Abstract
MicroRNAs (miRNAs) are increasingly recognized for their role in infection by bacterial pathogens, although the effect of each individual miRNA remains largely unknown. Here, we used a comparative genome-wide microscopy-based functional screening approach to identify miRNAs controlling infection by two bacterial pathogens—Salmonella enterica serovar Typhimurium and Shigella flexneri. Despite the similarities between these pathogens, we found infections to be controlled by largely non-overlapping subsets of miRNAs, seemingly reflecting different requirements prompted by their distinct intracellular lifestyles. By characterizing a small subset of miRNAs chosen among the strongest inhibitors of Shigella infection, we discovered that miR-3668, miR-4732-5p and miR-6073 exert a selective effect on Shigella infection by impairing bacterial actin-based motility by downregulating N-WASP. Additionally, by identifying let-7i-3p miRNA as a strong inhibitor of Salmonella replication and performing in-depth analysis of its mechanisms of action, we showed that this miRNA specifically inhibits Salmonella infection via modulation of endolysosomal trafficking and the vacuolar environment by targeting the host RGS2 protein. These findings illustrate two paradigms underlying miRNA-mediated regulation of bacterial infection, acting as part of the host response to infection, or as part of bacterial strategies to modulate the host environment and favour pathogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data supporting the findings of this study are available from the corresponding authors upon reasonable request. The demultiplexed FASTQ files and gene-wise quantifications have been deposited in NCBI’s Gene Expression Omnibus and are accessible through GEO Series accession numbers GSE53281 (mock-treated and Salmonella-infected samples) and GSE69660 (cells transfected with control miRNA or let-7i-3p mimic). Source data for Figs. 3e,i,k, 4f, 5f,g,i and 6h and Extended Data Figs. 3b,d, 5f,h,j and 6b are included in this article and its Supplementary Information files.
Code availability
Custom image analysis workflows implemented in Columbus or CellProfiler image analysis software are available upon reasonable request.
References
Bartel, D. P. Metazoan microRNAs. Cell 173, 20–51 (2018).
Jonas, S. & Izaurralde, E. Towards a molecular understanding of microRNA-mediated gene silencing. Nat. Rev. Genet. 16, 421–433 (2015).
Friedman, R. C., Farh, K. K., Burge, C. B. & Bartel, D. P. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 19, 92–105 (2009).
Aguilar, C., Mano, M. & Eulalio, A. MicroRNAs at the host–bacteria interface: host defense or bacterial offense. Trends Microbiol. 27, 206–218 (2019).
Duval, M., Cossart, P. & Lebreton, A. Mammalian microRNAs and long noncoding RNAs in the host-bacterial pathogen crosstalk. Semin. Cell. Dev. Biol. 65, 11–19 (2017).
Eulalio, A. & Mano, M. MicroRNA screening and the quest for biologically relevant targets. J. Biomol. Screen. 20, 1003–1017 (2015).
Lemons, D., Maurya, M. R., Subramaniam, S. & Mercola, M. Developing microRNA screening as a functional genomics tool for disease research. Front. Physiol. 4, 223 (2013).
Rodrigues Lopes, I., Silva, R. J., Caramelo, I., Eulalio, A. & Mano, M. Shedding light on microRNA function via microscopy-based screening. Methods 152, 55–64 (2019).
Huang, Q. et al. The microRNAs miR-373 and miR-520c promote tumour invasion and metastasis. Nat. Cell Biol. 10, 202–210 (2008).
Eulalio, A. et al. Functional screening identifies miRNAs inducing cardiac regeneration. Nature 492, 376–381 (2012).
Judson, R. L., Greve, T. S., Parchem, R. J. & Blelloch, R. MicroRNA-based discovery of barriers to dedifferentiation of fibroblasts to pluripotent stem cells. Nat. Struct. Mol. Biol. 20, 1227–1235 (2013).
Santhakumar, D. et al. Combined agonist–antagonist genome-wide functional screening identifies broadly active antiviral microRNAs. Proc. Natl Acad. Sci. USA 107, 13830–13835 (2010).
Maudet, C. et al. Functional high-throughput screening identifies the miR-15 microRNA family as cellular restriction factors for Salmonella infection. Nat. Commun. 5, 4718 (2014).
