Abstract
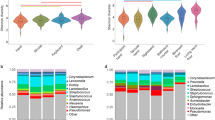
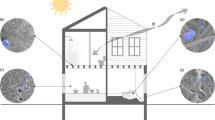
Urbanization represents a profound shift in human behaviour, and has considerable cultural and health-associated consequences1,2. Here, we investigate chemical and microbial characteristics of houses and their human occupants across an urbanization gradient in the Amazon rainforest, from a remote Peruvian Amerindian village to the Brazilian city of Manaus. Urbanization was found to be associated with reduced microbial outdoor exposure, increased contact with housing materials, antimicrobials and cleaning products, and increased exposure to chemical diversity. The degree of urbanization correlated with changes in the composition of house bacterial and microeukaryotic communities, increased house and skin fungal diversity, and an increase in the relative abundance of human skin-associated fungi and bacteria in houses. Overall, our results indicate that urbanization has large-scale effects on chemical and microbial exposures and on the human microbiota.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Mass spectrometry data have been deposited in MassIVE (accession number MSV000082924). Molecular networking jobs are available at https://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=549cfafcdaef4a7496768f45bb90771c (full housing dataset). GNPS molecular networking jobs for dataset matching are available at https://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=f1bc8144b7f648dc94215a34b94537df (Checherta), https://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=044f5c1437e4453eb9d47afafadb7cfb (Puerto Almendra), https://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=c3736e2379d1470f8b9e990a1afeb682 (Iquitos), https://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=8799f335311540bdb5af75da896bd87c (Manaus low income) and https://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=1070c36112c940a186fcf4454f845f08 (Manaus middle class; searches performed 19 August 2018). In silico structure annotation using NAP is provided at https://proteomics2.ucsd.edu/ProteoSAFe/status.jsp?task=af425ada55d54adca9c7b28a823af54c. The raw sequencing data and processed BIOM tables are available at Qiita (https://qiita.ucsd.edu/) under study ID 10333, and also at EMBL-EBI under submission number ERP107551.
Code availability
Instructions and source codes for replicating the bioinformatic analyses are provided at https://github.com/knightlab-analyses/amazon-urbanization.
Change history
24 January 2020
In the version of this Letter originally published, Maria G. Dominguez-Bello’s e-mail address was missing a hyphen; it has now been corrected to mg.dominguez-bello@rutgers.edu.
References
Oyebode, O. et al. Rural, urban and migrant differences in non-communicable disease risk-factors in middle income countries: a cross-sectional study of WHO-SAGE data. PLoS ONE 10, e0122747 (2015).
Ege, M. J. et al. Exposure to environmental microorganisms and childhood asthma. N. Engl. J. Med. 364, 701–709 (2011).
World Population Prospects: 2015 Revision (United Nations Publications, 2016).
Yatsunenko, T. et al. Human gut microbiome viewed across age and geography. Nature 486, 222–227 (2012).
De Filippo, C. et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc. Natl Acad. Sci. USA 107, 14691–14696 (2010).
Clemente, J. C. et al. The microbiome of uncontacted Amerindians. Sci. Adv. 1, e1500183 (2015).
Obregon-Tito, A. J. et al. Subsistence strategies in traditional societies distinguish gut microbiomes. Nat. Commun. 6, 6505 (2015).
Ruiz-Calderon, J. F. et al. Walls talk: microbial biogeography of homes spanning urbanization. Sci. Adv. 2, e1501061 (2016).
da Silva, R. R. et al. Propagating annotations of molecular networks using in silico fragmentation. PLoS Comput. Biol. 14, e1006089 (2018).
Perlin, D. S., Rautemaa-Richardson, R. & Alastruey-Izquierdo, A. The global problem of antifungal resistance: prevalence, mechanisms, and management. Lancet Infect. Dis. 17, e383–e392 (2017).
Human Microbiome Project Consortium Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Findley, K. et al. Topographic diversity of fungal and bacterial communities in human skin. Nature 498, 367–370 (2013).
