Abstract
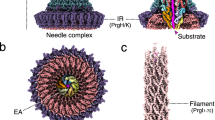
The bacterial injectisome is a syringe-shaped macromolecular nanomachine utilized by many pathogenic Gram-negative bacteria, including the causative agents of plague, typhoid fever, whooping cough, sexually transmitted infections and major nosocomial infections. Bacterial proteins destined for self-assembly and host-cell targeting are translocated by the injectisome in a process known as type III secretion (T3S). The core structure is the ~4 MDa needle complex (NC), built on a foundation of three highly oligomerized ring-forming proteins that create a hollow scaffold spanning the bacterial inner membrane (IM) (24-mer ring-forming proteins PrgH and PrgK in the Salmonella enterica serovar Typhimurium Salmonella pathogenicity island 1 (SPI-1) type III secretion system (T3SS)) and outer membrane (OM) (15-mer InvG, a member of the broadly conserved secretin pore family). An internalized helical needle projects from the NC and bacterium, ultimately forming a continuous passage to the host, for delivery of virulence effectors. Here, we have captured snapshots of the entire prototypical SPI-1 NC in four distinct needle assembly states, including near-atomic resolution, and local reconstructions in the absence and presence of the needle. These structures reveal the precise localization and molecular interactions of the internalized SpaPQR ‘export apparatus’ complex, which is intimately encapsulated and stabilized within the IM rings in the manner of a nanodisc, and to which the PrgJ rod directly binds and functions as an initiator and anchor of needle polymerization. We also describe the molecular details of the extensive and continuous coupling interface between the OM secretin and IM rings, which is remarkably facilitated by a localized 16-mer stoichiometry in the periplasmic-most coupling domain of the otherwise 15-mer InvG oligomer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request. Cryo-EM maps and atomic coordinates have been deposited with the EMDB and PDB with the following accession codes EMDB ID: EMD-20310, EMD-20311, EMD-20312, EMD-20313, EMD-20314, EMD-20315, EMD-20316, EMD-20317, EMD-20556 and PDB ID: 6PEE, 6PEM, 6PEP, 6Q14, 6Q15 and 6Q16.
References
Abrusci, P. et al. Architecture of the major component of the type III secretion system export apparatus. Nat. Struct. Mol. Biol. 20, 99–104 (2013).
Worrall, L. J. et al. Near-atomic-resolution cryo-EM analysis of the Salmonella T3S injectisome basal body. Nature 540, 597–601 (2016).
Wagner, S. et al. Organization and coordinated assembly of the type III secretion export apparatus. Proc. Natl Acad. Sci. USA 107, 17745–17750 (2010).
Schraidt, O. & Marlovits, T. C. Three-dimensional model of Salmonella’s needle complex at subnanometer resolution. Science 331, 1192–1195 (2011).
Zarivach, R., Vuckovic, M., Deng, W., Finlay, B. B. & Strynadka, N. C. J. Structural analysis of a prototypical ATPase from the type III secretion system. Nat. Struct. Mol. Biol. 14, 131–137 (2007).
Yip, C. K. et al. Structural characterization of the molecular platform for type III secretion system assembly. Nature 435, 702–707 (2005).
Spreter, T. et al. A conserved structural motif mediates formation of the periplasmic rings in the type III secretion system. Nat. Struct. Mol. Biol. 16, 468–476 (2009).
Bergeron, J. R. C. et al. A refined model of the prototypical Salmonella SPI-1 T3SS basal body reveals the molecular basis for its assembly. PLoS Pathog. 9, e1003307 (2013).
Bergeron, J. R. C. et al. The modular structure of the inner-membrane ring component PrgK facilitates assembly of the type III secretion system basal body. Structure 23, 161–172 (2015).
Majewski, D. D., Worrall, L. J. & Strynadka, N. C. Secretins revealed: structural insights into the giant gated outer membrane portals of bacteria. Curr. Opin. Struct. Biol. 51, 61–72 (2018).
Dietsche, T. et al. Structural and functional characterization of the bacterial type III secretion export apparatus. PLoS Pathog. 12, 1–25 (2016).
Kuhlen, L. et al. Structure of the core of the type III secretion system export apparatus. Nat. Struct. Mol. Biol. 25, 583–590 (2018).
Zilkenat, S. et al. Determination of the stoichiometry of the complete bacterial type III secretion needle complex using a combined quantitative proteomic approach. Mol. Cell. Proteom. 15, 1598–1609 (2016).
Hu, B., Lara-Tejero, M., Kong, Q., Galán, J. E. & Liu, J. In situ molecular architecture of the Salmonella Type III secretion machine. Cell 168, 1065–1074 (2017).
Hu, J. et al. Cryo-EM analysis of the T3S injectisome reveals the structure of the needle and open secretin. Nat. Commun. 9, 1–11 (2018).
