Abstract
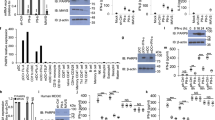
Outbreaks of viral infections are a global health burden. Although type I interferon (IFN-I) exerts broad-spectrum antiviral effects, its antiviral efficacy in host cells is largely restricted by viruses. How the antiviral efficacy of IFN-I can be improved remains to be explored. Here, we identified the ADP-ribosyltransferase poly(ADP-ribose) polymerase family member 11 (PARP11) as a potent regulator of IFN-I antiviral efficacy. PARP11 does not restrict IFN-I production induced by vesicular stomatitis virus or Sendai virus but inhibits the strength of IFN-I-activated signalling. Mechanistically, PARP11 mono-ADP-ribosylates the ubiquitin E3 ligase β-transducin repeat-containing protein (β-TrCP). Mono-ADP-ribosylation of β-TrCP promotes IFNα/β receptor subunit 1 (IFNAR1) ubiquitination and degradation. Moreover, PARP11 expression is upregulated by virus infections, including vesicular stomatitis virus, herpes simplex virus-1 and influenza A virus, thus promoting ADP-ribosylation-mediated viral evasion. We further highlight the potential for repurposing clinical ADP-ribosylation inhibitors. We found that rucaparib can target PARP11 to stabilize IFNAR1 and therefore exhibits efficient enhancement of IFN-I signalling and the host antiviral response. Consequently, rucaparib renders mice more resistant to viral infection. Our study updates the understanding of how β-TrCP regulates its substrates and may provide a druggable target for improving IFN antiviral efficacy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
References
Levy, D. E., Kessler, D. S., Pine, R., Reich, N. & Darnell, J. E. Jr. Interferon-induced nuclear factors that bind a shared promoter element correlate with positive and negative transcriptional control. Genes Dev. 2, 383–393 (1988).
Sadler, A. J. & Williams, B. R. Interferon-inducible antiviral effectors. Nat. Rev. Immunol. 8, 559–568 (2008).
Qian, J. et al. Pathogen recognition receptor signaling accelerates phosphorylation-dependent degradation of IFNAR1. PLoS Pathog. 7, e1002065 (2011).
Liu, J. et al. Virus-induced unfolded protein response attenuates antiviral defenses via phosphorylation-dependent degradation of the type I interferon receptor. Cell Host Microbe 5, 72–83 (2009).
Kumar, K. G. et al. SCF(HOS) ubiquitin ligase mediates the ligand-induced down-regulation of the interferon-α receptor. EMBO J. 22, 5480–5490 (2003).
Kumar, K. G., Krolewski, J. J. & Fuchs, S. Y. Phosphorylation and specific ubiquitin acceptor sites are required for ubiquitination and degradation of the IFNAR1 subunit of type I interferon receptor. J. Biol. Chem. 279, 46614–46620 (2004).
Zheng, H., Qian, J., Varghese, B., Baker, D. P. & Fuchs, S. Ligand-stimulated downregulation of the alpha interferon receptor: role of protein kinase D2. Mol. Cell. Biol. 31, 710–720 (2011).
Gibson, B. A. & Kraus, W. L. New insights into the molecular and cellular functions of poly(ADP-ribose) and PARPs. Nat. Rev. Mol. Cell Biol. 13, 411–424 (2012).
Schreiber, V., Dantzer, F., Ame, J. C. & de Murcia, G. Poly(ADP-ribose): novel functions for an old molecule. Nat. Rev. Mol. Cell Biol. 7, 517–528 (2006).
Audebert, M., Salles, B. & Calsou, P. Involvement of poly(ADP-ribose) polymerase-1 and XRCC1/DNA ligase III in an alternative route for DNA double-strand breaks rejoining. J. Biol. Chem. 279, 55117–55126 (2004).
Langelier, M. F., Planck, J. L., Roy, S. & Pascal, J. M. Structural basis for DNA damage-dependent poly(ADP-ribosyl)ation by human PARP-1. Science 336, 728–732 (2012).
