Abstract
To revitalize the antibiotic pipeline, it is critical to identify and validate new antimicrobial targets1. In Mycobacteria tuberculosis and Francisella tularensis, biotin biosynthesis is a key fitness determinant during infection2,3,4,5, making it a high-priority target. However, biotin biosynthesis has been overlooked for priority pathogens such as Acinetobacter baumannii, Klebsiella pneumoniae and Pseudomonas aeruginosa. This can be attributed to the lack of attenuation observed for biotin biosynthesis genes during transposon mutagenesis studies in mouse infection models6,7,8,9. Previous studies did not consider the 40-fold higher concentration of biotin in mouse plasma compared to human plasma. Here, we leveraged the unique affinity of streptavidin to develop a mouse infection model with human levels of biotin. Our model suggests that biotin biosynthesis is essential during infection with A. baumannii, K. pneumoniae and P. aeruginosa. Encouragingly, we establish the capacity of our model to uncover in vivo activity for the biotin biosynthesis inhibitor MAC13772. Our model addresses the disconnect in biotin levels between humans and mice, and explains the failure of potent biotin biosynthesis inhibitors in standard mouse infection models.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The crystal structure of E. coli BioA in complex with MAC13772 solved to 2.4 Å has been deposited to the Protein Data Bank with accession code 6ED7. The source data underlying Figs. 1d and 4a–c, are provided as Supplementary Data Tables 2–9. The data supporting the findings of this study are available from the corresponding author on reasonable request.
References
Brown, E. D. & Wright, G. D. Antibacterial drug discovery in the resistance era. Nature 529, 336–343 (2016).
Sassetti, C. M. & Rubin, E. J. Genetic requirements for mycobacterial survival during infection. Proc. Natl Acad. Sci. USA 100, 12989–12994 (2003).
Weiss, D. S. et al. In vivo negative selection screen identifies genes required for Francisella virulence. Proc. Natl Acad. Sci. USA 104, 6037–6042 (2007).
Napier, B. A. et al. Link between intraphagosomal biotin and rapid phagosomal escape in Francisella. Proc. Natl Acad. Sci. USA 109, 18084–18089 (2012).
Park, S. W. et al. Evaluating the sensitivity of Mycobacterium tuberculosis to biotin deprivation using regulated gene expression. PLoS Pathog. 7, e1002264 (2011).
Wang, N., Ozer, E. A., Mandel, M. J. & Hauser, A. R. Genome-wide identification of Acinetobacter baumannii genes necessary for persistence in the lung. mBio 5, e01163–14 (2014).
Subashchandrabose, S. et al. Acinetobacter baumannii genes required for bacterial survival during bloodstream infection. mSphere 1, e00013–e00015 (2016).
Bachman, M. A. et al. Genome-wide identification of Klebsiella pneumoniae fitness genes during lung infection. mBio 6, e00775–15 (2015).
Skurnik, D. et al. A comprehensive analysis of in vitro and in vivo genetic fitness of Pseudomonas aeruginosa using high-throughput sequencing of transposon libraries. PLoS Pathog. 9, e1003582 (2013).
Crofts, A. A. et al. Enterotoxigenic E. coli virulence gene regulation in human infections. Proc. Natl Acad. Sci. USA 115, 8968–8976 (2018).
Sheikh, A. et al. In vivo expression of Salmonella enterica serotype Typhi genes in the blood of patients with typhoid fever in Bangladesh. PLoS Negl. Trop. Dis. 5, e1419 (2011).
Subashchandrabose, S., Smith, S. N., Spurbeck, R. R., Kole, M. M. & Mobley, H. L. T. Genome-wide detection of fitness genes in uropathogenic Escherichia coli during systemic infection. PLoS Pathog. 9, e1003788 (2013).
Silva-Valenzuela, C. A. et al. Analysis of two complementary single-gene deletion mutant libraries of Salmonella Typhimurium in intraperitoneal infection of BALB/c mice. Front. Microbiol. 6, 1–9 (2016).
Valentino, M. D. et al. Genes contributing to Staphylococcus aureus fitness in abscess- and infection-related ecologies. mBio 5, e01729–14 (2014).
Streit, W. R. & Entcheva, P. Biotin in microbes, the genes involved in its biosynthesis, its biochemical role and perspectives for biotechnological production. Appl. Microbiol. Biotechnol. 61, 21–31 (2003).