Collaborators, G. B. D. D. D. Estimates of global, regional, and national morbidity, mortality, and aetiologies of diarrhoeal diseases: a systematic analysis for the Global Burden of Disease Study 2015. Lancet. Infect. Dis. 17, 909–948 (2017).
Haselbeck, A. H. et al. Current perspectives on invasive nontyphoidal Salmonella disease. Curr. Opin. Infect. Dis. 30, 498–503 (2017).
Kotloff, K. L., Riddle, M. S., Platts-Mills, J. A., Pavlinac, P. & Zaidi, A. K. M. Shigellosis. Lancet 391, 801–812 (2018).
Steele-Mortimer, O. The Salmonella-containing vacuole: moving with the times. Curr. Opin. Microbiol. 11, 38–45 (2008).
Mellouk, N. & Enninga, J. Cytosolic access of intracellular bacterial pathogens: the Shigella paradigm. Front. Cell. Infect. Microbiol. 6, 35 (2016).
Agaisse, H. Molecular and cellular mechanisms of Shigella flexneri dissemination. Front. Cell. Infect. Microbiol. 6, 29 (2016).
Lambrechts, A., Gevaert, K., Cossart, P., Vandekerckhove, J. & Van Troys, M. Listeria comet tails: the actin-based motility machinery at work. Trends Cell Biol. 18, 220–227 (2008).
Agarwal, V., Bell, G. W., Nam, J. W. & Bartel, D. P. Predicting effective microRNA target sites in mammalian mRNAs. eLife 4, 05005 (2015).
Egile, C. et al. Activation of the CDC42 effector N-WASP by the Shigella flexneri IcsA protein promotes actin nucleation by Arp2/3 complex and bacterial actin-based motility. J. Cell Biol. 146, 1319–1332 (1999).
Suzuki, T. et al. Neural Wiskott-Aldrich syndrome protein (N-WASP) is the specific ligand for Shigella VirG among the WASP family and determines the host cell type allowing actin-based spreading. Cell. Microbiol. 4, 223–233 (2002).
Boujemaa-Paterski, R. et al. Listeria protein ActA mimics WASp family proteins: it activates filament barbed end branching by Arp2/3 complex. Biochemistry 40, 11390–11404 (2001).
Bierne, H. et al. WASP-related proteins, Abi1 and Ena/VASP are required for Listeria invasion induced by the Met receptor. J. Cell Sci. 118, 1537–1547 (2005).
Lara-Tejero, M. & Galan, J. E. Salmonella enterica serovar Typhimurium pathogenicity island 1-encoded type III secretion system translocases mediate intimate attachment to nonphagocytic cells. Infect. Immun. 77, 2635–2642 (2009).
Misselwitz, B. et al. Salmonella enterica serovar Typhimurium binds to HeLa cells via Fim-mediated reversible adhesion and irreversible type three secretion system 1-mediated docking. Infect. Immun. 79, 330–341 (2011).
Schlumberger, M. C. et al. Real-time imaging of type III secretion: Salmonella SipA injection into host cells. Proc. Natl Acad. Sci. USA 102, 12548–12553 (2005).
Menard, R., Sansonetti, P. J. & Parsot, C. Nonpolar mutagenesis of the ipa genes defines IpaB, IpaC, and IpaD as effectors of Shigella flexneri entry into epithelial cells. J. Bacteriol. 175, 5899–5906 (1993).
Woodard, G. E., Jardin, I., Berna-Erro, A., Salido, G. M. & Rosado, J. A. Regulators of G-protein-signaling proteins: negative modulators of G-protein-coupled receptor signaling. Int. Rev. Cell Mol. Biol. 317, 97–183 (2015).
Sethakorn, N., Yau, D. M. & Dulin, N. O. Non-canonical functions of RGS proteins. Cell. Signal. 22, 1274–1281 (2010).
Tuli, A. & Sharma, M. How to do business with lysosomes: Salmonella leads the way. Curr. Opin. Microbiol. 47, 1–7 (2018).
Garcia-del Portillo, F. & Finlay, B. B. Targeting of Salmonella typhimurium to vesicles containing lysosomal membrane glycoproteins bypasses compartments with mannose 6-phosphate receptors. J. Cell Biol. 129, 81–97 (1995).
