Abstract
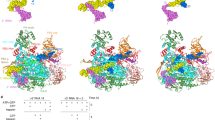
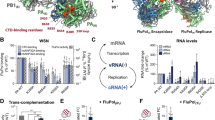
The influenza virus polymerase uses capped RNA primers to initiate transcription, and a combination of terminal and internal de novo initiations for the two-step replication process by binding the conserved viral genomic RNA (vRNA) or complementary RNA (cRNA) promoter. Here, we determined the apo and promoter-bound influenza D polymerase structures using cryo-electron microscopy and found the polymerase has an evolutionarily conserved stable core structure with inherently flexible peripheral domains. Strikingly, two conformations (mode A and B) of the vRNA promoter were observed where the 3ʹ-vRNA end can bind at two different sites, whereas the cRNA promoter only binds in the mode B conformation. Functional studies confirmed the critical role of the mode B conformation for vRNA synthesis via the intermediate cRNA but not for cRNA production, which is mainly regulated by the mode A conformation. Both conformations participate in the regulation of the transcription process. This work advances our understanding of the regulatory mechanisms for the synthesis of different RNA species by influenza virus polymerase and opens new opportunities for antiviral drug design.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The density maps have been deposited to the Electron Microscopy Data Bank under the accession numbers EMD-9577 (apo FluDPol), EMD-9578 (vRNA promoter-bound class A1), EMD-9579 (class A2), EMD-9581 (class B1), EMD-9580 (class B2), EMD-9582 (class B3), EMD-9887 (cRNA promoter-bound class 1) and EMD-9888 (class 2). The coordinates of the corresponding atomic models have been deposited to the Protein Data Bank with the entries 6AB7 (apo FluDPol), 6ABB (promoter-bound class A1), 6ABD (class A2), 6ABF (class B1), 6ABE (class B2), 6ABG (class B3), 6JU2 (cRNA promoter-bound class 1) and 6JU3 (class 2).
References
McCauley, J. W. et al. in Virus Taxonomy (eds King, A.M.Q. et al.) 749–762 (Elsevier, 2011).
Hause, B. M. et al. Isolation of a novel swine influenza virus from Oklahoma in 2011 which is distantly related to human influenza C viruses. PLoS Pathog. 9, e1003176 (2013).
Song, H. et al. An open receptor-binding cavity of hemagglutinin-esterase-fusion glycoprotein from newly-identified influenza D virus: basis for its broad cell tropism. PLoS Pathog. 12, e1005411 (2016).
Szewczyk, B., Bienkowska-Szewczyk, K. & Krol, E. Introduction to molecular biology of influenza A viruses. Acta Biochim. Pol. 61, 397–401 (2014).
Elderfield, R. & Barclay, W. Influenza pandemics. Adv. Exp. Med. Biol. 719, 81–103 (2011).
Paul Glezen, W., Schmier, J. K., Kuehn, C. M., Ryan, K. J. & Oxford, J. The burden of influenza B: a structured literature review. Am. J. Public Health 103, e43–e51 (2013).
Gao, G. F. From “A“IV to “Z“IKV: attacks from emerging and re-emerging pathogens. Cell 172, 1157–1159 (2018).
Muraki, Y. & Hongo, S. The molecular virology and reverse genetics of influenza C virus. Jpn. J. Infect. Dis. 63, 157–165 (2010).
Webster, R. G., Laver, W. G., Air, G. M. & Schild, G. C. Molecular mechanisms of variation in influenza viruses. Nature 296, 115–121 (1982).
Shi, Y., Wu, Y., Zhang, W., Qi, J. & Gao, G. F. Enabling the ‘host jump’: structural determinants of receptor-binding specificity in influenza A viruses. Nat. Rev. Microbiol. 12, 822–831 (2014).
Shen, Z., Lou, K. & Wang, W. New small-molecule drug design strategies for fighting resistant influenza A. Acta Pharm. Sin. B 5, 419–430 (2015).
Hussain, M., Galvin, H. D., Haw, T. Y., Nutsford, A. N. & Husain, M. Drug resistance in influenza A virus: the epidemiology and management. Infect. Drug Resist. 10, 121–134 (2017).
Amarelle, L., Lecuona, E. & Sznajder, J. I. Anti-influenza treatment: drugs currently used and under development. Arch. Bronconeumol. 53, 19–26 (2017).
Hurt, A. C., Ho, H. T. & Barr, I. Resistance to anti-influenza drugs: adamantanes and neuraminidase inhibitors. Expert Rev. Anti-Infective Ther. 4, 795–805 (2006).
Hatakeyama, S. & Kawaoka, Y. The molecular basis of resistance to anti-influenza drugs (in Japanese). Nihon Rinsho 64, 1845–1852 (2006).
Furuta, Y. et al. In vitro and in vivo activities of anti-influenza virus compound T-705. Antimicrob. Agents Chemother. 46, 977–981 (2002).
Stevaert, A. & Naesens, L. The influenza virus polymerase complex: an update on its structure, functions, and significance for antiviral drug design. Med. Res. Rev. 36, 1127–1173 (2016).
Mikulasova, A., Vareckova, E. & Fodor, E. Transcription and replication of the influenza A virus genome. Acta Virol. 44, 273–282 (2000).
Te Velthuis, A. J., Robb, N. C., Kapanidis, A. N. & Fodor, E. The role of the priming loop in influenza A virus RNA synthesis. Nat. Microbiol. 1, 16029 (2016).
Deng, T., Vreede, F. T. & Brownlee, G. G. Different de novo initiation strategies are used by influenza virus RNA polymerase on its cRNA and viral RNA promoters during viral RNA replication. J. Virol. 80, 2337–2348 (2006).