Lin, S., Hanson, R. E. & Cronan, J. E. Biotin synthesis begins by hijacking the fatty acid synthetic pathway. Nat. Chem. Biol. 6, 682–688 (2010).
Okami, Y. et al. Studies on a new amino acid antibiotic, amiclenomycin. J. Antibiot. 27, 656–664 (1974).
Kitahara, T., Hotta, K., Yoshida, M. & Okami, Y. Biological studies on amiclenomycin. J. Antibiot. 28, 215–222 (1975).
Dai, R., Wilson, D. J., Geders, T. W., Aldrich, C. C. & Finzel, B. C. Inhibition of Mycobacterium tuberculosis transaminase BioA by aryl hydrazines and hydrazides. Chembiochem 15, 575–586 (2014).
Park, S. W. et al. Target-based identification of whole-cell active inhibitors of biotin biosynthesis in Mycobacterium tuberculosis. Chem. Biol. 22, 76–86 (2015).
Liu, F. et al. Structure-based optimization of pyridoxal 5′-phosphate-dependent transaminase enzyme (BioA) inhibitors that target biotin biosynthesis in Mycobacterium tuberculosis. J. Med. Chem. 60, 5507–5520 (2017).
Salaemae, W., Booker, G. W. & Polyak, S. W. Genetic requirements for mycobacterial survival during infection. Microbiol. Spectr. 4, VMBF-0008-2015 (2016).
Mock, D. M. & Malik, M. I. Distribution of biotin in human plasma: most of the biotin is not bound to protein. Am. J. Clin. Nutr. 56, 427–432 (1992).
Trüeb, R. M. Serum biotin levels in women complaining of hair loss. Int. J. Trichology 8, 73–77 (2016).
Harthe, C. & Claustrat, B. A sensitive and practical competitive radioassay for plasma biotin. Ann. Clin. Biochem. 40, 259–263 (2003).
Perry, C. A. et al. Pregnancy and lactation alter biomarkers of biotin metabolism in women consuming a controlled diet. J. Nutr. 144, 1977–1984 (2014).
Wakabayashi, K. et al. Serum biotin in Japanese children: enzyme-linked immunosorbent assay measurement. Pediatr. Int. 58, 872–876 (2016).
Whiteside, M. D., Winsor, G. L., Laird, M. R. & Brinkman, F. S. L. OrtholugeDB: a bacterial and archaeal orthology resource for improved comparative genomic analysis. Nucleic Acids Res. 41, 366–376 (2013).
Zlitni, S., Ferruccio, L. F. & Brown, E. D. Metabolic suppression identifies new antibacterial inhibitors under nutrient limitation. Nat. Chem. Biol. 9, 796–804 (2013).
Tiwari, D. et al. Targeting protein biotinylation enhances tuberculosis chemotherapy. Sci. Transl. Med. 10, eaal1803 (2018).
El Zahed, S. S. & Brown, E. D. Chemical–chemical combinations map uncharted interactions in Escherichia coli under nutrient stress. iScience 2, 168–181 (2018).
Legrand, N. et al. Humanized mice for modeling human infectious disease: challenges, progress, and outlook. Cell Host Microbe 6, 5–9 (2009).
Lin, S. & Cronan, J. E. The BioC O-methyltransferase catalyzes methyl esterification of malonyl-acyl carrier protein, an essential step in biotin synthesis. J. Biol. Chem. 287, 37010–37020 (2012).
Ploux, O., Breyne, O., Carillon, S. & Marquet, A. Slow-binding and competitive inhibition of 8-amino-7-oxopelargonate synthase, a pyridoxal-5′-phosphate-dependent enzyme involved in biotin biosynthesis, by substrate and intermediate analogs. Eur. J. Biochem. 259, 63–70 (1999).
Hanka, L. J., Martin, D. G. & Reineke, L. M. Two new antimetabolites of biotin: α-methyldethiobiotin and α-methylbiotin. Antimicrob. Agents Chemother. 1, 135–138 (1972).
Eisenberg, M. A. & Hsiung, S. C. Mode of action of the biotin antimetabolites actithiazic acid and α-methyldethiobiotin. Antimicrob. Agents Chemother. 21, 5–10 (1982).
Alexeev, D. et al. Rational design of an inhibitor of dethiobiotin synthetase; interaction of 6-hydroxypyrimindin-4(3H)-one with the adenine base binding site. Tetrahedron 54, 15891–15898 (1998).
