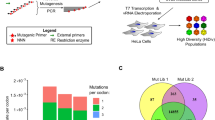
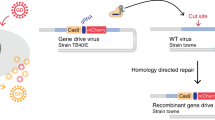
Abstract
RNA virus populations are composed of highly diverse individuals that form a cloud of related sequences commonly referred to as a ‘quasispecies’1,2,3. This diversity arises as a consequence of low-fidelity genome replication4,5. By contrast, DNA virus populations contain more uniform individuals with similar fitness6. Genome diversity is often correlated with increased fitness in RNA viruses, while DNA viruses are thought to require more faithful genome replication. During DNA replication, erroneously incorporated bases are removed by a 3′-5′ exonuclease, a highly conserved enzymatic function of replicative DNA but not RNA polymerases. This proofreading process enhances replication fidelity and ensures the genome integrity of DNA organisms, including large DNA viruses7. Here, we show that a herpesvirus can tolerate impaired exonucleolytic proofreading, resulting in DNA virus populations, which, as in RNA viruses8, are composed of highly diverse genotypes of variable individual fitness. This indicates that herpesvirus mutant diversity may compensate for individual fitness loss. Notably, in vivo infection with diverse virus populations results in a marked increase in virulence compared to genetically homogenous parental virus. While we cannot exclude that the increase in virulence is caused by selection of and/or interactions between individual genotypes, our findings are consistent with quasispecies dynamics. Our results contrast with traditional views of DNA virus replication and evolution, and indicate that a substantial increase in population diversity can lead to higher virulence.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The SNPs analysed in this study are supplied as Supplementary Tables. Illumina short reads (FASTQ) have been uploaded to the Short Read Archive and can be accessed under Bioproject no. PRJNA553690.
Code availability
The custom code used to analyse the sequencing data is available at the public repository github (https://github.com/NG-viro/Shannon_Entropy).
References
Andino, R. & Domingo, E. Viral quasispecies. Virology 479–480, 46–51 (2015).
Bordería, A. V., Rozen-Gagnon, K. & Vignuzzi, M. Fidelity variants and RNA quasispecies. Curr. Top. Microbiol. Immunol. 392, 303–322 (2016).
Domingo, E. & Schuster, P. What is a quasispecies? Historical origins and current scope. Curr. Top. Microbiol. Immunol. 392, 1–22 (2016).
Drake, J. W. The distribution of rates of spontaneous mutation over viruses, prokaryotes, and eukaryotes. Ann. N. Y. Acad. Sci. 870, 100–107 (1999).
Dolan, P. T., Whitfield, Z. J. & Andino, R. Mechanisms and concepts in RNA virus population dynamics and evolution. Annu. Rev. Virol. 5, 69–92 (2018).
Fornés, J., Tomás Lázaro, J., Alarcón, T., Elena, S. F. & Sardanyés, J. Viral replication modes in single-peak fitness landscapes: a dynamical systems analysis. J. Theor. Biol. 460, 170–183 (2019).
Bębenek, A. & Ziuzia-Graczyk, I. Fidelity of DNA replication: a matter of proofreading. Curr. Genet. 64, 985–996 (2018).
Villarreal, L. P. & Witzany, G. Rethinking quasispecies theory: from fittest type to cooperative consortia. World J. Biol. Chem. 4, 79–90 (2013).
Osterrieder, N., Kamil, J. P., Schumacher, D., Tischer, B. K. & Trapp, S. Marek’s disease virus: from miasma to model. Nat. Rev. Microbiol. 4, 283–294 (2006).
Trimpert, J. et al. A phylogenomic analysis of Marek’s disease virus reveals independent paths to virulence in Eurasia and North America. Evol. Appl. 10, 1091–1101 (2017).
Szpara, M. L. et al. Evolution and diversity in human herpes simplex virus genomes. J. Virol. 88, 1209–1227 (2014).
Weller, S. K. & Coen, D. M. Herpes simplex viruses: mechanisms of DNA replication. Cold Spring Harb. Perspect. Biol. 4, a013011 (2012).
Hwang, C. B.-C. in DNA Replication and Related Cellular Processes (ed. Kušić-Tišma, J.) 143–160 (InTech, 2011).