Bai, X., Rajendra, E., Yang, G., Shi, Y. & Scheres, S. H. W. Sampling the conformational space of the catalytic subunit of human γ-secretase. eLife 4, e11182 (2015).
Schraidt, O. et al. Topology and organization of the Salmonella typhimurium type III secretion needle complex components. PLoS Pathog. 6, e1000824 (2010).
Sanowar, S. et al. Interactions of the transmembrane polymeric rings of the Salmonella enterica serovar Typhimurium type III secretion system. mBio 1, 1–8 (2010).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Erhardt, M. et al. Mechanism of type-III protein secretion: regulation of FlhA conformation by a functionally critical charged-residue cluster. Mol. Microbiol. 104, 234–249 (2017).
Ward, E. et al. Type-III secretion pore formed by flagellar protein FliP. Mol. Microbiol. 107, 94–103 (2018).
Loquet, A. et al. Atomic model of the type III secretion system needle. Nature 486, 276–279 (2012).
Radics, J., Königsmaier, L. & Marlovits, T. C. Structure of a pathogenic type 3 secretion system in action. Nat. Struct. Mol. Biol. 21, 82–87 (2014).
Marlovits, T. C. et al. Structural insights into the assembly of the type III secretion needle complex. Science 306, 1040–1042 (2004).
Sal-man, N., Deng, W. & Finlay, B. B. EscI: a crucial component of the type III secretion system forms the inner rod structure in enteropathogenic Escherichia coli. Biochem J. 125, 119–125 (2012).
Zhong, D. et al. The Salmonella type III secretion system inner rod protein PrgJ is partially folded. J. Biol. Chem. 287, 25303–25311 (2012).
Lefebre, M. D. & Galan, J. E. The inner rod protein controls substrate switching and needle length in a Salmonella type III secretion system. Proc. Natl Acad. Sci. USA 111, 817–822 (2014).
El Hajjami, N. et al. The inner-rod component of Shigella flexneri type 3 secretion system, MxiI, is involved in the transmission of the secretion activation signal by its interaction with MxiC. Microbiologyopen 7, 1–11 (2018).
Wee, D. H. & Hughes, K. T. Molecular ruler determines needle length for the Salmonella Spi-1 injectisome. Proc. Natl Acad. Sci. USA 2015, 1–6 (2015).
Diepold, A., Amstutz, M., Sorg, I., Jenal, U. & Cornelis, G. R. Deciphering the assembly of the Yersinia type III secretion injectisome. EMBO J. 29, 1928–1940 (2010).
Diepold, A. & Wagner, S. Assembly of the bacterial type III secretion machinery. FEMS Microbiol. Rev. 38, 802–822 (2014).
Berger, C., Robin, G. P., Bonas, U. & Koebnik, R. Membrane topology of conserved components of the type III secretion system from the plant pathogen Xanthomonas campestris pv. vesicatoria. Microbiology 156, 1963–1974 (2010).
van Arnam, J. S., McMurry, J. L., Kihara, M. & Macnab, R. M. Analysis of an engineered Salmonella flagellar fusion protein, FliR-FlhB. J. Bacteriol. 186, 2495–2498 (2004).
Zarivach, R. et al. Structural analysis of the essential self-cleaving type III secretion proteins EscU and SpaS. Nature 453, 124–127 (2008).
Yan, Z., Yin, M., Xu, D., Zhu, Y. & Li, X. Structural insights into the secretin translocation channel in the type II secretion system. Nat. Struct. Mol. Biol. 24, 177–183 (2017).
Hay, I. D., Belousoff, M. J. & Lithgow, T. Structural basis of type 2 secretion system engagement between the inner and outer bacterial membranes. mBio 8, 1–6 (2017).
Hu, B. et al. Visualization of the type III secretion sorting platform of Shigella flexneri. Proc. Natl Acad. Sci. USA 112, 1047–1052 (2015).
Kowal, J. et al. Structure of the dodecameric Yersinia enterocolitica secretin YscC and its trypsin-resistant core. Structure 21, 2152–2161 (2013).
Koo, J., Burrows, L. L. & Lynne Howell, P. Decoding the roles of pilotins and accessory proteins in secretin escort services. FEMS Microbiol. Lett. 328, 1–12 (2012).
Chernyatina, A. & Harry, L. Architecture of a bacterial type II secretion system. Preprint at https://www.biorxiv.org/content/10.1101/397794 (2018).
Guilvout, I. et al. Independent domain assembly in a trapped folding intermediate of multimeric outer membrane secretins. Structure 22, 582–589 (2014).
Burkinshaw, B. J. et al. Structural analysis of a specialized type III secretion system peptidoglycan-cleaving enzyme. J. Biol. Chem. 290, 10406–10417 (2015).