Bockman, M. R. et al. Investigation of (S)-(−)-acidomycin: a selective antimycobacterial natural product that inhibits biotin synthase. ACS Infect. Dis. 5, 598–617 (2019).
Taira, J. et al. Identification of a novel class of small compounds with anti-tuberculosis activity by in silico structure-based drug screening. J. Antibiot. 70, 1057–1064 (2017).
Datta, S., Costantino, N. & Court, D. L. A set of recombineering plasmids for Gram-negative bacteria. Gene 379, 109–115 (2006).
Datsenko, Ka & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
Huang, T. W. et al. Capsule deletion via a λ-Red knockout system perturbs biofilm formation and fimbriae expression in Klebsiella pneumoniae MGH 78578. BMC Res. Notes 7, 13 (2014).
Gallagher, L. A. et al. Resources for genetic and genomic analysis of emerging pathogen Acinetobacter baumannii. J. Bacteriol. 197, 2027–2035 (2015).
Held, K., Ramage, E., Jacobs, M., Gallagher, L. & Manoil, C. Sequence-verified two-allele transposon mutant library for Pseudomonas aeruginosa PAO1. J. Bacteriol. 194, 6387–6389 (2012).
Fey, P. D. et al. A genetic resource for rapid and comprehensive phenotype screening of nonessential Staphylococcus aureus genes. mBio 4, e00537–12 (2013).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. Sect. D 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. Sect. D 60, 2126–2132 (2004).
Mann, S., Eveleigh, L., Lequin, O. & Ploux, O. A microplate fluorescence assay for DAPA aminotransferase by detection of the vicinal diamine 7,8-diaminopelargonic acid. Anal. Biochem. 432, 90–96 (2013).
Acknowledgements
We would like to thank S. McCusker from the Centre for Microbial Chemical Biology and G. Wright for bacterial strains from the Institute for Infectious Disease Research clinical collection, A. Fiebig-Comyn for amendments to the animal use protocol, National Institute of Allergy and Infectious Diseases preclinical services for pharmacokinetic testing, A. Eakin for advice and guidance (contract HHSN272201100022I awarded to SRI, International) and R. Melano at Public Health Ontario for bacterial strains GB687 and C0064. This research was supported by a Foundation grant from the Canadian Institutes for Health Research (FDN-143215), a donation from the Boris Family Foundation and by funding from the Ontario Research Fund Research Excellence program (RE07-048). E.D.B. was supported by a salary award from the Canada Research Chairs program. L.A.C. was supported by an Ontario Graduate Scholarship Award and a Canadian Institutes for Health Research scholarship.
Author information
Authors and Affiliations
Contributions
L.A.C. conceived the research, designed and carried out experiments and data analysis, and wrote the manuscript. C.R.M. assisted with data acquisition and interpretation and manuscript editing. L.A.C designed and conducted the in vitro assays to determine biotin requirement and in vitro enzyme assays. C.N.T. performed the orthologue search and phylogenetic analysis. L.A.C. and B.S.W. designed and performed the plasma growth assays. C.M.B., S.Z. and J.C. designed and performed crystallization experiments and C.M.B performed the analysis with input from M.S.J. L.A.C. and V.N.R. performed susceptibility testing. L.A.C and C.R.M. designed and performed in vivo infection model experiments. B.S.W., M.S.J. and B.K.C. assisted with data interpretation. E.D.B. conceived the research and assisted with data interpretation and manuscript editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Biotin requirements of BioA deficient strains represent the entirety of the biosynthetic pathway.
Analysis of the biotin concentrations required to restore growth of (a) A. baumannii transposon mutants in bioACDF and (b) E. coli ΔbioABCDF in M9 minimal media supplemented with amino acids after 18 hours. Points show the mean of 8 replicates ± s.d. and the solid lines show a four-parameter dose response curve. (c) Presence or absence of a high-affinity biotin transporter in a phylogenetic tree of pathogenic bacteria with an ortholog to E. coli BioA. The presence of an ortholog to biotin transporters, E. coli BioP and S. aureus BioY was determined using OrtholugeDB. Species with a predicted ortholog to E. coli BioP (green) or S. aureus BioY (blue) transporters are demarked by a circle.
Extended Data Fig. 2 Biotin levels are comparable among three common mouse strains.
