Abstract
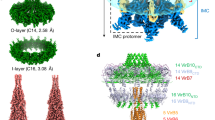
The type II secretion system (T2SS) is a multiprotein envelope-spanning assembly that translocates a wide range of virulence factors, enzymes and effectors through the outer membrane of many Gram-negative bacteria1,2,3. Here, using electron cryotomography and subtomogram averaging methods, we reveal the in vivo structure of an intact T2SS imaged within the human pathogen Legionella pneumophila. Although the T2SS has only limited sequence and component homology with the evolutionarily related type IV pilus (T4P) system4,5, we show that their overall architectures are remarkably similar. Despite similarities, there are also differences, including, for example, that the T2SS–ATPase complex is usually present but disengaged from the inner membrane, the T2SS has a much longer periplasmic vestibule and it has a short-lived flexible pseudopilus. Placing atomic models of the components into our electron cryotomography map produced a complete architectural model of the intact T2SS that provides insights into the structure and function of its components, its position within the cell envelope and the interactions between its different subcomplexes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The subtomogram averages of the L. pneumophila T2SS have been deposited in the Electron Microscopy Data Bank under the following accession codes: EMD-20713 (wild type, aligned on the OM part) and EMD-20712 (wild type, aligned on the IM part). All additional data/information are available from the authors upon request. The authors declare that all data supporting the findings of this study, including source data for Fig. 1, are available within the paper and its Supplementary Information.
Change history
19 February 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Costa, T. R. D. et al. Secretion systems in Gram-negative bacteria: structural and mechanistic insights. Nat. Rev. Microbiol. 13, 343–359 (2015).
Abby, S. S. et al. Identification of protein secretion systems in bacterial genomes. Sci. Rep. 6, 23080 (2016).
White, R. C. & Cianciotto, N. P. Assessing the impact, genomics and evolution of type II secretion across a large, medically important genus: the Legionella type II secretion paradigm. Microb. Genom. 5, e000273 (2019).
Hospenthal, M. K., Costa, T. R. D. & Waksman, G. A comprehensive guide to pilus biogenesis in Gram-negative bacteria. Nat. Rev. Microbiol. 15, 365–379 (2017).
Peabody, C. R. et al. Type II protein secretion and its relationship to bacterial type IV pili and archaeal flagella. Microbiology 149, 3051–3072 (2003).
Cianciotto, N. P. Type II secretion: a protein secretion system for all seasons. Trends Microbiol. 13, 581–588 (2005).
Cianciotto, N. P. & White, R. C. Expanding role of type II secretion in bacterial pathogenesis and beyond. Infect. Immun. 85, e00014–e00017 (2017).
Thomassin, J.-L., Santos Moreno, J., Guilvout, I., Tran Van Nhieu, G. & Francetic, O. The trans-envelope architecture and function of the type 2 secretion system: new insights raising new questions. Mol. Microbiol. 105, 211–226 (2017).
Filloux, A. & Voulhoux, R. Multiple structures disclose the secretins’ secrets. J. Bacteriol. 200, e00702-17 (2018).
Nivaskumar, M. & Francetic, O. Type II secretion system: a magic beanstalk or a protein escalator. Biochim. Biophys. Acta 1843, 1568–1577 (2014).
Nunn, D. Bacterial type II protein export and pilus biogenesis: more than just homologies? Trends Cell Biol. 9, 402–408 (1999).
López-Castilla, A. et al. Structure of the calcium-dependent type 2 secretion pseudopilus. Nat. Microbiol. 2, 1686–1695 (2017).
Yan, Z., Yin, M., Xu, D., Zhu, Y. & Li, X. Structural insights into the secretin translocation channel in the type II secretion system. Nat. Struct. Mol. Biol. 24, 177–183 (2017).
Hay, I. D., Belousoff, M. J., Dunstan, R. A., Bamert, R. S. & Lithgow, T. Structure and membrane topography of the Vibrio-type secretin complex from the type 2 secretion system of enteropathogenic Escherichia coli. J. Bacteriol. 200, e00521-17 (2017).
Yin, M., Yan, Z. & Li, X. Structural insight into the assembly of the type II secretion system pilotin–secretin complex from enterotoxigenic Escherichia coli. Nat. Microbiol. 3, 581–587 (2018).
Howard, S. P. et al. Structure and assembly of pilotin-dependent and -independent secretins of the type II secretion system. PLoS Pathog. 15, e1007731 (2019).
Chernyatina, A. A. & Low, H. H. Architecture of a bacterial type II secretion system. Preprint at https://www.biorxiv.org/content/10.1101/397794v1 (2018).