Kraus, W. L. & Lis, J. T. PARP goes transcription. Cell 113, 677–683 (2003).
Luo, X. & Kraus, W. L. On PAR with PARP: cellular stress signaling through poly(ADP-ribose) and PARP-1. Genes Dev. 26, 417–432 (2012).
Hassa, P. O. & Hottiger, M. O. The functional role of poly(ADP-ribose)polymerase 1 as novel coactivator of NF-kappaB in inflammatory disorders. Cell. Mol. Life Sci. 59, 1534–1553 (2002).
Bai, P. et al. PARP-1 inhibition increases mitochondrial metabolism through SIRT1 activation. Cell Metab. 13, 461–468 (2011).
Pellegrino, S. & Altmeyer, M. Interplay between ubiquitin, SUMO, and poly(ADP-Ribose) in the cellular response to genotoxic stress. Front. Genet. 7, 63 (2016).
Ekblad, T., Camaioni, E., Schuler, H. & Macchiarulo, A. PARP inhibitors: polypharmacology versus selective inhibition. FEBS J. 280, 3563–3575 (2013).
Jagtap, P. & Szabo, C. Poly(ADP-ribose) polymerase and the therapeutic effects of its inhibitors. Nat. Rev. Drug Discov. 4, 421–440 (2005).
Wahlberg, E. et al. Family-wide chemical profiling and structural analysis of PARP and tankyrase inhibitors. Nat. Biotechnol. 30, 283–288 (2012).
Lin, K. Y. & Kraus, W. L. PARP inhibitors for cancer therapy. Cell 169, 183 (2017).
Kim, G. et al. FDA approval summary: olaparib monotherapy in patients with deleterious germline BRCA-mutated advanced ovarian cancer treated with three or more lines of chemotherapy. Clin. Cancer Res. 21, 4257–4261 (2015).
Balasubramaniam, S. et al. FDA approval summary: rucaparib for the treatment of patients with deleterious BRCA mutation-associated advanced ovarian cancer. Clin. Cancer Res. 23, 7165–7170 (2017).
Erener, S. et al. Inflammasome-activated caspase 7 cleaves PARP1 to enhance the expression of a subset of NF-κB target genes. Mol. Cell 46, 200–211 (2012).
Verheugd, P. et al. Regulation of NF-κB signalling by the mono-ADP-ribosyltransferase ARTD10. Nat. Commun. 4, 1683 (2013).
Barkauskaite, E., Jankevicius, G. & Ahel, I. Structures and mechanisms of enzymes employed in the synthesis and degradation of PARP-dependent protein ADP-ribosylation. Mol. Cell 58, 935–946 (2015).
Zhang, Y. et al. PARP9-DTX3L ubiquitin ligase targets host histone H2BJ and viral 3C protease to enhance interferon signaling and control viral infection. Nat. Immunol. 16, 1215–1227 (2015).
Todorova, T., Bock, F. J. & Chang, P. PARP13 regulates cellular mRNA post-transcriptionally and functions as a pro-apoptotic factor by destabilizing TRAILR4 transcript. Nat. Commun. 5, 5362 (2014).
Zheng, H. et al. A BRISC-SHMT complex deubiquitinates IFNAR1 and regulates interferon responses. Cell Rep. 5, 180–193 (2013).
Vyas, S. et al. Family-wide analysis of poly(ADP-ribose) polymerase activity. Nat. Commun. 5, 4426 (2014).
Sowa, M. E., Bennett, E. J., Gygi, S. P. & Harper, J. W. Defining the human deubiquitinating enzyme interaction landscape. Cell 138, 389–403 (2009).
Otto, H. et al. In silico characterization of the family of PARP-like poly(ADP-ribosyl)transferases (pARTs). BMC Genomics 6, 139 (2005).
Kleine, H. et al. Substrate-assisted catalysis by PARP10 limits its activity to mono-ADP-ribosylation. Mol. Cell 32, 57–69 (2008).