(a) Quantification of biotin levels in purchased human and mouse plasma from Innovative Research. Each point represents a technical replicate (n = 4) and the line is the mean. (b) Quantification of biotin in the plasma of BALB/c (n = 6), CD-1 (n = 7), and C57Bl/6 (n = 8) mice. Each point represents the biotin levels of a single mouse and the line is the mean. (c) Fold change in viable cell counts of BioA deficient (grey) and wild-type (black) A. baumannii (AB), K. pneumoniae (KP), P. aeruginosa (PA), and S. Typhimurium (ST) growth after 24 hours in 50% human plasma supplemented with biotin (10 µg/mL). BioA deficient or wild-type strains were separately inoculated into plasma diluted with M9 salts supplemented with 25 mM of sodium bicarbonate. Viable cell counts were calculated at time 0 and 24 hours. Fold change was calculated by dividing the viable cell count at 24 hours by the viable cell count at time 0. The bars indicate the mean of 6 replicates ± s.d.
Extended Data Fig. 3 Biotin biosynthesis is dispensable for A. baumannii, K. pneumoniae, P. aeruginosa, S. aureus, and S. Typhimurium during standard infection models.
Bacterial load in the blood, spleen, kidneys, liver, and lungs following a systemic infection with (a) A. baumannii (AB; n = 5), (b) K. pneumoniae (KP; n=10), (c) P. aeruginosa (PA; n=9), (d) S. aureus (SA; n=6), and (e) S. Typhimurium (ST; n=5). Mice were co-infected with a mixed inoculum of wild- type (grey) and BioA deficient (black) bacteria inoculated by intraperitoneal injection. (f) Bacterial load in the lungs of mice co-infected with a mixed inoculum of wild-type (grey) and ΔbioA (black) bacteria. Infection was established by intranasal administration of 2×107 CFU by micropipette and mice (n=10) euthanized 36 hours later. (g) Bacterial load in the thighs of mice co-infected with a mixed inoculum of wild-type (grey) and BioA deficient (black) A. baumannii (AB; n=6), K. pneumoniae (KP; n=8), P. aeruginosa (PA; n=4), and S. aureus (SA; n=10). 5×105 (KP and SA) or 5×106 (AB and PA) CFU were injected into the thighs of mice. In all instances box plot whiskers show the minimum to maximum values, the box denotes the interquartile range, and the line in the box shows the median.
Extended Data Fig. 4 Determination of an optimal dose of streptavidin to mimic human biotin levels.
Quantification of biotin in the plasma of CD-1 mice following intraperitoneal administration of (a) streptavidin (1 mg/kg) or (b) streptavidin (4 mg/kg) treatment. Each point represents the biotin levels of a single mouse (n = 2–7) and the line indicates the mean. The grey area indicates the typical range of human biotin concentration in plasma accounting for daily fluctuations. (c) Streptavidin pretreatment has no effect on the ability of wild-type bacteria to cause infection. A. baumannii bacterial load in the blood, spleen, kidney, liver, and lungs in a standard infection model (n=6) or mice pretreated with streptavidin (2 mg/kg; n=4). Mice were infected by intraperitoneal injection and euthanized 10 hours post- infection. Each point represents the bacterial load from an individual mouse and the line indicates the mean.
Extended Data Fig. 5 Biotin biosynthesis is a critical fitness determinant for A. baumannii, K. pneumoniae, and P. aeruginosa during a systemic infection with human biotin levels.
Competitive index (CI) for the co-infection of BioA deficient and wild-type strains of (a) A. baumannii (n=5, 8), (b) K. pneumoniae (n=10, 8), (c) P. aeruginosa (n=9, 8), and (d) S. aureus (n=6, 9) in the blood, spleen, kidneys, liver, and lungs in a standard systemic infection model (light blue) or systemic infection in mice pretreated with streptavidin (2 mg/kg; blue) 1-hour prior to inoculation. CI is calculated by dividing the bacterial load of the query strain (BioA deficient) by the bacterial load of the wild-type strain. Each point represents the CI from an individual mouse and the line indicates the mean. The dotted line represents a CI of 1 indicating the two strains are proliferating equally in vivo. Groups were analyzed with an unpaired two-tailed t-test and corrected for multiple comparisons with a Holm-Sidak test, * indicates a p<0.01.
Extended Data Fig. 6 Biotin biosynthesis is a critical fitness determinant for A. baumannii, K. pneumoniae, and P. aeruginosa during a systemic infection with human biotin levels.
