Abstract
CRISPR–Cas systems provide bacteria with adaptive immunity against bacteriophages1. However, DNA modification2,3, the production of anti-CRISPR proteins4,5 and potentially other strategies enable phages to evade CRISPR–Cas. Here, we discovered a Serratia jumbo phage that evades type I CRISPR–Cas systems, but is sensitive to type III immunity. Jumbo phage infection resulted in a nucleus-like structure enclosed by a proteinaceous phage shell—a phenomenon only reported recently for distantly related Pseudomonas phages6,7. All three native CRISPR–Cas complexes in Serratia—type I-E, I-F and III-A—were spatially excluded from the phage nucleus and phage DNA was not targeted. However, the type III-A system still arrested jumbo phage infection by targeting phage RNA in the cytoplasm in a process requiring Cas7, Cas10 and an accessory nuclease. Type III, but not type I, systems frequently targeted nucleus-forming jumbo phages that were identified in global viral sequence datasets. The ability to recognize jumbo phage RNA and elicit immunity probably contributes to the presence of both RNA- and DNA-targeting CRISPR–Cas systems in many bacteria1,8. Together, our results support the model that jumbo phage nucleus-like compartments serve as a barrier to DNA-targeting, but not RNA-targeting, defences, and that this phenomenon is widespread among jumbo phages.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author on request. The genome sequence of bacteriophage PCH45 is available in GenBank under accession number MN334766. Data and R scripts for the bioinformatics analyses are available at https://github.com/JacksonLab/Jumbophages.
References
Hille, F. et al. The biology of CRISPR-Cas: backward and forward. Cell 172, 1239–1259 (2018).
Bryson, A. L. et al. Covalent modification of bacteriophage T4 DNA inhibits CRISPR-Cas9. mBio 6, e00648 (2015).
Vlot, M. et al. Bacteriophage DNA glucosylation impairs target DNA binding by type I and II but not by type V CRISPR–Cas effector complexes. Nucleic Acids Res. 46, 873–885 (2018).
Pawluk, A., Davidson, A. R. & Maxwell, K. L. Anti-CRISPR: discovery, mechanism and function. Nat. Rev. Microbiol. 16, 12–17 (2018).
Bondy-Denomy, J., Pawluk, A., Maxwell, K. L. & Davidson, A. R. Bacteriophage genes that inactivate the CRISPR/Cas bacterial immune system. Nature 493, 429–432 (2013).
Chaikeeratisak, V. et al. Assembly of a nucleus-like structure during viral replication in bacteria. Science 355, 194–197 (2017).
Chaikeeratisak, V. et al. The phage nucleus and tubulin spindle are conserved among large Pseudomonas phages. Cell Rep. 20, 1563–1571 (2017).
Makarova, K. S., Wolf, Y. I. & Koonin, E. V. Classification and nomenclature of CRISPR-Cas systems: where from here? CRISPR J. 1, 325–336 (2018).
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007).
Jackson, S. A. et al. CRISPR-Cas: adapting to change. Science 356, eaal5056 (2017).
Brouns, S. J. J. et al. Small CRISPR RNAs guide antiviral defense in prokaryotes. Science 321, 960–964 (2008).
Hale, C. R. et al. RNA-guided RNA cleavage by a CRISPR RNA-Cas protein complex. Cell 139, 945–956 (2009).
Garneau, J. E. et al. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468, 67–71 (2010).
Patterson, A. G. et al. Quorum sensing controls adaptive immunity through the regulation of multiple CRISPR-Cas systems. Mol. Cell 64, 1102–1108 (2016).
Yuan, Y. & Gao, M. Jumbo bacteriophages: an overview. Front. Microbiol. 8, 403 (2017).
Mesyanzhinov, V. V. et al. The genome of bacteriophage φKZ of Pseudomonas aeruginosa. J. Mol. Biol. 317, 1–19 (2002).
Ceyssens, P. J. et al. Development of giant bacteriophage φKZ is independent of the host transcription apparatus. J. Virol. 88, 10501–10510 (2014).
Jackson, S. A., Birkholz, N., Malone, L. M. & Fineran, P. C. Imprecise spacer acquisition generates CRISPR-Cas immune diversity through primed adaptation. Cell Host Microbe 25, 250–260 (2019).
