Abstract
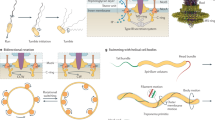
We describe Idionectes vortex gen. nov., sp. nov., a unicellular microeukaryote that swims by continuous inversion of its surface, similar to a vortex ring. This previously unreported mode of motility approximates a hypothetical concept called the ‘toroidal swimmer’, in which a doughnut-shaped object rotates around its circular axis and travels in the opposite direction to its outer surface motion. During swimming, the flagellum of Idionectes rotates relative to its cell body, which is normally a hallmark of prokaryotic rather than eukaryotic flagella.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The SSU ribosomal RNA gene sequence of I. vortex is available in GenBank under accession number MK736991, and transcriptomic RNA sequence read data are available for download from the NCBI Sequence Read Archive database under BioProject accession PRJNA531640.
Code availability
The custom code used for the phylogenomic analyses is available from the authors upon request.
References
Cavalier-Smith, T., Chao, E. E. & Lewis, R. Mol. Phylogenet. Evol. 99, 275–296 (2016).
Kang, S. et al. Mol. Biol. Evol. 34, 2258–2270 (2017).
Kudryavtsev, A. & Pawlowski, J. Protist 164, 13–23 (2013).
Lahr, D. J., Grant, J., Molestina, R., Katz, L. A. & Anderson, O. R. J. Eukaryot. Microbiol. 62, 444–453 (2015).
Brennen, C. & Winet, H. Annu. Rev. Fluid Mech. 9, 339–398 (1977).
Sleigh, M. A. Protoplasma 164, 45–53 (1991).
Purcell, E. M. Am. J. Phys. 45, 3–11 (1977).
Taylor, G. I. Proc. R. Soc. Lond. A 211, 225–239 (1952).
Thaokar, R. M., Schiessel, H. & Kulic, I. M. Eur. Phys. J. B 60, 325–336 (2007).
Leshansky, A. M. & Kenneth, O. Phys. Fluids 20, 063104 (2008).
Huang, J. & Fauci, L. Phys. Rev. E 95, 043102 (2017).
Sleigh, M. A. Comp. Biochem. Physiol. 94, 359–364 (1989).
Day, M. A. Erkenntnis 33, 285–296 (1990).
Berg, H. C. & Anderson, R. A. Nature 245, 380–382 (1973).
Yubuki, N. & Leander, B. S. Plant J. 75, 230–244 (2013).
Walker, G., Simpson, A. G., Edgcomb, V., Sogin, M. L. & Patterson, D. J. Eur. J. Protistol. 37, 25–49 (2001).
Singer, S. J. & Nicolson, G. L. Science 175, 720–731 (1972).
Tamm, S. L. & Tamm, S. Proc. Natl Acad. Sci. USA 71, 4589–4593 (1974).
Omoto, C. K. & Witman, G. B. Nature 290, 708–710 (1981).
Chwang, A. T. & Wu, T. Y. Proc. R. Soc. Lond. B 178, 327–346 (1971).
McFadden, G. I. & Melkonian, M. Phycologia 25, 551–557 (1986).
Sensen, C. W., Heimann, K. & Melkonian, M. Eur. J. Phycol. 28, 93–97 (1993).
Melkonian, M. & Weber, A. Z. Pflanzenphysiol. 76, 120–129 (1975).
Schindelin, J. et al. Nat. Methods 9, 676–682 (2012).
Reynolds, E. S. J. Cell Biol. 17, 208–212 (1963).
Marin, B., Palm, A., Klingberg, M. & Melkonian, M. Protist 154, 99–145 (2003).
Hultman, T., Bergh, S., Moks, T. & Uhlén, M. BioTechniques 10, 84–93 (1991).
Marin, B., Klingberg, M. & Melkonian, M. Protist 149, 265–276 (1998).
Stamatakis, A. Bioinformatics 22, 2688–2690 (2006).
Bolger, A. M., Lohse, M. & Usadel, B. Bioinformatics 30, 2114–2120 (2014).