Song, L., Chaudhuri, M., Knopf, C. W. & Parris, D. S. Contribution of the 3′- to 5′-exonuclease activity of herpes simplex virus type 1 DNA polymerase to the fidelity of DNA synthesis. J. Biol. Chem. 279, 18535–18543 (2004).
Lawler, J. L. & Coen, D. M. HSV-1 DNA polymerase 3′-5′ exonuclease-deficient mutant D368A exhibits severely reduced viral DNA synthesis and polymerase expression. J. Gen. Virol. 99, 1432–1437 (2018).
Hall, J. D., Orth, K. L., Sander, K. L., Swihart, B. M. & Senese, R. A. Mutations within conserved motifs in the 3′−5′ exonuclease domain of herpes simplex virus DNA polymerase. J. Gen. Virol. 76, 2999–3008 (1995).
Hwang, C. B., Ruffner, K. L. & Coen, D. M. A point mutation within a distinct conserved region of the herpes simplex virus DNA polymerase gene confers drug resistance. J. Virol. 66, 1774–1776 (1992).
Hwang, Y. T., Liu, B. Y., Hong, C. Y., Shillitoe, E. J. & Hwang, C. B. Effects of exonuclease activity and nucleotide selectivity of the herpes simplex virus DNA polymerase on the fidelity of DNA replication in vivo. J. Virol. 73, 5326–5332 (1999).
Crotty, S., Cameron, C. E. & Andino, R. RNA virus error catastrophe: direct molecular test by using ribavirin. Proc. Natl Acad. Sci. USA 98, 6895–6900 (2001).
Lauring, A. S., Frydman, J. & Andino, R. The role of mutational robustness in RNA virus evolution. Nat. Rev. Microbiol. 11, 327–336 (2013).
Taddei, F. et al. Role of mutator alleles in adaptive evolution. Nature 387, 700–702 (1997).
Mansky, L. M. & Cunningham, K. S. Virus mutators and antimutators: roles in evolution, pathogenesis and emergence. Trends Genet. 16, 512–517 (2000).
Xu, M., Fitzgerald, S. D., Zhang, H., Karcher, D. M. & Heidari, M. Very virulent plus strains of MDV induce an acute form of transient paralysis in both susceptible and resistant chicken lines. Viral Immunol. 25, 306–323 (2012).
Pandey, U. et al. DNA from dust: comparative genomics of large DNA viruses in field surveillance samples. mSphere 1, e00132-16 (2016).
Jaramillo, N. et al. Evidence of Muller’s ratchet in herpes simplex virus type 1. J. Gen. Virol. 94, 366–375 (2013).
Sanjuán, R., Nebot, M. R., Chirico, N., Mansky, L. M. & Belshaw, R. Viral mutation rates. J. Virol. 84, 9733–9748 (2010).
Xiao, Y. et al. RNA recombination enhances adaptability and is required for virus spread and virulence. Cell Host Microbe 19, 493–503 (2016).
Vignuzzi, M., Stone, J. K., Arnold, J. J., Cameron, C. E. & Andino, R. Quasispecies diversity determines pathogenesis through cooperative interactions in a viral population. Nature 439, 344–348 (2006).
Fitzsimmons, W. J. et al. A speed-fidelity trade-off determines the mutation rate and virulence of an RNA virus. PLoS Biol. 16, e2006459 (2018).
Holmes, E. C. The RNA virus quasispecies: fact or fiction? J. Mol. Biol. 400, 271–273 (2010).
Bull, J. J., Meyers, L. A. & Lachmann, M. Quasispecies made simple. PLoS Comput. Biol. 1, e61 (2005).
Lauring, A. S. & Andino, R. Quasispecies theory and the behavior of RNA viruses. PLoS Pathog. 6, e1001005 (2010).
Leeks, A., Segredo-Otero, E. A., Sanjuán, R. & West, S. A. Beneficial coinfection can promote within-host viral diversity. Virus Evol. 4, vey028 (2018).
Jarosinski, K. W., Arndt, S., Kaufer, B. B. & Osterrieder, N. Fluorescently tagged pUL47 of Marek’s disease virus reveals differential tissue expression of the tegument protein in vivo. J. Virol. 86, 2428–2436 (2012).
Schat, K. A. & Purchase, H. in A Laboratory Manual for the Isolation and Identification of Avian Pathogens (ed. Swayne, D. E.) 223–234 (American Association of Avian Pathologists, 1998).