Kraemer, J. A. et al. A phage tubulin assembles dynamic filaments by an atypical mechanism to center viral DNA within the host cell. Cell 149, 1488–1499 (2012).
Erb, M. L. et al. A bacteriophage tubulin harnesses dynamic instability to center DNA in infected cells. eLife 3, e03197 (2014).
Samai, P. et al. Co-transcriptional DNA and RNA cleavage during type III CRISPR-Cas immunity. Cell 161, 1164–1174 (2015).
Goldberg, G. W., Jiang, W., Bikard, D. & Marraffini, L. A. Conditional tolerance of temperate phages via transcription-dependent CRISPR–Cas targeting. Nature 514, 633–637 (2014).
Jia, N. et al. Type III-A CRISPR-Cas Csm complexes: assembly, periodic RNA cleavage, DNase activity regulation, and autoimmunity. Mol. Cell 73, 264–277 (2019).
Liu, T. Y., Iavarone, A. T. & Doudna, J. A. RNA and DNA targeting by a reconstituted Thermus thermophilus type III-A CRISPR-Cas system. PLoS ONE 12, e0170552 (2017).
Niewoehner, O. et al. Type III CRISPR–Cas systems produce cyclic oligoadenylate second messengers. Nature 548, 543–548 (2017).
Kazlauskiene, M., Kostiuk, G., Venclovas, Č., Tamulaitis, G. & Siksnys, V. A cyclic oligonucleotide signaling pathway in type III CRISPR-Cas systems. Science 357, 605–609 (2017).
Rouillon, C., Athukoralage, J. S., Graham, S., Gruschow, S. & White, M. F. Control of cyclic oligoadenylate synthesis in a type III CRISPR system. eLife 7, e36734 (2018).
Varble, A. & Marraffini, L. A. Three new Cs for CRISPR: collateral, communicate, cooperate. Trends Genet. 35, 446–456 (2019).
Lau, R. K. et al. Structure and mechanism of a cyclic trinucleotide-activated bacterial endonuclease mediating bacteriophage immunity. Preprint at https://doi.org/10.1101/694703 (2019).
Silas, S. et al. Direct CRISPR spacer acquisition from RNA by a natural reverse transcriptase-Cas1 fusion protein. Science 351, aad4234 (2016).
Hynes, A. P., Villion, M. & Moineau, S. Adaptation in bacterial CRISPR-Cas immunity can be driven by defective phages. Nat. Commun. 5, 4399 (2014).
Mendoza, S. D. et al. A nucleus-like compartment shields bacteriophage DNA from CRISPR-Cas and restriction nucleases. Preprint at https://doi.org/10.1101/370791 (2018).
Watson, B. N. J., Staals, R. H. J. & Fineran, P. C. CRISPR-Cas-mediated phage resistance enhances horizontal gene transfer by transduction. mBio 9, e02406-17 (2018).
Frampton, R. A. et al. Identification of bacteriophages for biocontrol of the kiwifruit canker phytopathogen Pseudomonas syringae pv. actinidiae. Appl. Environ. Microbiol. 80, 2216–2228 (2014).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Brettin, T. et al. RASTtk: a modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes. Sci. Rep. 5, 8365 (2015).
Lowe, T. M. & Chan, P. P. tRNAscan-SE On-line: integrating search and context for analysis of transfer RNA genes. Nucleic Acids Res. 44, W54–W57 (2016).
Carver, T., Thomson, N., Bleasby, A., Berriman, M. & Parkhill, J. DNAPlotter: circular and linear interactive genome visualization. Bioinformatics 25, 119–120 (2009).
Bao, Y., Chetvernin, V. & Tatusova, T. Improvements to pairwise sequence comparison (PASC): a genome-based web tool for virus classification. Arch. Virol. 159, 3293–3304 (2014).
Sullivan, M. J., Petty, N. K. & Beatson, S. A. Easyfig: a genome comparison visualizer. Bioinformatics 27, 1009–1010 (2011).
Meier-Kolthoff, J. P. & Göker, M. VICTOR: genome-based phylogeny and classification of prokaryotic viruses. Bioinformatics 33, 3396–3404 (2017).
Pawluk, A. et al. Inactivation of CRISPR-Cas systems by anti-CRISPR proteins in diverse bacterial species. Nat. Microbiol. 1, 16085 (2016).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Hampton, H. G. et al. CRISPR-Cas gene-editing reveals RsmA and RsmC act through FlhDC to repress the SdhE flavinylation factor and control motility and prodigiosin production in Serratia. Microbiology 162, 1047–1058 (2016).