Rathman, M., Barker, L. P. & Falkow, S. The unique trafficking pattern of Salmonella typhimurium-containing phagosomes in murine macrophages is independent of the mechanism of bacterial entry. Infect. Immun. 65, 1475–1485 (1997).
Santos, J. C. et al. The COPII complex and lysosomal VAMP7 determine intracellular Salmonella localization and growth. Cell. Microbiol. 17, 1699–1720 (2015).
Hang, H. C. et al. Mechanism-based probe for the analysis of cathepsin cysteine proteases in living cells. ACS Chem. Biol. 1, 713–723 (2006).
Meresse, S., Steele-Mortimer, O., Finlay, B. B. & Gorvel, J. P. The rab7 GTPase controls the maturation of Salmonella typhimurium-containing vacuoles in HeLa cells. EMBO J. 18, 4394–4403 (1999).
Brown, W. J., DeWald, D. B., Emr, S. D., Plutner, H. & Balch, W. E. Role for phosphatidylinositol 3-kinase in the sorting and transport of newly synthesized lysosomal enzymes in mammalian cells. J. Cell Biol. 130, 781–796 (1995).
Davidson, H. W. Wortmannin causes mistargeting of procathepsin D. Evidence for the involvement of a phosphatidylinositol 3-kinase in vesicular transport to lysosomes. J. Cell Biol. 130, 797–805 (1995).
Gaffet, P., Jones, A. T. & Clague, M. J. Inhibition of calcium-independent mannose 6-phosphate receptor incorporation into trans-Golgi network-derived clathrin-coated vesicles by wortmannin. J. Biol. Chem. 272, 24170–24175 (1997).
Ikeda, K. et al. Drug-induced phospholipidosis is caused by blockade of mannose 6-phosphate receptor-mediated targeting of lysosomal enzymes. Biochem. Biophys. Res. Commun. 377, 268–274 (2008).
Reaves, B. J., Bright, N. A., Mullock, B. M. & Luzio, J. P. The effect of wortmannin on the localisation of lysosomal type I integral membrane glycoproteins suggests a role for phosphoinositide 3-kinase activity in regulating membrane traffic late in the endocytic pathway. J. Cell Sci. 109, 749–762 (1996).
Brumell, J. H., Tang, P., Zaharik, M. L. & Finlay, B. B. Disruption of the Salmonella-containing vacuole leads to increased replication of Salmonella enterica serovar typhimurium in the cytosol of epithelial cells. Infect. Immun. 70, 3264–3270 (2002).
Collado-Romero, M., Arce, C., Ramirez-Boo, M., Carvajal, A. & Garrido, J. J. Quantitative analysis of the immune response upon Salmonella typhimurium infection along the porcine intestinal gut. Vet. Res. 41, 23 (2010).
Denzler, R. et al. Impact of microRNA levels, target-site complementarity, and cooperativity on competing endogenous RNA-regulated gene expression. Mol. Cell 64, 565–579 (2016).
Hon, L. S. & Zhang, Z. The roles of binding site arrangement and combinatorial targeting in microRNA repression of gene expression. Genome Biol. 8, R166 (2007).
Campellone, K. G. & Welch, M. D. A nucleator arms race: cellular control of actin assembly. Nat. Rev. Mol. Cell Biol. 11, 237–251 (2010).
Sunkavalli, U. et al. Analysis of host microRNA function uncovers a role for miR-29b-2-5p in Shigella capture by filopodia. PLoS Pathog. 13, e1006327 (2017).
Hume, P. J., Singh, V., Davidson, A. C. & Koronakis, V. Swiss Army pathogen: the Salmonella entry toolkit. Front. Cell. Infect. Microbiol. 7, 348 (2017).
Van Nhieu, G. T. & Romero, S. Common themes in cytoskeletal remodeling by intracellular bacterial effectors. Handb. Exp. Pharmacol. 235, 207–235 (2017).
Sullivan, B. M. et al. RGS4 and RGS2 bind coatomer and inhibit COPI association with Golgi membranes and intracellular transport. Mol. Biol. Cell 11, 3155–3168 (2000).
McGourty, K. et al. Salmonella inhibits retrograde trafficking of mannose-6-phosphate receptors and lysosome function. Science 338, 963–967 (2012).