Hay, I. D., Belousoff, M. J. & Lithgow, T. Structural basis of type 2 secretion system engagement between the inner and outer bacterial membranes. mBio 8, e01344-17 (2017).
Yamagata, A. & Tainer, J. A. Hexameric structures of the archaeal secretion ATPase GspE and implications for a universal secretion mechanism. EMBO J. 26, 878–890 (2007).
Lu, C. et al. Hexamers of the type II secretion ATPase GspE from Vibrio cholerae with increased ATPase activity. Structure 21, 1707–1717 (2013).
Korotkov, K. V. et al. Structural and functional studies on the interaction of GspC and GspD in the type II secretion system. PLoS Pathog. 7, e1002228 (2011).
Korotkov, K. V. & Hol, W. G. J. Structure of the GspK–GspI–GspJ complex from the enterotoxigenic Escherichia coli type 2 secretion system. Nat. Struct. Mol. Biol. 15, 462–468 (2008).
Zhang, Y. et al. Structure-guided disruption of the pseudopilus tip complex inhibits the type II secretion in Pseudomonas aeruginosa. PLoS Pathog. 14, e1007343 (2018).
Abendroth, J., Rice, A. E., McLuskey, K., Bagdasarian, M. & Hol, W. G. J. The crystal structure of the periplasmic domain of the type II secretion system protein EpsM from Vibrio cholerae: the simplest version of the ferredoxin fold. J. Mol. Biol. 338, 585–596 (2004).
Abendroth, J., Kreger, A. C. & Hol, W. G. J. The dimer formed by the periplasmic domain of EpsL from the type 2 secretion system of Vibrio parahaemolyticus. J. Struct. Biol. 168, 313–322 (2009).
Abendroth, J. et al. The three-dimensional structure of the cytoplasmic domains of EpsF from the type 2 secretion system of Vibrio cholerae. J. Struct. Biol. 166, 303–315 (2009).
Ghosal, D. et al. Molecular architecture, polar targeting and biogenesis of the Legionella Dot/Icm T4SS. Nat. Microbiol. 4, 1173–1182 (2019).
Kaplan, M. et al. The presence and absence of periplasmic rings in bacterial flagellar motors correlates with stator type. eLife 8, e43487 (2019).
Ghosal, D., Kaplan, M., Chang, Y.-W. & Jensen, G. J. In situ imaging and structure determination of bacterial toxin delivery systems using electron cryotomography. Methods Mol. Biol. 1921, 249–265 (2019).
Oikonomou, C. M., Chang, Y.-W. & Jensen, G. J. A new view into prokaryotic cell biology from electron cryotomography. Nat. Rev. Micro. 14, 205–220 (2016).
Hu, B. et al. Visualization of the type III secretion sorting platform of Shigella flexneri. Proc. Natl Acad. Sci. USA 112, 1047–1052 (2015).
Liles, M. R., Viswanathan, V. K. & Cianciotto, N. P. Identification and temperature regulation of Legionella pneumophila genes involved in type IV pilus biogenesis and type II protein secretion. Infect. Immun. 66, 1776–1782 (1998).
Chang, Y.-W. et al. Architecture of the Vibrio cholerae toxin-coregulated pilus machine revealed by electron cryotomography. Nat. Microbiol. 2, 16269 (2017).
Chang, Y.-W. et al. Architecture of the type IVa pilus machine. Science 351, aad2001 (2016).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. E. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Adebali, O., Ortega, D. R. & Zhulin, I. B. CDvist: a webserver for identification and visualization of conserved domains in protein sequences. Bioinformatics 31, 1475–1477 (2015).
D’Imprima, E. et al. Cryo-EM structure of the bifunctional secretin complex of Thermus thermophilus. eLife 6, e30483 (2017).
Cisneros, D. A., Bond, P. J., Pugsley, A. P., Campos, M. & Francetic, O. Minor pseudopilin self-assembly primes type II secretion pseudopilus elongation: role of minor pseudopilins in type II secretion. EMBO J. 31, 1041–1053 (2012).
Gu, S., Shevchik, V. E., Shaw, R., Pickersgill, R. W. & Garnett, J. A. The role of intrinsic disorder and dynamics in the assembly and function of the type II secretion system. Biochim. Biophys. Acta Proteins Proteom. 1865, 1255–1266 (2017).
Strozen, T. G. et al. Involvement of the GspAB complex in assembly of the type II secretion system secretin of Aeromonas and Vibrio species. J. Bacteriol. 193, 2322–2331 (2011).