Oh, J. et al. Biogeography and individuality shape function in the human skin metagenome. Nature 514, 59–64 (2014).
Wang, M. et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat. Biotechnol. 34, 828–837 (2016).
McCall, L.-I. & McKerrow, J. H. Determinants of disease phenotype in trypanosomatid parasites. Trends Parasitol. 30, 342–349 (2014).
Stagaman, K. et al. Market integration predicts human gut microbiome attributes across a gradient of economic development. mSystems 3, e00122-17 (2018).
Adams, R. I. et al. Ten questions concerning the microbiomes of buildings. Build. Environ. 109, 224–234 (2016).
Caporaso, J. G. et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl Acad. Sci. USA 108(Suppl 1), 4516–4522 (2011).
Thompson, L. R. et al. A communal catalogue reveals Earth’s multiscale microbial diversity. Nature 551, 457–463 (2017).
Salter, S. J. et al. Reagent and laboratory contamination can critically impact sequence-based microbiome analyses. BMC Biol. 12, 87 (2014).
Liu, C. M. et al. BactQuant: an enhanced broad-coverage bacterial quantitative real-time PCR assay. BMC Microbiol. 12, 56 (2012).
Gil-Serna, J., González-Salgado, A., González-Jaén, M. A. T., Vázquez, C. & Patiño, B. ITS-based detection and quantification of Aspergillus ochraceus and Aspergillus westerdijkiae in grapes and green coffee beans by real-time quantitative PCR. Int. J. Food Microbiol. 131, 162–167 (2009).
Schrader, C., Schielke, A., Ellerbroek, L. & Johne, R. PCR inhibitors—occurrence, properties and removal. J. Appl. Microbiol. 113, 1014–1026 (2012).
Bouslimani, A. et al. Molecular cartography of the human skin surface in 3D. Proc. Natl Acad. Sci. USA 112, E2120–E2129 (2015).
Bouslimani, A. et al. Lifestyle chemistries from phones for individual profiling. Proc. Natl Acad. Sci. USA 113, E7645–E7654 (2016).
Protsyuk, I. et al. 3D molecular cartography using LC–MS facilitated by Optimus and ’ili software. Nat. Protoc. 13, 134–154 (2018).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Anderson, M. J. A new method for non-parametric multivariate analysis of variance. Austral Ecol. 26, 32–46 (2001).
Watrous, J. et al. Mass spectral molecular networking of living microbial colonies. Proc. Natl Acad. Sci. USA 109, E1743–E1752 (2012).
Scheubert, K. et al. Significance estimation for large scale metabolomics annotations by spectral matching. Nat. Commun. 8, 1494 (2017).
Djoumbou Feunang, Y. et al. ClassyFire: automated chemical classification with a comprehensive, computable taxonomy. J. Cheminform. 8, 61 (2016).
Ernst, M. et al. MolNetEnhancer: enhanced molecular networks by integrating metabolome mining and annotation tools. Metabolites 9, E144 (2019).
Sumner, L. W. et al. Proposed minimum reporting standards for chemical analysis. Metabolomics 3, 211–221 (2007).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Gonzalez, A. et al. Qiita: rapid, web-enabled microbiome meta-analysis. Nat. Methods 15, 796–798 (2018).
Amir, A. et al. Deblur rapidly resolves single-nucleotide community sequence patterns. mSystems 2, e00191-16 (2017).
Kopylova, E., Noé, L. & Touzet, H. SortMeRNA: fast and accurate filtering of ribosomal RNAs in metatranscriptomic data. Bioinformatics 28, 3211–3217 (2012).
McDonald, D. et al. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 6, 610–618 (2012).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Kõljalg, U. et al. Towards a unified paradigm for sequence-based identification of fungi. Mol. Ecol. 22, 5271–5277 (2013).
Falony, G. et al. Population-level analysis of gut microbiome variation. Science 352, 560–564 (2016).
Oksanen, J et al. vegan: Community ecology package. R package version 2.5-6 https://CRAN.R-project.org/package=vegan (2019).
Gower, J. C. Generalized procrustes analysis. Psychometrika 40, 33–51 (1975).