Marlovits, T. C. et al. Assembly of the inner rod determines needle length in the type III secretion injectisome. Nature 441, 637–640 (2006).
Wagner, S., Stenta, M., Metzger, L. C., Dal, M. & Cornelis, G. R. Length control of the injectisome needle requires only one molecule of Yop secretion protein P (YscP). Proc. Natl Acad. Sci. USA 107, 13860–13865 (2010).
Erhardt, M., Singer, H. M., Wee, D. H., Keener, J. P. & Hughes, K. T. An infrequent molecular ruler controls flagellar hook length in Salmonella enterica. EMBO J. 30, 2948–2961 (2011).
Kenjale, R. et al. The needle component of the type III secreton of Shigella regulates the activity of the secretion apparatus. J. Biol. Chem. 280, 42929–42937 (2005).
Deng, W. et al. Assembly, structure, function and regulation of type III secretion systems. Nat. Rev. Microbiol. 15, 323–337 (2017).
Kimbrough, T. G. & Miller, S. I. Contribution of Salmonella typhimurium type III secretion components to needle complex formation. Proc. Natl Acad. Sci. USA 97, 11008–11013 (2000).
Eichelberg, K. & Galán, J. E. Differential regulation of Salmonella typhimurium type III secreted proteins by pathogenicity island 1 (SPI-1)-encoded transcriptional activators InvF and HilA. Infect. Immun. 67, 4099–4105 (1999).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, 1–22 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. CryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. Sect. D 66, 213–221 (2010).
Wang, R. Y. R. et al. Automated structure refinement of macromolecular assemblies from cryo-EM maps using Rosetta. eLife 5, 1–22 (2016).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. Sect. D 66, 486–501 (2010).
Barad, B. A. et al. EMRinger: side chain-directed model and map validation for 3D cryo-electron microscopy. Nat. Methods 12, 943–946 (2015).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. Sect. D 66, 12–21 (2010).
The PyMOL Molecular Graphics System v.2.0 (Schrödinger, LLC, 2017).
Zimmermann, L. et al. A completely reimplemented MPI bioinformatics toolkit with a new HHpred server at its core. J. Mol. Biol. 430, 2237–2243 (2018).
Edwards, R. A., Keller, L. H. & Schi, D. M. Improved allelic exchange vectors and their use to analyze 987P fimbria gene expression. Gene 207, 149–157 (1998).
Ferrie, L. et al. Silent mischief: bacteriophage Mu insertions contaminate products of Escherichia coli random mutagenesis performed using suicidal transposon delivery plasmids mobilized by broad-host-range RP4 conjugative machinery. J. Bacteriol. 192, 6418–6427 (2010).
Acknowledgements
We thank S. Miller for providing S. Typhimurium deletion strains and plasmids, as well as the InvG antibody. This work was funded by operating grants from CIHR, to N.C.J.S. and B.B.F. and the Howard Hughes International Senior Scholar Program, to N.C.J.S. B.B.F. is the UBC Peter Wall Distinguished Professor. N.C.J.S. is a Tier I Canada Research Chair in Antibiotic Discovery.
Author information
Authors and Affiliations
Contributions
M.V. performed all cloning and protein purification. C.H. performed single-particle cryo-EM grid preparation and data collection with assistance from Z.Y. J.H. performed data processing and map generation with assistance from L.J.W. and C.E.A. L.J.W. performed model building, refinement and structural analysis with help from J.H. W.D. made the Salmonella deletion mutants and expression strains and carried out secretion assays with assistance from B.B.F. L.J.W., J.H. and N.C.J.S. wrote the manuscript with input from all.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Table 1, Supplementary Figs. 1–10, Supplementary Video Legends and Supplementary References.
Supplementary Video 1
Allosteric step of InvG periplasmic gate opening.
Supplementary Video 2
Steric step of InvG periplasmic gate opening.
Rights and permissions
About this article
Cite this article
Hu, J., Worrall, L.J., Vuckovic, M. et al. T3S injectisome needle complex structures in four distinct states reveal the basis of membrane coupling and assembly. Nat Microbiol 4, 2010–2019 (2019). https://doi.org/10.1038/s41564-019-0545-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0545-z
This article is cited by
-
Substrate-engaged type III secretion system structures reveal gating mechanism for unfolded protein translocation
Nature Communications (2021)
-
CryoEM structure of the outer membrane secretin channel pIV from the f1 filamentous bacteriophage
Nature Communications (2021)
-
Molecular structure of the intact bacterial flagellar basal body
Nature Microbiology (2021)
-
Essential role of Salmonella Enteritidis DNA adenine methylase in modulating inflammasome activation
BMC Microbiology (2020)