Abstract
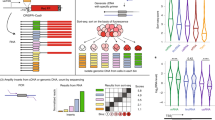
The ability to determine full-length nucleotide composition of individual RNA molecules is essential for understanding the architecture and function of a transcriptome. However, experimental approaches capable of capturing the sequences of both 5′ and 3′ termini of the same transcript remain scarce. In the present study, simultaneous 5′ and 3′ end sequencing (SEnd-seq)—a high-throughput and unbiased method that simultaneously maps transcription start and termination sites with single-nucleotide resolution—is presented. Using this method, a comprehensive view of the Escherichia coli transcriptome was obtained, which displays an unexpected level of complexity. SEnd-seq notably expands the catalogue of transcription start sites and termination sites, defines unique transcription units and detects prevalent antisense RNA. Strikingly, the results of the present study unveil widespread overlapping bidirectional terminators located between opposing gene pairs. Furthermore, it has been shown that convergent transcription is a major contributor to highly efficient bidirectional termination both in vitro and in vivo. This finding highlights an underappreciated role of RNA polymerase conflicts in shaping transcript boundaries and suggests an evolutionary strategy for modulating transcriptional output by arranging gene orientation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
SEnd-seq and standard RNA-seq datasets from this study have been deposited in the Gene Expression Omnibus (GEO) with the accession number GSE117737.
Code availability
The custom scripts used in this study are available on Github (https://github.com/LiuLab-codes/SEnd_seq_analysis). Other data that support the findings of this study are available from the corresponding author upon request.
References
Morris, K. V. & Mattick, J. S. The rise of regulatory RNA. Nat. Rev. Genet. 15, 423–437 (2014).
Wang, Z., Gerstein, M. & Snyder, M. RNA-Seq: a revolutionary tool for transcriptomics. Nat. Rev. Genet. 10, 57–63 (2009).
Sharma, C. M. et al. The primary transcriptome of the major human pathogen Helicobacter pylori. Nature 464, 250–255 (2010).
Wurtzel, O. et al. A single-base resolution map of an archaeal transcriptome. Genome Res. 20, 133–141 (2010).
Dar, D. et al. Term-seq reveals abundant ribo-regulation of antibiotics resistance in bacteria. Science 352, aad9822 (2016).
Babski, J. et al. Genome-wide identification of transcriptional start sites in the haloarchaeon Haloferax volcanii based on differential RNA-Seq (dRNA-Seq). BMC Genom. 17, 629 (2016).
Lalanne, J. B. et al. Evolutionary convergence of pathway-specific enzyme expression stoichiometry. Cell 173, 749–761 (2018).
Ettwiller, L., Buswell, J., Yigit, E. & Schildkraut, I. A novel enrichment strategy reveals unprecedented number of novel transcription start sites at single base resolution in a model prokaryote and the gut microbiome. BMC Genom. 17, 199 (2016).
Matteau, D. & Rodrigue, S. Precise identification of genome-wide transcription start sites in bacteria by 5′-rapid amplification of cDNA ends (5′-RACE). Methods Mol. Biol. 1334, 143–159 (2015).
Goodwin, S., McPherson, J. D. & McCombie, W. R. Coming of age: ten years of next-generation sequencing technologies. Nat. Rev. Genet. 17, 333–351 (2016).
Hor, J., Gorski, S. A. & Vogel, J. Bacterial RNA biology on a genome scale. Mol. Cell 70, 785–799 (2018).
Guell, M., Yus, E., Lluch-Senar, M. & Serrano, L. Bacterial transcriptomics: what is beyond the RNA horiz-ome? Nat. Rev. Microbiol. 9, 658–669 (2011).
Gama-Castro, S. et al. RegulonDB version 9.0: high-level integration of gene regulation, coexpression, motif clustering and beyond. Nucleic Acids Res. 44, D133–D143 (2016).
Ruan, X. & Ruan, Y. Genome wide full-length transcript analysis using 5′ and 3′ paired-end-tag next generation sequencing (RNA-PET). Methods Mol. Biol. 809, 535–562 (2012).
Pelechano, V., Wei, W. & Steinmetz, L. M. Extensive transcriptional heterogeneity revealed by isoform profiling. Nature 497, 127–131 (2013).
Lama, L. & Ryan, K. Adenylylation of small RNA sequencing adapters using the TS2126 RNA ligase I. RNA 22, 155–161 (2016).
Lin-Chao, S., Wei, C. L. & Lin, Y. T. RNase E is required for the maturation of ssrA RNA and normal ssrA RNA peptide-tagging activity. Proc. Natl Acad. Sci. USA 96, 12406–12411 (1999).
Ruff, E. F., Record, M. T. Jr. & Artsimovitch, I. Initial events in bacterial transcription initiation. Biomolecules 5, 1035–1062 (2015).