Grabherr, M. G. et al. Nat. Biotechnol. 29, 644–652 (2011).
Harding, T., Brown, M. W., Simpson, A. G. & Roger, A. J. Genome Biol. Evol. 8, 2241–2258 (2016).
Katoh, K. & Standley, D. M. Mol. Biol. Evol. 30, 772–780 (2013).
Criscuolo, A. & Gribaldo, S. BMC Evol. Biol. 10, 210 (2010).
Nguyen, L. T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. Mol. Biol. Evol. 32, 268–274 (2015).
Minh, B. Q., Nguyen, M. A. T. & von Haeseler, A. Mol. Biol. Evol. 30, 1188–1195 (2013).
Wang, H. C., Minh, B. Q., Susko, E. & Roger, A. J. Syst. Biol. 67, 216–235 (2017).
Acknowledgements
We thank M. Kreutz and K. Hoef-Emden for providing natural samples, M. Melkonian and the CCAC for laboratory resources and algal cultures, Labor Dr. Schäffner (Solingen) for access to a field emission scanning electron microscope, and R. Goldstein, A. Rutenberg, A. Speers, A. Leshansky, G. Witman and S. Geimer for discussions and comments on the manuscript. This research was supported by the Studienstiftung des Deutschen Volkes (fellowship to S.H.) and German Research Foundation (DFG grant HE 7560/1-1 to S.H.). The work carried out in A.J.R.’s laboratory was supported by a Discovery Grant (2017-06792) from the Natural Sciences and Engineering Research Council of Canada (NSERC) awarded to A.J.R. The work conducted in A.G.B.S.’s laboratory was supported by NSERC Discovery Grant 298366-2014.
Author information
Authors and Affiliations
Contributions
S.H. performed the experiments and analysed the microscopy data. S.H. and L.E. performed the phylogenomic analyses. A.G.B.S. and A.J.R. provided the laboratory resources. S.H. and A.G.B.S. wrote the manuscript with input from all co-authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–4, Suppementary Tables 1–7, Supplementary Video legends and Supplementary References.
Supplementary Table 8
Taxa and proteins analysed in phylogenomic analyses.
Supplementary Video 1
Tumbling cells of I. vortex, imaged by DIC and shown at natural speed.
Supplementary Video 2
Smooth swimmers of I. vortex, imaged by DIC and shown at natural speed and in slow motion (1/4 natural speed).
Supplementary Video 3
Behaviour of deflected particles at the flagellum of I. vortex, imaged by DIC and shown in slow motion (1/3 natural speed).
Supplementary Video 4
Flagellar fluid layer of I. vortex during swimming, imaged by DIC and shown in slow motion (1/3 natural speed).
Supplementary Video 5
Flow fields of swimming cells visualized by motion tracking of latex microbeads, imaged by DIC with differential background subtraction and shown in slow motion (1/2 natural speed).
Supplementary Video 6
Relative flagellar rotation in I. vortex, imaged by DIC and shown at natural speed and in slow motion (1/4 natural speed).
Supplementary Video 7
Oscillating flagellar tip in rotating cells of I. vortex, imaged by DIC and shown in slow motion (1/3 natural speed).
Supplementary Video 8
Fluid flow around the static flagellum of I. vortex, imaged by DIC with motion tracking and shown at natural speed.
Rights and permissions
About this article
Cite this article
Hess, S., Eme, L., Roger, A.J. et al. A natural toroidal microswimmer with a rotary eukaryotic flagellum. Nat Microbiol 4, 1620–1626 (2019). https://doi.org/10.1038/s41564-019-0478-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0478-6
This article is cited by
-
Minimum entropy production by microswimmers with internal dissipation
Nature Communications (2023)
-
Experimental evidence for enzymatic cell wall dissolution in a microbial protoplast feeder (Orciraptor agilis, Viridiraptoridae)
BMC Biology (2022)
-
Different serotypes of Escherichia coli flagellin exert identical adjuvant effects
BMC Veterinary Research (2022)