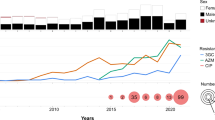
Abstract
Despite the sporadic detection of fluoroquinolone-resistant Shigella in Asia in the early 2000s and the subsequent global spread of ciprofloxacin-resistant (cipR) Shigella sonnei from 2010, fluoroquinolones remain the recommended therapy for shigellosis1,2,3,4,5,6,7. The potential for cipR S. sonnei to develop resistance to alternative second-line drugs may further limit future treatment options8. Here, we aim to understand the evolution of novel antimicrobial resistant (AMR) S. sonnei variants after introduction into Vietnam. We found that cipR S. sonnei displaced the resident ciprofloxacin-susceptible (cipS) lineage while rapidly acquiring additional resistance to multiple alternative antimicrobial classes. We identified several independent acquisitions of extensively drug-resistant/multidrug-resistant-inducing plasmids, probably facilitated by horizontal transfer from commensals in the human gut. By characterizing commensal Escherichia coli from Shigella-infected and healthy children, we identified an extensive array of AMR genes and plasmids, including an identical multidrug-resistant plasmid isolated from both S. sonnei and E. coli in the gut of a single child. We additionally found that antimicrobial usage may impact plasmid transfer between commensal E. coli and S. sonnei. These results suggest that, in a setting with high antimicrobial use and a high prevalence of AMR commensals, cipR S. sonnei may be propelled towards pan-resistance by adherence to outdated international treatment guidelines.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All raw genomic data that support the findings of this study have been deposited in the European Nucleotide Archive (ENA, project PRJEB30967) and can be accessed via this link. Details about the accession numbers of S. sonnei isolates are provided in Supplementary Table 1. The S. sonnei reference genome Ss046 chromosome (accession no. NC_007382), virulence plasmid pSs046 (accession no. NC_007385.1)<q> and three small plasmids commonly found in global lineage III S. sonnei—spA (accession no. NC_009345.1), spB (accession no. NC_009346.1) and spC (accession no. NC_009347.1)—were downloaded from GenBank. Raw MinION reads for plasmid p01_0123 are deposited in ENA (accession no. ERS3050922). Source data for the main figures (Figs. 2a, 3a and 4) and Extended Data Figs. 1 and 2 are provided with the paper. Additional data that support the findings of this study are available from the corresponding author upon request.
References
Pazhani, G. P., Ramamurthy, T., Mitra, U., Bhattacharya, S. K. & Niyogi, S. K. Species diversity and antimicrobial resistance of Shigella spp. isolated between 2001 and 2004 from hospitalized children with diarrhoea in Kolkata (Calcutta), India. Epidemiol. Infect. 133, 1089–1095 (2005).
Shakya, G., Acharya, J., Adhikari, S. & Rijal, N. Shigellosis in Nepal: 13 years review of nationwide surveillance. J. Health Popul. Nutr. 35, 36 (2016).
Von Seidlein, L. et al. A multicentre study of Shigella diarrhoea in six Asian countries: disease burden, clinical manifestations and microbiology. PLoS Med. 3, 1556–1569 (2006).
Pazhani, G. P. et al. Molecular characterization of multidrug-resistant Shigella species isolated from epidemic and endemic cases of shigellosis in India. J. Med. Microbiol. 57, 856–863 (2008).
De Lappe, N., O’Connor, J., Garvey, P., McKeown, P. & Cormican, M. Ciprofloxacin-resistant Shigella sonnei associated with travel to India. Emerg. Infect. Dis. 21, 894–896 (2015).
Bowen, A. et al. Importation and domestic transmission of Shigella sonnei resistant to ciprofloxacin—United States, May 2014–February 2015. MMWR Morb. Mortal. Wkly Rep. 64, 318–320 (2015).
Nüesch-Inderbinen, M. et al. Shigella antimicrobial drug resistance mechanisms, 2004–2014. Emerg. Infect. Dis. 22, 1083–1085 (2016).
Chung The, H. et al. Introduction and establishment of fluoroquinolone-resistant Shigella sonnei into Bhutan. Microb. Genom. 1, 000042 (2015).
Rajpara, N. et al. Molecular analysis of multidrug resistance in clinical isolates of Shigella spp. from 2001–2010 in Kolkata, India: role of integrons, plasmids and topoisomerase mutations. Infect. Drug Resist. 11, 87–102 (2018).
Chung The, H. et al. South Asia as a reservoir for the global spread of ciprofloxacin-resistant Shigella sonnei: a cross-sectional study. PLoS Med. 13, 1–12 (2016).
Holt, K. E. et al. Tracking the establishment of local endemic populations of an emergent enteric pathogen. Proc. Natl Acad. Sci. USA 110, 17522–17527 (2013).
Praszkier, J. & Pittard, A. J. Control of replication in I-complex plasmids. Plasmid 53, 97–112 (2005).
Seral, C. et al. Characterisation of a CTX-M-15-producing Shigella sonnei in a Spanish patient who had not travelled abroad. Enferm. Infecc. Microbiol. Clin. 30, 469–471 (2012).
