. Biochem. 57, 839–872 (1988).
Werner, F. & Grohmann, D. Evolution of multisubunit RNA polymerases in the three domains of life. Nat. Rev. Microbiol. 9, 85–98 (2011).
Yuan, J., Jin, F., Glatter, T. & Sourjik, V. Osmosensing by the bacterial PhoQ/PhoP two-component system. Proc. Natl Acad. Sci. USA 114, E10792–E10798 (2017).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Acknowledgements
The authors thank L. Søgaard-Andersen and T. Mascher for helpful comments on the manuscript. This work was supported by the Max Planck Society (S.R.), the LOEWE Program of the State of Hesse (SYNMIKRO) (G.F.), and the German Federal Ministry of Education and Research (grant no. 031L0010B) funded in the framework of the ERASynBio initiative (G.F.).
Author information
Authors and Affiliations
Contributions
S.C.I. carried out the majority of the experimental work. T.G. and P.M. contributed to the experimental work and its data analyses. G.F. and D.C.-P. carried out the bioinformatics analysis, and S.C.I. and S.R. contributed to the bioinformatics analysis. S.R. and G.F. conceived the study. S.R., G.F., S.C.I. and D.C.-P. designed the research and experiments and analysed the data. D.K. contributed to the data analyses. K.S. contributed to the data interpretation and experimental design. S.R. and G.F. wrote the manuscript. S.C.I., D.C.-P. and K.S. contributed to writing the manuscript. All authors read and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Phosphorylation of EcfP by membrane spanning kinase PknT.
a, Schematic showing the domain architecture of PknT. b, Membrane topology assay of PknT. E. coli cells expressing PknT N- and C-terminally translationally fused to PhoA-LacZα. The position of the fusion is indicated in the figure. Cells were propagated on indicator medium including the two chromogenic substrates, Red-Gal (indicating LacZ galactosidase activity) and X-Pho (indicating phosphatase activity). Blue coloration of the colonies is the result of high phosphatase activity, indicating a periplasmic location of the fusion point. Red coloration of the colonies shows a high galactosidase activity, indicating cytosolic location of the fusion point. c, Schematic showing the orientation of PknT in the inner membrane. d, Bar graph showing summed detected peptide intensities of total V. parahaemolyticus σ factors, EcfP and PknT. ND, non-detected. e, Bar graph showing the ratio between non-phosphorylated and phosphorylated EcfP peptide in wild-type cells and cells overexpressing PknT (wt + PknT). f, Extracted ion chromatogram of a non-phosphorylated EcfP peptide and detected EcfP phosphopeptide in wild-type and ΔpknT strain backgrounds. b, f, Each experiment was repeated independently a minimum of three times with similar or identical results. d-e, Data are mean ± s.e.m., n = 3 independent biological replicates. Statistical significance was calculated in relation to the indicated samples using a two-sided unpaired t-test (***P<0.001).
Extended Data Fig. 2 The PknT/EcfP system is specifically required for polymyxin resistance of V. parahaemolyticus.
a, Spot dilution assay of V. parahaemolyticus wild-type and mutant variants on LB growth medium in the presence and absence of indicated stress compounds, or spotted on LB medium subsequent to heat stress. b, Spot dilution assay of V. parahaemolyticus wild-type and mutant variants on LB growth medium in the presence and absence of Polymyxin E. c, EcfP and PknT function in the same signaling pathway. Spot dilution assay of V. parahaemolyticus wild-type and mutant variants on LB growth medium in the presence and absence of Polymyxin B. a-c, Each experiment was repeated independently a minimum of three times with similar or identical results.
Extended Data Fig. 3 Regulation of gene expression by the PknT/EcfP system.
a, MA plots for gene expression in wild-type versus ΔpknT, wherein M = log2(R/G) and A = 0.5*log2(RG). R and G values were calculated as the mean RPKM of two biological replicates of the mutant and wild-type respectively. Genes highlighted in black are up- or downregulated more than four fold. Gene highlighted in green represents vpa0879. b, Venn diagrams showing overlaps among genes with up-regulated or down-regulated transcript abundance in response to deletion of ecfP or pknT relative to wild-type. c, Table listing the genes down-regulated in wild-type by a minimum of 4 fold with respect to ΔecfP and ΔpknT. d, Table listing the genes up-regulated in wild-type with respect to ΔecfP and ΔpknT. P-value was determined by Students t-test. e, Bar graph showing the fold downregulation of VPA0879 protein levels in the absence of PknT. Data are mean ± s.e.m., n = 4 independent biological replicates. Statistical significance was calculated in relation to wild-type using a two-sided unpaired t-test (***P<0.001).
Extended Data Fig. 4 PknT catalytic activity is required for PknT mediated phosphorylation and activation of EcfP.
a, EcfPT63A prevents phosphorylation at EcfP residue Thr63. Upon IP-MS experiments of cells expressing sfGFP-EcfPT63A, label-free quantification was performed. Bar graph shows summed peptide intensities of total EcfPT63A and phosphorylated EcfPT63A peptides. ND, non-detected. b, A strain expressing EcfPT63E is less sensitive to polymyxin stress than an ecfPT63A strain. Spot dilution assay of V. parahaemolyticus wild-type and mutant variants on LB growth medium in the presence and absence of Polymyxin B. c, Bar graph showing the ratio of VPA0879 levels between wt and EcfPT63 variants. d, Catalytic activity of PknT is required for its phosphorylation of EcfP. Extracted ion chromatograms of a non-phosphorylated EcfP peptide and detected EcfP phosphopeptide in wild-type and pknTD202A strain backgrounds. a, c, Data are mean ± s.e.m., n = 3 independent biological replicates. Statistical significance was calculated in relation to indicated samples using a two-sided unpaired t-test (*P<0.05, ***P<0.001). b, d, Each experiment was repeated independently a minimum of three times with similar or identical results.