Tawk, C. et al. Stress-induced host membrane remodeling protects from infection by non-motile bacterial pathogens. EMBO J. 37, e98529 (2018).
Papenfort, K. et al. Specific and pleiotropic patterns of mRNA regulation by ArcZ, a conserved, Hfq-dependent small RNA. Mol. Microbiol. 74, 139–158 (2009).
Eulalio, A., Frohlich, K. S., Mano, M., Giacca, M. & Vogel, J. A candidate approach implicates the secreted Salmonella effector protein SpvB in P-body disassembly. PLoS ONE 6, e17296 (2011).
Garcia-Feliz, C. et al. Salmonella enterica infections in Spanish swine fattening units. Zoonoses Public Health 54, 294–300 (2007).
Hussain, N. K. et al. Endocytic protein intersectin-l regulates actin assembly via Cdc42 and N-WASP. Nat. Cell Biol. 3, 927–932 (2001).
Sancak, Y. et al. The Rag GTPases bind raptor and mediate amino acid signaling to mTORC1. Science 320, 1496–1501 (2008).
McQuin, C. et al. CellProfiler 3.0: Next-generation image processing for biology. PLoS Biol. 16, e2005970 (2018).
Collado-Romero, M. et al. Quantitative proteomics and bioinformatic analysis provide new insight into the dynamic response of porcine intestine to Salmonella Typhimurium. Front. Cell. Infect. Microbiol. 5, 64 (2015).
Forstner, K. U., Vogel, J. & Sharma, C. M. READemption-a tool for the computational analysis of deep-sequencing-based transcriptome data. Bioinformatics 30, 3421–3423 (2014).
Otto, C., Stadler, P. F. & Hoffmann, S. Lacking alignments? The next-generation sequencing mapper segemehl revisited. Bioinformatics 30, 1837–1843 (2014).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Sturn, A., Quackenbush, J. & Trajanoski, Z. Genesis: cluster analysis of microarray data. Bioinformatics 18, 207–208 (2002).
Acknowledgements
C.A. is a recipient of a fellowship from the Bayerischen Gleichstellungsförderung (BGF) through the SCIENTIA Program; I.R.L. and R.J.S. are recipients of PhD fellowships (PD/BD/146464/2019 and PD/BD/129294/2017) of the Doctoral Programme in Experimental Biology and Biomedicine of the Center for Neuroscience and Cell Biology, University of Coimbra. The authors acknowledge the animal facility staff at the University of Leon, Spain, in particular A. Carvajal from the Department of Animal Health. This work was supported by grants from the Bavarian Ministry of Sciences, Research and the Arts in the framework of the Bavarian Molecular Biosystems Research Network (BioSysNet), DFG project BR 4837/1-1 and the Portuguese Foundation for Science and Technology (FCT Investigator Programme IF/01105/2015).
Author information
Authors and Affiliations
Contributions
C.A., A.R.C., C.M., M.M. and A.E. designed the experiments; A.R.C., M.M. and A.E. performed the high-throughput screening and validation experiments; C.A., I.R.L., C.M., U.S., R.J.S. and C.L. performed the biochemical and cell biology assays for characterization of the role of miRNAs in infection; M.S. performed the analysis of the RNA-sequencing datasets; S.Z.-L. and J.J.G. performed the in vivo Salmonella infections; M.G. provided reagents and access to equipment; M.M. and A.E. wrote the manuscript, with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Screenings for miRNAs controlling infection by Salmonella and Shigella.
a. Microscopy images and corresponding image segmentation outlines of HeLa cells treated with control miRNA and infected with Salmonella or Shigella. Cells marked in green in the image segmentation panels correspond to cells with high intracellular bacterial load; cells touching the borders of the images were excluded from the analysis. Scale bar, 100 μm. b and c. Pairwise correlation between the three independent runs of the screenings to identify miRNAs that regulate Salmonella (b) and Shigella (c) infection. Results are based on quantification of the percentage of infected cells following treatment with the library of miRNA mimics and are shown as fold change compared to control miRNA (log2 scale). Spearman rank correlation coefficients are shown in the right bottom corner of graphs. d and e. Comparison of the results obtained in the Salmonella (d) and Shigella (e) screenings for the 110 selected miRNAs, with those obtained for the corresponding time point in the time-course experiments (20 hpi, Salmonella; 6 hpi, Shigella). Results shown are the mean of the three independent runs of each screen or time-course experiments. Colours in panels d and e correspond to the different miRNA categories identified in Figure 2a. Spearman rank correlation coefficients are shown in the right bottom corner of graphs.