Kapono, C. A. et al. Creating a 3D microbial and chemical snapshot of a human habitat. Sci. Rep. 8, 3669 (2018).
Roberts, D. W. labdsv: Ordination and multivariate analysis for ecology. R package version 1.8-0 (2007).
Acknowledgements
We thank the collaborators in Peru, the late L. Bocca and the interpreter J. J. Semu for their support and information sources; in Manaus, A. Vasconcelos and J. Machado for help with the fieldwork, and the support of the community leader M. Aparecida Lisboa, Director of Association Fazendo Amigos (AFA), Manaus; the nurses that accompanied the researchers in the jungle; I. Fajardo Neddermann for helping us with the architectural work; D. Vargas and M. Magris, who helped to prepare the urbanization score survey; D. McDonald, J. Morton, R. da Silva and L. Jiang, who helped with DNA and MS data analysis. A. Cai helped to determine the sources of microorganisms correlated with cleaning products; in Peru, staff and community members participating within the Malaria Immunology and Genetics in the Amazon Project with the Ministry of Health of Peru for support. This study was supported by the Sloan Foundation (to M.G.D.-B., R.K. and P.C.D.), C&D Fund, and Emch Fund for Microbial Diversity (to M.G.D.-B.). Partial support was also provided by the NIH Research Initiative for Scientific Enhancement Program 2R25GM061151-13 (to J.F.R.-C.). L.-I.M. was partially supported by a fellowship from the Canadian Institutes of Health Research (grant no. 338511; http://www.cihr-irsc.gc.ca/). C.C. was supported by the Belgian American Educational Foundation and the Research Foundation Flanders. We acknowledge the NIH for providing the MS and MS data analysis infrastructure P41-GM103484 and GMS10RR029121 (to P.C.D.).
Author information
Authors and Affiliations
Contributions
M.G.D.-B., P.C.D. and R.K. conceived and designed the study. M.G.D.-B., J.F.R.-C., H.S.P., J.H., R.R., O.H.B., M.J.B., L.C.P., A.N. and H.C. collected the samples and metadata. A.B. acquired LC–MS data. L.-I.M. led the LC–MS data analysis. C.C. led the taxonomy and metadata analysis. Q.Z. led the DNA data and multi-omics analysis. J.J.M. performed qPCR. S.J.S., M.E., H.C., A.N., A.B. and J.J.M. provided additional contributions to data analysis. L.-I.M., C.C., Q.Z. and M.G.D.-B. wrote the manuscript with contributions from R.K. and P.C.D. All of the authors reviewed the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Notes, Supplementary Tables 1–6, Supplementary Figs. 1–22, Supplementary References and legends for Supplementary Data 1–5.
Supplementary Data 1
Cleaning product metabolite feature table.
Supplementary Data 2
Pearson correlation analysis between relative abundances of cleaning products and bacteria (n = 256).
Supplementary Data 3
Pearson correlation analysis between relative abundances of cleaning products and fungi (n = 140).
Supplementary Data 4
Pearson correlation analysis between relative abundances of cleaning products and microeukaryotes (n = 82).
Supplementary Data 5
Urbanization score calculation tool.
Rights and permissions
About this article
Cite this article
McCall, LI., Callewaert, C., Zhu, Q. et al. Home chemical and microbial transitions across urbanization. Nat Microbiol 5, 108–115 (2020). https://doi.org/10.1038/s41564-019-0593-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0593-4
This article is cited by
-
Functional host-specific adaptation of the intestinal microbiome in hominids
Nature Communications (2024)
-
Early-life interactions between the microbiota and immune system: impact on immune system development and atopic disease
Nature Reviews Immunology (2023)
-
Impact of Hurricane Maria on mold levels in the homes of Piñones, Puerto Rico
Air Quality, Atmosphere & Health (2023)
-
Ecological Processes and Human Behavior Provide a Framework for Studying the Skin Microbial Metacommunity
Microbial Ecology (2022)
-
Challenges in exploring and manipulating the human skin microbiome
Microbiome (2021)