Conway, T. et al. Unprecedented high-resolution view of bacterial operon architecture revealed by RNA sequencing. mBio 5, e01442–14 (2014).
Yan, B., Boitano, M., Clark, T. A. & Ettwiller, L. SMRT-Cappable-seq reveals complex operon variants in bacteria. Nat. Commun. 9, 3676 (2018).
Ray-Soni, A., Bellecourt, M. J. & Landick, R. Mechanisms of bacterial transcription termination: all good things must end. Annu. Rev. Biochem 85, 319–347 (2016).
Hui, M. P., Foley, P. L. & Belasco, J. G. Messenger RNA degradation in bacterial cells. Annu Rev. Genet. 48, 537–559 (2014).
Dar, D. & Sorek, R. High-resolution RNA 3′-ends mapping of bacterial Rho-dependent transcripts. Nucleic Acids Res. 46, 6797–6805 (2018).
Zwiefka, A., Kohn, H. & Widger, W. R. Transcription termination factor rho: the site of bicyclomycin inhibition in Escherichia coli. Biochemistry 32, 3564–3570 (1993).
Chen, Y. J. et al. Characterization of 582 natural and synthetic terminators and quantification of their design constraints. Nat. Methods 10, 659–664 (2013).
Wang, X. et al. Processing generates 3′ ends of RNA masking transcription termination events in prokaryotes. Proc. Natl Acad. Sci. USA 116, 4440–4445 (2019).
Goliger, J. A., Yang, X. J., Guo, H. C. & Roberts, J. W. Early transcribed sequences affect termination efficiency of Escherichia coli RNA polymerase. J. Mol. Biol. 205, 331–341 (1989).
Telesnitsky, A. P. & Chamberlin, M. J. Sequences linked to prokaryotic promoters can affect the efficiency of downstream termination sites. J. Mol. Biol. 205, 315–330 (1989).
Thomason, M. K. et al. Global transcriptional start site mapping using differential RNA sequencing reveals novel antisense RNAs in Escherichia coli. J. Bacteriol. 197, 18–28 (2015).
Dornenburg, J. E., Devita, A. M., Palumbo, M. J. & Wade, J. T. Concerns about recently identified widespread antisense transcription in Escherichia coli. mBio 1, e00106–10 (2010).
Peters, J. M., Vangeloff, A. D. & Landick, R. Bacterial transcription terminators: the RNA 3′-end chronicles. J. Mol. Biol. 412, 793–813 (2011).
Dar, D. & Sorek, R. Extensive reshaping of bacterial operons by programmed mRNA decay. PLoS Genet. 14, e1007354 (2018).
Mondal, S., Yakhnin, A. V., Sebastian, A., Albert, I. & Babitzke, P. NusA-dependent transcription termination prevents misregulation of global gene expression. Nat. Microbiol. 1, 15007 (2016).
Zhang, J. & Landick, R. A two-way street: regulatory Interplay between RNA polymerase and nascent RNA structure. Trends Biochem. Sci. 41, 293–310 (2016).
Zhao, C., Liu, F. & Pyle, A. M. An ultraprocessive, accurate reverse transcriptase encoded by a metazoan group II intron. RNA 24, 183–195 (2018).
Callen, B. P., Shearwin, K. E. & Egan, J. B. Transcriptional interference between convergent promoters caused by elongation over the promoter. Mol. Cell 14, 647–656 (2004).
Horowitz, H. & Platt, T. Regulation of transcription from tandem and convergent promoters. Nucleic Acids Res. 10, 5447–5465 (1982).
Elledge, S. J. & Davis, R. W. Position and density effects on repression by stationary and mobile DNA-binding proteins. Genes Dev. 3, 185–197 (1989).
Shearwin, K. E., Callen, B. P. & Egan, J. B. Transcriptional interference—a crash course. Trends Genet. 21, 339–345 (2005).
Sameshima, J. H., Wek, R. C. & Hatfield, G. W. Overlapping transcription and termination of the convergent ilvA and ilvY genes of Escherichia coli. J. Biol. Chem. 264, 1224–1231 (1989).
Postle, K. & Good, R. F. A bidirectional rho-independent transcription terminator between the E. coli tonB gene and an opposing gene. Cell 41, 577–585 (1985).
Sneppen, K. et al. A mathematical model for transcriptional interference by RNA polymerase traffic in Escherichia coli. J. Mol. Biol. 346, 399–409 (2005).
Brantl, S. & Wagner, E. G. An antisense RNA-mediated transcriptional attenuation mechanism functions in Escherichia coli. J. Bacteriol. 184, 2740–2747 (2002).