Bacterial load in the blood, spleen, kidneys, liver, and lungs following a systemic infection with human biotin levels with (a) A. baumannii (n=8), (b) K. pneumoniae (n=8), (c) P. aeruginosa (n=8), and (d) S. aureus (n=9). Mice were pretreated with streptavidin (2 mg/kg) 1-hour prior to co- infection with a mixed inoculum of wild-type (grey) and BioA deficient (black) bacteria. Box plot whiskers show the minimum to maximum values, the box denotes the interquartile range, and the line in the box shows the median of each group. Groups were analyzed with an unpaired two-tailed t-test and corrected for multiple comparisons with a Holm-Sidak test, * indicates a p<0.01.
Extended Data Fig. 7 Biotin biosynthesis is dispensable during colonization of a systemic murine infection mimicking human biotin levels.
Bacterial load in the blood, spleen, kidneys, liver, and lungs 1- hour post-infection following a systemic infection with human biotin levels with (a) A. baumannii (AB; n=3), (b) K. pneumoniae (KP; n=2), and (c) P. aeruginosa (PA; n=3). Mice were pretreated with streptavidin (2 mg/kg) 1-hour prior to co- infection with a mixed inoculum of wild-type (grey) and BioA deficient (black) bacteria. Each point represents the bacterial load from an individual mouse and the line indicates the mean.
Extended Data Fig. 8 Resistance to inhibition of biotin biosynthesis imparts a fitness cost.
Kinetics of (7,8)-di-amino-pelargonic acid production. Initial velocities of (7,8)-di-amino-pelargonic acid production with BioA, BioAW52A, BioAK274A, BioAF393A, and BioAY398A were determined at various (a) (S)-keto-amino-pelargonic acid (KAPA; n=4) and (b) S-adenosyl-L-methionine (SAM; n=6) concentrations. Points show the mean of replicates ± s.d. and the solid lines show a Michaelis-Menten curve. (c) Analysis of the biotin concentrations required to restore growth of BioA deficient A. baumannii passaged into the lowest biotin concentration supporting growth for 14 days. Points show the mean of 8 replicates ± s.d. and the solid lines show a four- parameter dose response curve.
Extended Data Fig. 9 MAC13772 is efficacious against A. baumannii in a systemic infection with human biotin levels.
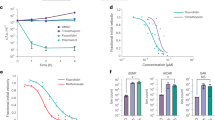
Bacterial load of A. baumannii in the blood, spleen, kidneys, liver, and lungs following a systemic infection with (a) standard biotin levels and (b) human biotin levels, treated with MAC13772 (15 mg/kg). Mice were pretreated with streptavidin (2 mg/kg) 1-hour prior infection and treated with MAC13772 (15 mg/kg) 1-hour following infection. Groups were analyzed with an unpaired two-tailed t-test and corrected for multiple comparisons with a Holm-Sidak test, * indicates a p<0.01. (c) Time course of plasma concentrations of MAC13772 in male (black) and female (grey) Sprague Dawley rats, after intravenous administration. Each data point represents the mean ± s.d. concentration from n=3 rats, the dotted line indicates the limit of quantification (LOQ).
Supplementary information
Supplementary Information
Supplementary Tables 1, 3–6 and 8–11, legends for Supplementary Tables 2 and 7, Supplementary Figures 1–3 and Supplementary References.
Supplementary Tables 2 and 7
Supplemental Table 2: OrtholugeDB predictions for the presence of an orthologue to E. coli BioP and S. aureus BioY. Supplemental Table 7: prediction of the presence of Phe393, Tyr144, Trp52, Tyr398 and Lys274 in 100 bacteria.
Rights and permissions
About this article
Cite this article
Carfrae, L.A., MacNair, C.R., Brown, C.M. et al. Mimicking the human environment in mice reveals that inhibiting biotin biosynthesis is effective against antibiotic-resistant pathogens. Nat Microbiol 5, 93–101 (2020). https://doi.org/10.1038/s41564-019-0595-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0595-2
This article is cited by
-
Dual proteomics of infected macrophages reveal bacterial and host players involved in the Francisella intracellular life cycle and cell to cell dissemination by merocytophagy
Scientific Reports (2024)
-
Inhibiting fatty acid synthesis overcomes colistin resistance
Nature Microbiology (2023)
-
Unrealized targets in the discovery of antibiotics for Gram-negative bacterial infections
Nature Reviews Drug Discovery (2023)
-
Biotin-dependent cell envelope remodelling is required for Mycobacterium abscessus survival in lung infection
Nature Microbiology (2023)