Extended Data Fig. 2 Shigella intracellular motility and intercellular spreading are inhibited by miR-3668, miR-4732-5p and miR-6073.
a. Fluorescence microscopy images extracted from the time-lapse microscopy analysis of HeLa cells infected with Shigella wild-type, upon treatment with control miRNA, miR-3668, miR-4732-5p or miR-6073 mimics. Full time-lapse sequence is included as Supplementary Videos 1-4. Scale bar, 100 μm. b and c. Representative images (b) and area of Shigella infection foci (c) in HCT-8 cells transfected with control miRNA, miR-3668, miR-4732-5p or miR-6073 mimics and infected with Shigella, analyzed at 3 hpi. Shigella ΔicsA mutant is shown for comparison. Scale bar, 25 µm. d. Area of Listeria infection foci in HeLa cells transfected with the indicated miRNAs, analyzed at 8 hpi. e. Sequences of mature human miR-3668, miR-4732-5p and miR-6073. The 7-mer seed sequences of the miRNAs are highlighted in red. f. N-WASP levels, determined by western blot, in HeLa cells transfected with control miRNA, miR-3668, miR-4732-5p or miR-6073 mimics, or with control siRNA or N-WASP siRNA. g and h. Representative images (g) and percentage of bacteria forming actin tails (h) in HeLa cells transfected with control siRNA or N-WASP siRNA, analyzed at 1.5 hpi. Staining of F-actin using fluorescently labeled phalloidin allows the visualization of actin tails. Scale bar, 10 µm. i and j. Area of Shigella infection foci in HeLa (i) and HCT-8 (j) cells transfected with control miRNA or N-WASP siRNA, analyzed at 3 hpi. Shigella ΔicsA mutant is shown for comparison. k. Area of Listeria infection foci in HeLa cells transfected with control miRNA or N-WASP siRNA, analyzed at 3 hpi. Shigella infection was performed at MOI 25 (b,g,h), 10 (a,i) or 5 (c,j); Listeria infection was performed at MOI 20. Results are shown as mean ± s.e.m. of n=4 (h) or n=5 (c,f,i-k) biologically independent experiments; microscopy images are representative of n=3 (a), n=4 (g) or n=5 (b) biologically independent experiments; centre values indicate means (c,d,h-j); ***P<0.001 (statistical analyses are detailed in Supplementary Table 4).
Extended Data Fig. 3 Targeting of N-WASP by miR-3668, miR-4732-5p and miR-6073 inhibits Shigella intercellular spreading.
a. Area of Shigella infection foci (in parental HeLa cells, HeLa cells stably expressing a siRNA resistant version of N-WASP fused with EGFP (EGFP-N-WASPr) or wild-type N-WASP fused with EGFP (EGFP-N-WASP), transfected with a control siRNA or a siRNA targeting N-WASP, analyzed at 3 hpi. b. N-WASP expression, determined by western blot, in HeLa cells (parental, EGFP-N-WASPr, EGFP-N-WASP) transfected with a control siRNA or a siRNA targeting N-WASP. c. Area of Shigella infection foci in HeLa cells transfected with control siRNA or 4 independent siRNAs targeting N-WASP, analyzed at 3 hpi. d. N-WASP expression, determined by western blot, in HeLa cells transfected with control siRNA or 4 independent siRNAs targeting N-WASP. e. Sequences of the six active binding sites for miR-3668 (2 sites), miR-4732-5p (1 site) and miR-6073 (3 sites) identified in the 3’UTR of N-WASP. Complementarity of the binding sites with the miRNAs, as predicted by TargetScan, is shown. f. N-WASP levels, determined by western blot, in HeLa cells mock-treated or infected with Shigella, at different times post-infection (0.5, 3, 6 and 8 hpi). g. Expression of mature miR-4732-5p or miR-6073, determined by RT-qPCR, in HeLa cells transfected with control miRNA inhibitor or inhibitors of miR-4732-5p and miR-6073, separately or simultaneously. h. N-WASP levels, determined by western blot, in HeLa cells mock-treated or infected with Shigella, and transfected with control miRNA inhibitor or inhibitors of miR-4732-5p and miR-6073 simultaneously, analyzed at 8 hpi. Shigella infection was performed at MOI 100 (f,h) or 10 (a,c). Results are shown as mean ± s.e.m. of n=3 (g,h) or n=5 (a,c,f) biologically independent experiments; western blots (b,d) are representative of n=3 biologically independent experiments; centre values indicate means (a,d); **P<0.01, ***P<0.001 (statistical analyses are detailed in Supplementary Table 4).