Kaniga, K., Delor, I. & Cornelis, G. R. A wide-host-range suicide vector for improving reverse genetics in Gram-negative bacteria: inactivation of the blaA gene of Yersinia enterocolitica. Gene 109, 137–141 (1991).
Watson, B. N. J. et al. Type I-F CRISPR-Cas resistance against virulent phage infection triggers abortive infection and provides population-level immunity. Preprint at https://doi.org/10.1101/679308 (2019).
Silva-Rocha, R. et al. The Standard European Vector Architecture (SEVA): a coherent platform for the analysis and deployment of complex prokaryotic phenotypes. Nucleic Acids Res. 41, D666–D675 (2013).
Biswas, A., Staals, R. H., Morales, S. E., Fineran, P. C. & Brown, C. M. CRISPRDetect: a flexible algorithm to define CRISPR arrays. BMC Genomics 17, 356 (2016).
Nicholson, T. J. et al. Bioinformatic evidence of widespread priming in type I and II CRISPR-Cas systems. RNA Biol. 16, 566–576 (2019).
Paez-Espino, D. et al. IMG/VR v.2.0: an integrated data management and analysis system for cultivated and environmental viral genomes. Nucleic Acids Res. 47, D678–D686 (2019).
Rizk, G. & Lavenier, D. GASSST: global alignment short sequence search tool. Bioinformatics 26, 2534–2540 (2010).
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Johnson, L. S., Eddy, S. R. & Portugaly, E. Hidden Markov model speed heuristic and iterative HMM search procedure. BMC Bioinformatics 11, 431 (2010).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Finn, R. D., Clements, J. & Eddy, S. R. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 39, W29–W37 (2011).
Acknowledgements
This work was supported by the Marsden Fund from the Royal Society of New Zealand and a University of Otago Research Grant. L.M.M. was supported by a University of Otago Doctoral Scholarship. We thank staff of the Otago Micro and Nano Imaging facility for assistance with electron and confocal microscopy and the Otago Genomics Facility for genome sequencing. We thank members of the Fineran laboratory for helpful discussions, S. Shehreen, T. Nicholson and X. Morgan for bioinformatics advice. We also acknowledge the use of the New Zealand eScience Infrastructure (NeSI) high-performance computing facilities in this research. NeSI’s facilities are provided by and funded jointly by NeSI’s collaborator institutions and through the Ministry of Business, Innovation and Employment’s Research Infrastructure programme.
Author information
Authors and Affiliations
Contributions
L.M.M., S.L.W., C.W. and L.F.G. performed the experiments. L.M.M., S.L.W., S.A.J. and C.W. generated the strains and plasmids. L.M.M., S.L.W. and L.F.G. performed the microscopy. L.M.M. and S.A.J. performed the bioinformatics analysis with input from P.P.G. and P.C.F. L.M.M. and P.C.F. conceived the project with input from all authors. P.C.F. supervised the project. L.M.M. and P.C.F. wrote the manuscript. All authors edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
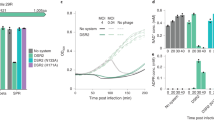
Extended Data Fig. 1 The jumbo phage is resistant to type I CRISPR-Cas immunity.
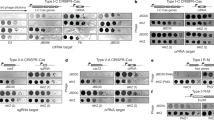
a. tblastx alignment of PCH45 with phages PhiEaH1 and 2050HW. b. Target location of chromosomally expressed anti-PCH45 type I-E (S4-7) and type I-F (S8) spacers targeting the major capsid gene (gp033). Phage resistance measured by c. EOP or d. plate reader assays for Serratia strains with type I-E, (S4, PCF591; S5, PCF593; S6, PCF545; S7, PCF544) and type I-F (S8, PCF548) infected with PCH45. Phage resistance measured by e. EOP or f. plate reader assays for Serratia carrying a type I-E (S4, pPF1460) spacer in mini-CRISPR arrays, infected with PCH45. Phage resistance measured by g. EOP or h. plate reader assays for Serratia carrying a type I-F (S8, pPF1461) spacer in a plasmid mini-CRISPR array, infected with PCH45. In c, e and g MOI=0.001. In c-h data are presented as mean ± s.d. (n=3 biologically independent samples). i. Restriction length fragment polymorphism (RLFP) analysis of phage gDNA treated with restriction enzymes MfeI, EcoRI, DpnI*, KpnI, AluI and NruI. Undigested PCH45 gDNA was used as a negative control. In parenthesis the number of restriction sites found in the genome of PCH45. (*): Cleaves only when the recognition motif is methylated. This experiment was performed three times with similar results and a representative gel is shown.