Ma, Q. et al. A waterborne outbreak of Shigella sonnei with resistance to azithromycin and third-generation cephalosporins in China in 2015. Antimicrob. Agents Chemother. 61, e00308-17 (2017).
Kim, J. S. et al. Complete nucleotide sequence of the IncI1 plasmid pSH4469 encoding CTX-M-15 extended-spectrum β-lactamase in a clinical isolate of Shigella sonnei from an outbreak in the Republic of Korea. Int. J. Antimicrob. Agents 44, 533–537 (2014).
Folster, J. P. et al. Identification and characterization of CTX-M-producing Shigella isolates in the United States. Antimicrob. Agents Chemother. 54, 2269–2270 (2010).
Allué-Guardia, A. et al. Closed genome and comparative phylogenetic analysis of the clinical multidrug resistant Shigella sonnei strain 866. Genome Biol. Evol. 30, 469–471 (2018).
Zurita, J., Ortega-Paredes, D. & Barba, P. First description of Shigella sonnei harboring bla CTX-M-55 outside Asia. J. Microbiol. Biotechnol. 26, 2224–2227 (2016).
Lee, W. et al. CTX-M-55-type extended-spectrum β-lactamase-producing Shigella sonnei isolated from a Korean patient who had travelled to China. Ann. Lab. Med. 33, 141–144 (2013).
Qu, F. et al. Plasmid-encoding extended-spectrum β-lactamase CTX-M-55 in a clinical Shigella sonnei strain, China. Future Microbiol. 9, 1143–1150 (2014).
Bratoeva, M. P. & John, J. F. In vivo R–plasmid transfer in a patient with a mixed infection of Shigella dysentery. Epidemiol. Infect. 112, 247–252 (1994).
Rashid, H. & Rahman, M. Possible transfer of plasmid mediated third generation cephalosporin resistance between Escherichia coli and Shigella sonnei in the human gut. Infect. Genet. Evol. 30, 15–18 (2015).
Duong, V. T. et al. No clinical benefit of empirical antimicrobial therapy for pediatric diarrhea in a high-usage, high-resistance setting. Clin. Infect. Dis. 66, 504–511 (2018).
Magiorakos, A. P. et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 18, 268–281 (2012).
Thiem, V. D. et al. Detection of Shigella by a PCR assay targeting the ipaH gene suggests increased prevalence of shigellosis in Nha Trang, Vietnam. J. Clin. Microbiol. 42, 2031–2035 (2004).
Thompson, C. N. et al. A cohort study to define the age-specific incidence and risk factors of Shigella diarrhoeal infections in Vietnamese children: A study protocol. BMC Public Health 14, 1289 (2014).
Holt, K. E. et al. Shigella sonnei genome sequencing and phylogenetic analysis indicate recent global dissemination from Europe. Nat. Genet. 44, 1056–1059 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Croucher, N. J. et al. Rapid phylogenetic analysis of large samples of recombinant bacterial whole genome sequences using Gubbins. Nucleic Acids Res. 43, e15 (2015).
Nguyen, L. T., Schmidt, H. A., Von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Rambaut, A., Lam, T. T., Max Carvalho, L. & Pybus, O. G. Exploring the temporal structure of heterochronous sequences using TempEst (formerly Path-O-Gen). Virus Evol. 2, vew007 (2016).
Drummond, A. J. & Rambaut, A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol. Biol. 7, 214 (2007).
Drummond, A. J., Rambaut, A., Shapiro, B. & Pybus, O. G. Bayesian coalescent inference of past population dynamics from molecular sequences. Mol. Biol. Evol. 22, 1185–1192 (2005).
Drummond, A. J., Ho, S. Y. W., Phillips, M. J. & Rambaut, A. Relaxed phylogenetics and dating with confidence. PLoS Biol. 4, e88 (2006).
Xie, W., Lewis, P. O., Fan, Y., Kuo, L. & Chen, M. H. Improving marginal likelihood estimation for Bayesian phylogenetic model selection. Syst. Biol. 60, 150–160 (2011).
Lartillot, N. & Philippe, H. Computing Bayes factors using thermodynamic integration. Syst. Biol. 55, 195–207 (2006).
Inouye, M. et al. SRST2: rapid genomic surveillance for public health and hospital microbiology labs. Genome Med. 6, 90 (2014).
Gupta, S. K. et al. ARG-annot, a new bioinformatic tool to discover antibiotic resistance genes in bacterial genomes. Antimicrob. Agents Chemother. 58, 212–220 (2014).
Carattoli, A. et al. In silico detection and typing of plasmids using plasmidfinder and plasmid multilocus sequence typing. Antimicrob. Agents Chemother. 58, 3895–3903 (2014).
García-Fernández, A. et al. Multilocus sequence typing of IncI1 plasmids carrying extended-spectrum β-lactamases in Escherichia coli and Salmonella of human and animal origin. J. Antimicrob. Chemother. 61, 1229–1233 (2008).