Extended Data Fig. 4 Disruption of actin stress fibers reverts the inhibitory effect of let-7i-3p on Salmonella binding to host cells.
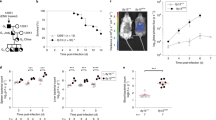
a. Sequences of the mature miRNAs derived from the 5’ and 3’ arm of the precursors of the let-7 family. Common seed sequence of the let-7 miRNA family (-5p miRNAs) is highlighted in blue and that of let-7i-3p is highlighted in green. b, c, e and f. Representative images (b,e) and quantification by cfu (c,f) of Salmonella bound to HeLa cells transfected with control miRNA or let-7i-3p mimics, and incubated with Salmonella wild-type or Δ4 mutant strain for 10 min. Cells were pre-treated with cytochalasin D for 1 h (b,c). Salmonella was previously coated with PL (e,f). d, g and h. Representative images (d,g) and quantification by cfu (h) of Shigella bound to HeLa cells transfected with control miRNA or let-7i-3p mimics, and incubated with Shigella wild-type or ΔipaB mutant strains for 25 min. Shigella was previously coated with PL (g,h). i-l. Representative images (i,k) and quantification by cfu (j,l) of Listeria bound to (i,j) or intracellular Listeria (k,l) of HeLa cells transfected with control miRNA or let-7i-3p mimics. Binding was performed for 25 min, and intracellular bacteria analyzed at 1 and 20 hpi. m-p. Expression of mature let-7i-3p (m,n) and RGS2 (o,p), evaluated by RT-qPCR, in HeLa cells infected with Shigella (m,o) or Listeria (n,p). Shigella and Listeria infections were analyzed at 6 or 20 hpi, respectively. Salmonella infection was performed at MOI 50. Shigella infection was performed at MOI 50 (d,g,h) or MOI 100 (m,o). Listeria infection was performed at MOI 25 (k,l,n,p) or 50 (i,j). Scale bars, 50 μm. Results are shown as mean ± s.e.m. of n=5 (c,f,h,l-p) or n=6 (j) biologically independent experiments; microscopy images are representative of n=3 (i,k) or n=5 (b,d,e,g) biologically independent experiments; ***P<0.001 (statistical analyses are detailed in Supplementary Table 4).
Extended Data Fig. 5 Let-7i-3p targets the coding sequence of RGS2.
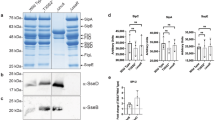
a. Schematic representation of RGS2 3′UTR, CDS, and CDS fragments and mutant constructs used for miRNA binding site reporter assays. The RGS2 3’UTR was cloned downstream of a Renilla luciferase reporter (psiCHECK2-RGS2 3’UTR) and the RGS2 CDS and CDS fragments were cloned as a C-terminal fusion of green fluorescent protein (EGFP-RGS2 CDS). b-d. Results of the reporter assays using the RGS2 3’UTR (b), RGS2 CDS (c), or RGS2 CDS fragments and mutant (d). Results in cells treated with the let-7i-3p mimic are shown compared to the control miRNA. e. Sequence of the binding site for let-7i-3p in fragment F1 of the N-WASP CDS. Complementarity of the miRNA with the binding site of human RGS2 sequence is shown. f. RGS2 expression, determined by western blot, in HeLa cells transfected with control siRNA or RGS2 siRNA. g. Quantification of intracellular Salmonella by cfu in parental HeLa cells, HeLa cells stably expressing a siRNA resistant version of RGS2 fused with EGFP (EGFP-RGS2r) or wild-type RGS2 fused with EGFP (EGFP-RGS2), transfected with a control siRNA or a siRNA targeting RGS2, analyzed at 20 hpi. h. RGS2 expression, determined by western blot, in HeLa cells (parental, EGFP-RGS2r or EGFP-RGS2), transfected with a control siRNA or a siRNA targeting RGS2. i. Quantification of intracellular Salmonella by cfu in HeLa cells transfected with control siRNA or 4 independent siRNAs targeting RGS2, analyzed at 20 hpi. j. RGS2 expression, determined by western blot, in HeLa cells transfected with control siRNA or 4 independent siRNAs targeting RGS2. Salmonella infection was performed at MOI 25. Results are shown as mean ± s.e.m. of n=3 (b), n=5 (g,i), n=7 (d) or n=10 (c) biologically independent experiments; western blots (f,h,j) are representative of n=3 biologically independent experiments; **P<0.01, ***P<0.001 (statistical analyses are detailed in Supplementary Table 4).