Michel-Souzy, S. et al. Direct interactions between the secreted effector and the T2SS components GspL and GspM reveal a new effector-sensing step during type 2 secretion. J. Biol. Chem. 293, 19441–19450 (2018).
Abendroth, J., Murphy, P., Sandkvist, M., Bagdasarian, M. & Hol, W. G. J. The X-ray structure of the type II secretion system complex formed by the N-terminal domain of EpsE and the cytoplasmic domain of EpsL of Vibrio cholerae. J. Mol. Biol. 348, 845–855 (2005).
Py, B., Loiseau, L. & Barras, F. An inner membrane platform in the type II secretion machinery of Gram-negative bacteria. EMBO Rep. 2, 244–248 (2001).
Arts, J. et al. Interaction domains in the Pseudomonas aeruginosa type II secretory apparatus component XcpS (GspF). Microbiology 153, 1582–1592 (2007).
Waters, R. C., O’Toole, P. W. & Ryan, K. A. The FliK protein and flagellar hook-length control. Protein Sci. 16, 769–780 (2007).
Jeong, K. C., Ghosal, D., Chang, Y.-W., Jensen, G. J. & Vogel, J. P. Polar delivery of Legionella type IV secretion system substrates is essential for virulence. Proc. Natl Acad. Sci. USA 114, 8077–8082 (2017).
Ghosal, D., Chang, Y.-W., Jeong, K. C., Vogel, J. P. & Jensen, G. J. In situ structure of the Legionella Dot/Icm type IV secretion system by electron cryotomography. EMBO Rep. 18, 726–732 (2017).
Vincent, C. D. et al. Identification of the core transmembrane complex of the Legionella Dot/Icm type IV secretion system. Mol. Microbiol. 62, 1278–1291 (2006).
White, R. C. et al. Type II secretion-dependent aminopeptidase LapA and acyltransferase PlaC are redundant for nutrient acquisition during Legionella pneumophila intracellular infection of amoebas. mBio 9, e00528-18 (2018).
Rossier, O., Starkenburg, S. R. & Cianciotto, N. P. Legionella pneumophila type II protein secretion promotes virulence in the A/J mouse model of Legionnaires’ disease pneumonia. Infect. Immun. 72, 310–321 (2004).
Rossier, O. & Cianciotto, N. P. Type II protein secretion is a subset of the PilD-dependent processes that facilitate intracellular infection by Legionella pneumophila. Infect. Immun. 69, 2092–2098 (2001).
Truchan, H. K., Christman, H. D., White, R. C., Rutledge, N. S. & Cianciotto, N. P. Type II secretion substrates of Legionella pneumophila translocate out of the pathogen-occupied vacuole via a semipermeable membrane. mBio 8, e00870-17 (2017).
Zheng, S. Q. et al. UCSF tomography: an integrated software suite for real-time electron microscopic tomographic data collection, alignment and reconstruction. J. Struct. Biol. 157, 138–147 (2007).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Agulleiro, J.-I. & Fernandez, J.-J. Tomo3D 2.0-exploitation of advanced vector extensions (AVX) for 3D reconstruction. J. Struct. Biol. 189, 147–152 (2015).
Nicastro, D. et al. The molecular architecture of axonemes revealed by cryoelectron tomography. Science 313, 944–948 (2006).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Korotkov, K. V., Sandkvist, M. & Hol, W. G. J. The type II secretion system: biogenesis, molecular architecture and mechanism. Nat. Rev. Microbiol. 10, 336–351 (2012).
Leighton, T. L., Yong, D. H., Howell, P. L. & Burrows, L. L. Type IV pilus alignment subcomplex proteins PilN and PilO form homo- and heterodimers in vivo. J. Biol. Chem. 291, 19923–19938 (2016).
Schneidman-Duhovny, D., Inbar, Y., Nussinov, R. & Wolfson, H. J. PatchDock and SymmDock: servers for rigid and symmetric docking. Nucleic Acids Res. 33, W363–W367 (2005).
Douzi, B. et al. The XcpV/GspI pseudopilin has a central role in the assembly of a quaternary complex within the T2SS pseudopilus. J. Biol. Chem. 284, 34580–34589 (2009).
Acknowledgements
This work was supported by National Institutes of Health grants AI127401 to G.J.J. and AI043987 to N.P.C. ECT data were recorded at the Beckman Institute Resource Center for Transmission Electron Microcopy at Caltech and the cryo-EM facility at Janelia Research Campus. We thank D. Ortega, Y.-W. Chang and R.C. White for helpful discussions. M.K. is supported by a postdoctoral Rubicon fellowship from De Nederlandse Organisatie voor Wetenschappelijk Onderzoek.