Pflug, A., Lukarska, M., Resa-Infante, P., Reich, S. & Cusack, S. Structural insights into RNA synthesis by the influenza virus transcription-replication machine. Virus Res. 234, 103–117 (2017).
Pflug, A., Guilligay, D., Reich, S. & Cusack, S. Structure of influenza A polymerase bound to the viral RNA promoter. Nature 516, 355–360 (2014).
Chang, S. et al. Cryo-EM structure of influenza virus RNA polymerase complex at 4.3 Å resolution. Mol. Cell 57, 925–935 (2015).
Hengrung, N. et al. Crystal structure of the RNA-dependent RNA polymerase from influenza C virus. Nature 527, 114–117 (2015).
Reich, S. et al. Structural insight into cap-snatching and RNA synthesis by influenza polymerase. Nature 516, 361–366 (2014).
Thierry, E. et al. Influenza polymerase can adopt an alternative configuration involving a radical repacking of PB2 domains. Mol. Cell 61, 125–137 (2016).
Serna Martin, I. et al. A mechanism for the activation of the influenza virus transcriptase. Mol. Cell 70, 1101–1110 (2018).
Te Velthuis, A. J. W. & Oymans, J. Initiation, elongation and realignment during influenza virus mRNA synthesis. J. Virol. 92, 1–12 (2017).
Gerlach, P., Malet, H., Cusack, S. & Reguera, J. Structural insights into Bunyavirus replication and its regulation by the vRNA promoter. Cell 161, 1267–1279 (2015).
Reich, S., Guilligay, D. & Cusack, S. An in vitro fluorescence based study of initiation of RNA synthesis by influenza B polymerase. Nucleic Acids Res. 45, 3353–3368 (2017).
Kumar, N., Xin, Z. T., Liang, Y. H., Ly, H. & Liang, Y. Y. NF-κB signaling differentially regulates influenza virus RNA synthesis. J. Virol. 82, 9880–9889 (2008).
Sugiyama, K., Kawaguchi, A., Okuwaki, M. & Nagata, K. pp32 and APRIL are host cell-derived regulators of influenza virus RNA synthesis from cRNA. eLife 4, 1–19 (2015).
York, A., Hengrung, N., Vreede, F. T., Huiskonen, J. T. & Fodor, E. Isolation and characterization of the positive-sense replicative intermediate of a negative-strand RNA virus. Proc. Natl Acad. Sci. USA 110, E4238–E4245 (2013).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Kimanius, D., Forsberg, B. O., Scheres, S. H. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, e18722 (2016).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Acknowledgements
We thank all staff at the National Center for Protein Science Shanghai cryo-EM department and the Center of Biological Imaging, Institute of Biophysics for assistance with data collection. We are grateful to G. Wang (The Core Facilities at School of Life Sciences, Peking University), T. Yang and the staff in the EM department of the State Key Laboratory of Membrane Biology, Institute of Zoology, CAS for their technical support in the operation of the electron microscope. The ForteBio Octet experiment was supported by the Research Facility Center at Beijing Institutes of Life Science, CAS. This study was supported by the Strategic Priority Research Program of the CAS (grant no. XDB29010000), the National Science and Technology Major Project (grant no. 2018ZX10101004) and the External Cooperation Program of the CAS (grant no. 153211KYSB20160001). R.P. was supported by the Young Elite Scientist Sponsorship Program from the China Association for Science and Technology (grant no. 2018QNRC001). M.W. and J.Y. were also supported by the National Science and Technology Major Project (grant no. 2018ZX09711003). G.F.G. was partly supported as a leading principal investigator of the NSFC Innovative Research Group (grant no. 81621091). Y.S. was supported by the Excellent Young Scientist Program from the National Natural Science Foundation of China (grant no. 81622031), the Excellent Young Scientist Program of the CAS and the Youth Innovation Promotion Association CAS (grant no. 2015078).
Author information
Authors and Affiliations
Contributions
Y.S., R.P. and G.F.G. designed the project. Q.P. and Y.L. purified the protein samples and conducted the biochemical studies. Q.P., R.P. and S.L. prepared the cryo-EM samples and collected the data. R.P. and X.Z. performed image processing and reconstruction. R.P. and J.Q. built the atomic models. Q.P., R.P., M.W. and Y.S. analysed the structure. Q.P., Y.L., Y.C., H.S. and M.H. conducted the radioactive-labelled polymerase activity assays. Q.P., M.W., W.Y. and T.D. performed the replicon-based polymerase activity assays. Y.S., R.P. and G.F.G. wrote the paper. M.W., M.H., H.S., T.D., P.W., J.Y. and B.Z. revised the manuscript and were involved in intensive discussions of the data. Y.S. supervised all of the research.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–18 and Supplementary Tables 1–3.
Rights and permissions
About this article
Cite this article
Peng, Q., Liu, Y., Peng, R. et al. Structural insight into RNA synthesis by influenza D polymerase. Nat Microbiol 4, 1750–1759 (2019). https://doi.org/10.1038/s41564-019-0487-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0487-5
This article is cited by
-
Generation of a recombinant temperature-sensitive influenza D virus
Scientific Reports (2023)
-
Molecular mechanism of de novo replication by the Ebola virus polymerase
Nature (2023)
-
An intermediate state allows influenza polymerase to switch smoothly between transcription and replication cycles
Nature Structural & Molecular Biology (2023)
-
Structures of active Hantaan virus polymerase uncover the mechanisms of Hantaviridae genome replication
Nature Communications (2023)
-
Structural and functional analysis of the minimal orthomyxovirus-like polymerase of Tilapia Lake Virus from the highly diverged Amnoonviridae family
Nature Communications (2023)