Extended Data Fig. 5 Different members of ECF43 are associated to a distinct protein kinase domain architecture.
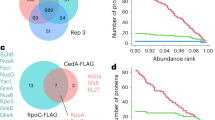
The phylogenetic tree on the left side is a fragment of the ECF tree of the homologues of EcfP show in Fig. 3a. For simplicity, only the closest variants to EcfP are shown. Group #1 depicts the position and the Pfam domains found in the STK associated to each ECF. Group #2 shows the order of the organisms of origin of the proteins. The extracytoplasmic input domains, located in C-termini from the protein kinase domain, vary across protein kinases, indicating a potential response to different stimuli or a different sensing mechanism. The number of genomes and methods used for generation of the tree can be found in the Methods section. Raw version of the tree in Newick format is found in Supplementary Data 1. The genomes can be found in Supplementary Data 2. The number of genomes used is 1,510. The number of marker genes is 1,532, all homologues of EcfP with a STK in encoded in < 5,000bp from their coding sequence. In order to limit the redundancy of this dataset, only one representative from each 98% identity group (see Mothods section) is considered for the phylogenetic tree. This makes a total of 89 98% identity groups represented in this view. IQ-TREE 1.6.1 was used to generate the tree.
Extended Data Fig. 6 The negatively charged DAED motif has been replaced by non-charged amino acids and a highly conserved Thr in EcfP and group ECF43.
Conservation of the DAED motif and other negatively charged residues of σ2.2, responsible for mediating interaction to the β’ subunit of the core RNAP enzyme. Amino acid coordinates refer to SigH from M. tuberculosis. E55 of SigH corresponds to Thr63 of EcfP.
Extended Data Fig. 7 Phylogenetic tree of species encoding EcfP homologs in their genomes.
The phylogenetic tree represents the phylogenetic distance (as calculated by IQ-tree using default parameters over a MSA computed by ClustalO) of the 16S rDNA sequences of representative and reference organisms (as defined by NCBI), containing an ECF σ factor with Ser or Thr in a position equivalent to Thr63 in EcfP of V. parahaemolyticus. In most of the cases there is a perfect correlation between tree architecture and class of the organims, indicated by the shades of the tree and the accompanying labels. The tree was rooted in the 16S rDNA of Bacillus subtilis (used as outlier), since Firmicutes do not contain any ECF sequence with Ser/Thr63. The heatmap on the right shows the frequency of ECF variants with S63, T63, with an extended region between σ2.1 and σ2.2 and their membership to group ECF43. Frequencies identical to 1 (=100%) indicate that all ECFs in this species exhibit the respective feature, while frequencies below 1 indicate that there exist other EcfP homologs in the species, which do not exhibit the given feature. The number of genomes and methods used in generation of the tree can be found in the Methods section. Raw version of the tree in Newick format is found in Supplementary Data 3. The genomes can be found in Supplementary Data 4. The number of genomes used is 138 genomes + outgroup Bacillus subtilis subsp. subtilis str. 168 (NCBI Assembly GCF_000009045.1). The number of marker genes is DNA annotated as ‘16S ribosomal RNA’, selecting only one per genome and discarding the ones with ambiguous characters (N) or with a length ≤ 1200bp. IQ-TREE 1.6.1 was used to generate the tree.
Supplementary information
Supplementary Information
Supplementary Tables 1–3 and Supplementary References.
Supplementary Tables
Supplementary Tables 4–11.
Supplementary Data 1
Raw version of the tree in Fig. 3a in Newick format. The genomes can be found in Supplementary Data 2.
Supplementary Data 2
ECF σ factors and STKs retrieved during this study. The ECF group is only shown for groups known to be linked to STKs (ECF43, ECF59 and ECF60). ‘Length extended region’ refers to the length of the region between dashed lines in Fig. 3b. ‘aa63’ refers to the amino acid in the position equivalent to T63 in EcfP. ‘DAED’ refers to the amino acids that occupy the same position as consensus DAED in canonical ECFs and STTA in EcfP. The ‘98% identity group’ groups together the pairs of ECF-STK with a combined identity of ≥98% (see Methods). One example from each 98% identity group was chosen for the phylogenetic tree in Fig. 3a.
Supplementary Data 3
Raw version of the tree in Extended Data Fig. 7 in Newick format. The genomes can be found in Supplementary Data 4.
Supplementary Data 4
16S rDNA sequence, organisms and genome assemblies used for the phylogenetic tree of Extended Data Fig. 7.
Source data
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 2
Unprocessed western blots.
Source Data Fig. 4
Unprocessed western blots.
Rights and permissions
About this article
Cite this article
Iyer, S.C., Casas-Pastor, D., Kraus, D. et al. Transcriptional regulation by σ factor phosphorylation in bacteria. Nat Microbiol 5, 395–406 (2020). https://doi.org/10.1038/s41564-019-0648-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0648-6