Author information
Authors and Affiliations
Contributions
D.G. and G.J.J. conceived the project. D.G. prepared samples and recorded and processed tomography data. D.G., K.W.K. and G.J.J. analysed data. H.Z., H.K.T., A.E.L., I.E.M., N.P.C. and J.P.V. generated and characterized L. pneumophila mutants. M.K. helped in model building. D.G., K.W.K., N.P.C. and G.J.J. wrote the manuscript with input from all the authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1
Comparison between the T2SS and related molecular machines T4aP and T4bP systems.
Extended Data Fig. 2 T2SS flexibility.
Subtomogram averages of all particles aligned on (a) the OM-associated complex and (b) the IM-associated complex. The distribution of the green dots in (b) indicates the translations imposed on the OM complexes to align the IM complexes. (c) A composite average using the upper and lower halves of (a) and (b), respectively. d) Local resolution of (c) calculated by Resmap. (e) Focused alignment near the base of the secretin channel revealed the presence of a plug-like structure. 20% of the particles with highest cross-correlation showed this distinct density. In the rest of the particles, the plug density is either not present or so dynamic that including them makes the plug almost invisible. (f) Previously reported in situ averages of the T4aP (WT, piliated) and T4bP (WT, piliated and ∆tcpR mutant) machines in states with cytoplasmic dome, ring and disks for comparison33,34. White arrows indicate cytoplasmic disks. Scale bars, 10 nm (a–c), (e), (f). Panel reproduced from: f, ref. 33, Springer Nature Ltd; ref. 34, AAAS.
Extended Data Fig. 3 Position of the T2SS secretin with respect to the OM.
(a, b) Atomic models of the V. cholerae (PDB ID: 5WQ8) and E. coli (PDB ID: 5WQ7) T2SS secretins superimposed on our subtomogram average based on the position of the gate. (c) Positions of the OM on these structures as suggested in earlier publications13,15. The widths of the suggested OM spanning regions were only ~ 1.8 nm, but real membranes are known to be 5–7 nm wide. In all reported atomic models, the secretin channel is suggested to extend beyond the OM13,9. However, when we overlaid the secretin atomic models on our subtomogram average, it only reached through the inner leaflet of the OM. Scale bars, 10 nm. (d) Tomographic slices of mutant L. pneumophila cells lacking all major and minor pilins (ΔlspGHIJK). Showing representative individual T2SS particles. No pseudopilus or lower-periplasmic ring is visible. A similar result was obtained when we examined a L. pneumophila ΔlspHIJK mutant. Scale bar, 10 nm (d). For each strain, number of tomograms recorded and number of particles found are listed in the SI Table-1.
Extended Data Fig. 4 Position of the T2SS with respect to the PG layer in L. pneumophila.
(a, b) Tomographic slices through L. pneumophila cells showing T2SS particles (red arrowheads) and the peptidoglycan layer (PG, yellow arrowheads). (c, d) Tomographic slices through L. pneumophila cells showing T4BSS particles (red arrowheads) and the peptidoglycan layer (PG, yellow arrowheads). (e) Tomographic slice through a L. pneumophila cell showing both T4SS and T2SS particles (red arrowheads) and the peptidoglycan (PG, yellow arrowheads) in the same cell. (f, g) Subtomogram averages of the T4BSS and T2SS, respectively. DotK (shown as green arrow) in the T4BSS is known to interact with the PG layer confirming its location just a few nm below the OM (f). We therefore conclude that the PG layer surrounds the T2SS at approximately the level of the gate (g). For each strain, number of tomograms recorded and number of particles found are listed in the SI Table-1. Scale bars, 100 nm (a–e), 10 nm (f, g). Panel f reproduced from: b, ref. 27, Springer Nature Ltd.
Extended Data Fig. 5
Cryo-EM data collection, refinement and validation statistics.
Supplementary information
Supplementary Information
Supplementary Table 1.
Source data
Source Data Fig. 2
Unprocessed Western Blots
Rights and permissions
About this article
Cite this article
Ghosal, D., Kim, K.W., Zheng, H. et al. In vivo structure of the Legionella type II secretion system by electron cryotomography. Nat Microbiol 4, 2101–2108 (2019). https://doi.org/10.1038/s41564-019-0603-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0603-6
This article is cited by
-
Bdellovibrio predation cycle characterized at nanometre-scale resolution with cryo-electron tomography
Nature Microbiology (2023)
-
Membrane translocation process revealed by in situ structures of type II secretion system secretins
Nature Communications (2023)
-
CryoEM structure of the type IVa pilus secretin required for natural competence in Vibrio cholerae
Nature Communications (2020)