Extended Data Fig. 6 Inhibition of let-7i-3p favours Salmonella infection.
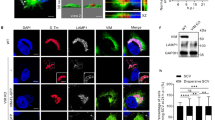
a. Expression of mature let-7i-3p, determined by RT-qPCR, in HeLa cells transfected with control miRNA or let-7i-3p inhibitors. b. RGS2 expression, determined by western blot, in HeLa cells transfected with control miRNA or let-7i-3p inhibitors; non-transfected and mock-transfected cells are shown for comparison. c. Representative images of HeLa cells transfected with control miRNA or let-7i-3p inhibitors and infected with Salmonella. Infection was analyzed at 1, 4 and 20 hpi. Salmonella infection was performed at MOI 25. Scale bar, 50 μm. d-f. Representative images of HeLa cells transfected with control miRNA or let-7i-3p inhibitors and stained with Lysotracker (d), for LAMP-1 (e), or with DQ Red BSA (f). Scale bar, 50 μm. Results are shown as mean ± s.e.m. of n=4 (a) biologically independent experiments; western blots and microscopy images are representative of n=3 (b,d-f) or n=5 (c) biologically independent experiments; ***P<0.001 (statistical analyses are detailed in Supplementary Table 4).
Supplementary information
Supplementary Video 1
Time-lapse microscopy of Shigella infection in HeLa cells transfected with control miRNA. Data are representative of n = 3 biologically independent experiments.
Supplementary Video 2
Time-lapse microscopy of Shigella infection in HeLa cells transfected with miR-3668. Data are representative of n = 3 biologically independent experiments.
Supplementary Video 3
Time-lapse microscopy of Shigella infection in HeLa cells transfected with miR-4732-5p. Data are representative of n = 3 biologically independent experiments.
Supplementary Video 4
Time-lapse microscopy of Shigella infection in HeLa cells transfected with miR-6073. Data are representative of n = 3 biologically independent experiments.
Supplementary Tables
Supplementary Tables 1–4.
Source data
Source Data Fig. 3
Unprocessed Western Blots
Source Data Fig. 4
Unprocessed Western Blots
Source Data Fig. 5
Unprocessed Western Blots
Source Data Fig. 6
Unprocessed Western Blots
Source Data Extended Data Fig. 3
Unprocessed Western Blots
Source Data Extended Data Fig. 5
Unprocessed Western Blots
Source Data Extended Data Fig. 6
Unprocessed Western Blots
Rights and permissions
About this article
Cite this article
Aguilar, C., Cruz, A.R., Rodrigues Lopes, I. et al. Functional screenings reveal different requirements for host microRNAs in Salmonella and Shigella infection. Nat Microbiol 5, 192–205 (2020). https://doi.org/10.1038/s41564-019-0614-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0614-3
This article is cited by
-
Study of microRNA expression in Salmonella Typhimurium-infected porcine ileum reveals miR-194a-5p as an important regulator of the TLR4-mediated inflammatory response
Veterinary Research (2022)
-
Microscopy-based phenotypic profiling of infection by Staphylococcus aureus clinical isolates reveals intracellular lifestyle as a prevalent feature
Nature Communications (2022)
-
Cross-species RNA-seq for deciphering host–microbe interactions
Nature Reviews Genetics (2021)
-
Reprogramming of microRNA expression via E2F1 downregulation promotes Salmonella infection both in infected and bystander cells
Nature Communications (2021)