Extended Data Fig. 2 The shell and tubulin proteins in Serratia jumbo phage PCH45 possess low sequence similarity to homologues encoded by other jumbo phages.
Phylogenetic tree of a. the shell protein and b. PhuZ protein encoded by jumbo phages. The maximum likelihood trees were built for phage encoded shell and tubulin-like proteins (n=9 and n=11, respectively) using RaxML with 100 bootstrap replicates. The scale bar represents the approximate number of changes per amino acid position. c. EOP assay for Serratia strains PCF761 (mCherry2-cas8e), PCF763 (mCherry2-cas8f) and PCF765 (mCherry2-cas10) carrying type I-E, I-F and III-A anti-JS26 spacers in plasmids (pPF1485, pPF1489 and pPF1473 respectively). In c data are presented as mean ± s.d. (n=3 biologically independent samples).
Extended Data Fig. 3 Type III RNA targeting provides protection against jumbo phage infection.
a. EOP assay for type III-A mutant strains: cas10H17A, N18A (HD domain), cas10D618A, D619A (Palm domain), cas7D34A, and the accessory nuclease knock out carrying an anti-PCH45 spacers (RNA polymerase beta subunit, S9; anti-terminase S10; and anti-helicase, S12) overexpressed in trans from a plasmid mini-CRISPR array. b. Conjugation efficiency assay (transconjugants/recipients) of plasmids pPF781 (untargeted control) and pPF1043 targeted by the type III-A CRISPR-Cas systems for Serratia strains. The type III-A mutants: PCF683 (cas7D34A), PCF690 (cas10 HD mutant), PCF691 (cas10 Palm mutant) PCF686 (Δ accessory nuclease), and c. the chromosomal complementation with wild-type copies of the genes in PCF684 (cas7), PCF688 (cas10) and PCF687 (accessory nuclease). All data are presented as mean ± s.d. (n=3 biologically independent samples).
Extended Data Fig. 4 Type III CRISPR arrays are enriched in jumbo phage-targeting spacers.
a. Workflow used to obtain spacer-phage hits. Scores for spacer-target matches for targeted (black) and shuffled (grey) databases for b. type III c. type I-E and d. type I-F CRISPR-Cas systems. Scores with a false positive rate (FPR) < 0.01 were used as a cut-off to determine the spacer-protospacer hits; the FPR is defined as (the number of hits above the scoring threshold to the shuffled target database)/(the number of hits above the scoring threshold to the target database). e. Number of unique spacers in type I-E, I-F or type III systems matching nucleus-forming phages.
Extended Data Fig. 5
Spacer list in native CRISPR arrays.
Extended Data Fig. 6
Spacers expressed from mini-array in plasmid.
Supplementary information
Supplementary Information
Supplementary Tables 3–5.
Supplementary Data
Supplementary File 1.
Supplementary Data
Supplementary File 2.
Supplementary Tables
Supplementary Tables 1 and 2.
Rights and permissions
About this article
Cite this article
Malone, L.M., Warring, S.L., Jackson, S.A. et al. A jumbo phage that forms a nucleus-like structure evades CRISPR–Cas DNA targeting but is vulnerable to type III RNA-based immunity. Nat Microbiol 5, 48–55 (2020). https://doi.org/10.1038/s41564-019-0612-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0612-5
This article is cited by
-
Phage proteins target and co-opt host ribosomes immediately upon infection
Nature Microbiology (2024)
-
Inhibitors of bacterial immune systems: discovery, mechanisms and applications
Nature Reviews Genetics (2024)
-
Conservation and similarity of bacterial and eukaryotic innate immunity
Nature Reviews Microbiology (2024)
-
Nucleus-forming vibriophage cocktail reduces shrimp mortality in the presence of pathogenic bacteria
Scientific Reports (2023)
-
Identification of the bacteriophage nucleus protein interaction network
Nature Structural & Molecular Biology (2023)