Zerbino, D. R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Assefa, S., Keane, T. M., Otto, T. D., Newbold, C. & Berriman, M. ABACAS: algorithm-based automatic contiguation of assembled sequences. Bioinformatics 25, 1968–1969 (2009).
Carver, T. et al. Artemis and ACT: viewing, annotating and comparing sequences stored in a relational database. Bioinformatics 24, 2672–2676 (2008).
Wood, D. E. & Salzberg, S. L. Kraken: ultrafast metagenomic sequence classification using exact alignments. Genome Biol. 15, R46 (2014).
Kado, C. I. & Liu, S. T. Rapid procedure for detection and isolation of large and small plasmids. J. Bacteriol. 145, 1365–1373 (1981).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Acknowledgements
This work was funded by a Senior Research Fellowship funded by the Wellcome Trust to S.B. (100087/Z/12/Z) and an Oak Leader Fellowship to D.P.T.
Author information
Authors and Affiliations
Contributions
P.T.D., M.A.R. and S.B. designed the study. P.T.D. performed data analysis and interpreted the results under the scientific guidance of M.A.R. and S.B. P.T.D. drafted the paper, with M.A.R. and S.B. revising and structuring the paper. P.T.D. and H.N.D.T. performed whole genome sequencing. P.T.D. and T.N.T.N. performed plasmid isolation, digestion and sequencing. P.T.D., F.A. and T.N.T.N. performed the plasmid conjugation experiments. H.T.T. performed basic microbiology work. H.C.T. and C.B. assisted in the design of laboratory experiments. D.V.T. recruited patients and performed the clinical work required for the study. H.C.T., C.B. and G.E.T. contributed to the editing of the paper. All authors read and approved the final draft.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Plasmid profiling of Shigella sonnei in Vietnam, 2014–2016.
Dendrogram shows the difference in plasmid electrophoresis patterns between the resident S. sonnei clade and the cipR S. sonnei clade in Vietnam. Black lines separate lanes that were not contiguous in a gel. Data were obtained from a single experiment. Cluster analysis was performed with Bionumerics by using the Jaccard coefficient and the unweighted pair group mathematical average (UPGMA) clustering algorithm.
Extended Data Fig. 2 Distribution of antimicrobial resistance genes and plasmid groups in Shigella sonnei and human commensal bacteria.
The first column highlights the four different isolate types in different colors. Fecal/rectal swab cultures with ciprofloxacin-resistant and ESBL-producing isolates are highlighted in turquoise and red (second and third columns, respectively). The remaining columns show the distribution of all antimicrobial resistance genes and plasmid groups identified in commensal bacteria and S. sonnei. AMR genes are grouped together based on the class of antimicrobial agents to which they are resistant, with different variants of an AMR gene shown in different shades of blue.
Extended Data Fig. 3 Root-to-tip regression for the maximum likelihood tree of Shigella sonnei in Vietnam, 2014–2016.
Each point on the plot corresponds to a measurement of genetic distance from the inferred root to each tip in the tree (n = 81 biologically independent samples). The solid line is the regression line fitted using the ordinary least squares method. The slope of the line is a crude estimate of the evolutionary rate, the x-intercept corresponds to the inferred date of the most recent common ancestor, and the R2 value measures the degree of clock-like behavior.
Supplementary information
Supplementary Table 1
Shigella sonnei isolates and their corresponding metadata.
Source data
Source Data Fig. 2a
Raw unprocessed gels associated with Fig. 2a.
Source Data Fig. 3a
Source data of antimicrobial resistance genes and plasmids in Shigella sonnei and human commensal bacteria associated with Fig. 3a.
Source Data Fig. 4
Source data of the conjugation efficiency of ESBL-encoding plasmids from human commensal E. coli to ciprofloxacin-resistant Shigella sonnei with and without supplementation with ciprofloxacin associated with Fig. 4.
Source Data Extended Data Fig. 1
Raw unprocessed gels associated with Extended Data Fig.1.
Source Data Extended Data Fig. 2
Source data of distribution of antimicrobial resistance genes and plasmid groups in Shigella sonnei and human commensal bacteria associated with Extended Data Fig. 2.
Rights and permissions
About this article
Cite this article
Thanh Duy, P., Thi Nguyen, T.N., Vu Thuy, D. et al. Commensal Escherichia coli are a reservoir for the transfer of XDR plasmids into epidemic fluoroquinolone-resistant Shigella sonnei. Nat Microbiol 5, 256–264 (2020). https://doi.org/10.1038/s41564-019-0645-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0645-9
This article is cited by
-
The impact of antibiotics on the gut microbiota of children recovering from watery diarrhoea
npj Antimicrobials and Resistance (2024)
-
The genomic epidemiology of shigellosis in South Africa
Nature Communications (2023)
-
Antimicrobial-resistant Shigella: where do we go next?
Nature Reviews Microbiology (2023)
-
Rapid emergence of extensively drug-resistant Shigella sonnei in France
Nature Communications (2023)
-
Emerging Increase in Colistin Resistance Rates in Escherichia coli and Salmonella enterica from Türkiye
Current Microbiology (2023)