Longo, P. A., Kavran, J. M., Kim, M.-S. & Leahy, D. J. Transient mammalian cell transfection with polyethylenimine (PEI). Methods Enzymol. 529, 227–240 (2013).
Schumacher, D., Tischer, B. K., Fuchs, W. & Osterrieder, N. Reconstitution of Marek’s disease virus serotype 1 (MDV-1) from DNA cloned as a bacterial artificial chromosome and characterization of a glycoprotein B-negative MDV-1 mutant. J. Virol. 74, 11088–11098 (2000).
Schumacher, D., Tischer, B. K., Trapp, S. & Osterrieder, N. The protein encoded by the US3 orthologue of Marek’s disease virus is required for efficient de-envelopment of perinuclear virions and involved in actin stress fiber breakdown. J. Virol. 79, 3987–3997 (2005).
Tischer, B. K., Smith, G. A. & Osterrieder, N. En passant mutagenesis: a two step markerless red recombination system. Methods Mol. Biol. 634, 421–430 (2010).
Blin, N. & Stafford, D. W. A general method for isolation of high molecular weight DNA from eukaryotes. Nucleic Acids Res. 3, 2303–2308 (1976).
Sambrook, J. & Russell, D. W. Molecular Cloning: a Laboratory Manual 3rd edn (Cold Spring Harbor Lab. Press, 2001).
Hirt, B. Selective extraction of polyoma DNA from infected mouse cell cultures. J. Mol. Biol. 26, 365–369 (1967).
Baigent, S. J. et al. Herpesvirus of turkey reconstituted from bacterial artificial chromosome clones induces protection against Marek’s disease. J. Gen. Virol. 87, 769–776 (2006).
Tischer, B. K., von Einem, J., Kaufer, B. & Osterrieder, N. Two-step red-mediated recombination for versatile high-efficiency markerless DNA manipulation in Escherichia coli. BioTechniques 40, 191–197 (2006).
Wu, S. Y. & Chiang, C. M. Expression and purification of epitope-tagged multisubunit protein complexesfrom mammalian cells. Curr. Protoc. Mol. Biol. Chapter 16, Unit 16.13 (2002).
Kühn, F. J. & Knopf, C. W. Herpes simplex virus type 1 DNA polymerase. Mutational analysis of the 3′−5′-exonuclease domain. J. Biol. Chem. 271, 29245–29254 (1996).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Garrison, E. & Marth, G. Haplotype-based variant detection from short-read sequencing. Preprint at https://arxiv.org/pdf/1207.3907.pdf (2012).
R Core Team. R: a language and environment for statistical computing. R package version 3.1.0 (R Foundation for Statistical Computing, 2014).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Huson, D. H. & Scornavacca, C. Dendroscope 3: an interactive tool for rooted phylogenetic trees and networks. Syst. Biol. 61, 1061–1067 (2012).
Zagordi, O., Däumer, M., Beisel, C. & Beerenwinkel, N. Read length versus depth of coverage for viral quasispecies reconstruction. PLoS ONE 7, e47046 (2012).
Acknowledgments
We thank the animal caretakers of the Robert-von-Ostertag-Haus for their expert assistance. J.T. was supported by a stipend from the Studienstiftung des Deutschen Volkes.
Author information
Authors and Affiliations
Contributions
J.T. performed and analysed the virological, genetic, biochemical, sequencing and animal experiments. N.G. performed the sequence analyses. S.H. and D.P.M. performed the phylogenetic analyses. K.E. and N.O. determined some of the virus growth properties. D.P.M., D.K. and N.O. supervised the study. J.T., D.P.M. and N.O. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–5, Supplementary Tables 1–12.
Rights and permissions
About this article
Cite this article
Trimpert, J., Groenke, N., Kunec, D. et al. A proofreading-impaired herpesvirus generates populations with quasispecies-like structure. Nat Microbiol 4, 2175–2183 (2019). https://doi.org/10.1038/s41564-019-0547-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0547-x
This article is cited by
-
Ribozyme Mutagenic Evolution: Mechanisms of Survival
Origins of Life and Evolution of Biospheres (2021)
-
Herpesvirus DNA Polymerase Mutants—How Important Is Faithful Genome Replication?
Current Clinical Microbiology Reports (2019)