Landick, R., Carey, J. & Yanofsky, C. Translation activates the paused transcription complex and restores transcription of the trp operon leader region. Proc. Natl Acad. Sci. USA 82, 4663–4667 (1985).
Proshkin, S., Rahmouni, A. R., Mironov, A. & Nudler, E. Cooperation between translating ribosomes and RNA polymerase in transcription elongation. Science 328, 504–508 (2010).
Ma, J., Bai, L. & Wang, M. D. Transcription under torsion. Science 340, 1580–1583 (2013).
Crampton, N., Bonass, W. A., Kirkham, J., Rivetti, C. & Thomson, N. H. Collision events between RNA polymerases in convergent transcription studied by atomic force microscopy. Nucleic Acids Res. 34, 5416–5425 (2006).
Hobson, D. J., Wei, W., Steinmetz, L. M. & Svejstrup, J. Q. RNA polymerase II collision interrupts convergent transcription. Mol. Cell 48, 365–374 (2012).
Prescott, E. M. & Proudfoot, N. J. Transcriptional collision between convergent genes in budding yeast. Proc. Natl Acad. Sci. USA 99, 8796–8801 (2002).
Eszterhas, S. K., Bouhassira, E. E., Martin, D. I. & Fiering, S. Transcriptional interference by independently regulated genes occurs in any relative arrangement of the genes and is influenced by chromosomal integration position. Mol. Cell Biol. 22, 469–479 (2002).
Creecy, J. P. & Conway, T. Quantitative bacterial transcriptomics with RNA-seq. Curr. Opin. Microbiol. 23, 133–140 (2015).
Jensen, S. I., Lennen, R. M., Herrgard, M. J. & Nielsen, A. T. Seven gene deletions in seven days: fast generation of Escherichia coli strains tolerant to acetate and osmotic stress. Sci. Rep. 5, 17874 (2015).
Peters, J. M. et al. Rho directs widespread termination of intragenic and stable RNA transcription. Proc. Natl Acad. Sci. USA 106, 15406–15411 (2009).
McClure, R. et al. Computational analysis of bacterial RNA-Seq data. Nucleic Acids Res. 41, e140 (2013).
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Celesnik, H., Deana, A. & Belasco, J. G. Initiation of RNA decay in Escherichia coli by 5′ pyrophosphate removal. Mol. Cell 27, 79–90 (2007).
Lorenz, R. et al. ViennaRNA Package 2.0. Algorithms Mol. Biol. 6, 26 (2011).
Crooks, G. E., Hon, G., Chandonia, J. M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Acknowledgements
We thank S. Darst and E. Campbell for help with the in vitro transcription experiments and critical reading of the manuscript, K. Ryan, A. Pyle and R. Landick for sharing reagents and E. Cheng for help with data analysis. This work was supported by a C.H. Li Memorial Scholar Fund Award (X.J.), the Robertson Foundation, the Quadrivium Foundation, a Monique Weill-Caulier Career Scientist Award, a March of Dimes Basil O’Connor Starter Scholar Award, a Kimmel Scholar Award, and National Institute of Health grants R00GM107365 and DP2HG010510 (S.L.).
Author information
Authors and Affiliations
Contributions
S.L. conceived of and oversaw the project. X.J. performed the experiments and data analysis. D.L. contributed to the development of SEnd-seq workflow. S.L. and X.J. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The Rockefeller University has filed a provisional patent application encompassing aspects of the SEnd-seq technology on which S.L. and X.J. are listed as inventors.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–23, Supplementary Tables 1–5 and Supplementary References.
Supplementary Table 1
Genomic position of TSSs identified by SEnd-seq.
Supplementary Table 2
Genomic position and predicted secondary structure of TTSs identified by SEnd-seq.
Supplementary Table 3
Transcription units defined by SEnd-seq.
Supplementary Table 4
Genomic position and structural analysis of overlapping bidirectional TTSs and the E. coli genes involved.
Rights and permissions
About this article
Cite this article
Ju, X., Li, D. & Liu, S. Full-length RNA profiling reveals pervasive bidirectional transcription terminators in bacteria. Nat Microbiol 4, 1907–1918 (2019). https://doi.org/10.1038/s41564-019-0500-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0500-z
This article is cited by
-
Incomplete transcripts dominate the Mycobacterium tuberculosis transcriptome
Nature (2024)
-
Co-transcriptional gene regulation in eukaryotes and prokaryotes
Nature Reviews Molecular Cell Biology (2024)
-
TRS: a method for determining transcript termini from RNAtag-seq sequencing data
Nature Communications (2023)
-
Improved RNA stability estimation indicates that transcriptional interference is frequent in diverse bacteria
Communications Biology (2023)
-
Extensive diversity in RNA termination and regulation revealed by transcriptome mapping for the Lyme pathogen Borrelia burgdorferi
Nature Communications (2023)