Iwata, H. et al. PARP9 and PARP14 cross-regulate macrophage activation via STAT1 ADP-ribosylation. Nat. Commun. 7, 12849 (2016).
Gagne, J. P. et al. Proteome-wide identification of poly(ADP-ribose) binding proteins and poly(ADP-ribose)-associated protein complexes. Nucleic Acids Res. 36, 6959–6976 (2008).
Pleschke, J. M., Kleczkowska, H. E., Strohm, M. & Althaus, F. R. Poly(ADP-ribose) binds to specific domains in DNA damage checkpoint proteins. J. Biol. Chem. 275, 40974–40980 (2000).
Rosenthal, F. et al. Macrodomain-containing proteins are new mono-ADP-ribosylhydrolases. Nat. Struct. Mol. Biol. 20, 502–507 (2013).
Kalisch, T., Ame, J. C., Dantzer, F. & Schreiber, V. New readers and interpretations of poly(ADP-ribosyl)ation. Trends Biochem. Sci. 37, 381–390 (2012).
Feijs, K. L., Forst, A. H., Verheugd, P. & Luscher, B. Macrodomain-containing proteins: regulating new intracellular functions of mono(ADP-ribosyl)ation. Nat. Rev. Mol. Cell Biol. 14, 443–451 (2013).
Galan, J. M. & Peter, M. Ubiquitin-dependent degradation of multiple F-box proteins by an autocatalytic mechanism. Proc. Natl Acad. Sci. USA 96, 9124–9129 (1999).
Zhou, P. & Howley, P. M. Ubiquitination and degradation of the substrate recognition subunits of SCF ubiquitin-protein ligases. Mol. Cell 2, 571–580 (1998).
Li, Y., Gazdoiu, S., Pan, Z. Q. & Fuchs, S. Y. Stability of homologue of Slimb F-box protein is regulated by availability of its substrate. J. Biol. Chem. 279, 11074–11080 (2004).
Harding, S. M. et al. Mitotic progression following DNA damage enables pattern recognition within micronuclei. Nature 548, 466–470 (2017).
Ghosh, R., Roy, S. & Franco, S. PARP1 depletion induces RIG-I-dependent signaling in human cancer cells. PloS ONE 13, e0194611 (2018).
Meyer-Ficca, M. L. et al. Spermatid head elongation with normal nuclear shaping requires ADP-ribosyltransferase PARP11 (ARTD11) in mice. Biol. Reprod. 92, 80 (2015).
Zhang, Y. et al. RNF146 is a poly(ADP-ribose)-directed E3 ligase that regulates axin degradation and Wnt signalling. Nat. Cell Biol. 13, 623–629 (2011).
Kang, H. C. et al. Iduna is a poly(ADP-ribose) (PAR)-dependent E3 ubiquitin ligase that regulates DNA damage. Proc. Natl Acad. Sci. USA 108, 14103–14108 (2011).
Fuchs, S. Y., Spiegelman, V. S. & Kumar, K. G. The many faces of β-TrCP E3 ubiquitin ligases: reflections in the magic mirror of cancer. Oncogene 23, 2028–2036 (2004).
Acknowledgements
We thank S. Y. Fuchs at the University of Pennsylvania for valuable discussions, and E. Y. Chinn for the critical reading of this manuscript. We also thank X. Zhu for technical support. This work was supported by the National Natural Science Foundation of China (31570865 and 31770177 for H.Z., and 31501139 for T.G.), and the program of 1000 Young Talents (2014).
Author information
Authors and Affiliations
Contributions
H.Z. conceived the study. H.Z., T.G., S.X., and C.D. designed the experiments. T.G., Y.Z., L.Q. and J.L. performed most of the experiments and analysed the data. Y.Y., Y.M. and Q.F. assisted with the RNAi screening, transfection and lentivirus experiments, and tissue processing; K.X. assisted with the PCR analyses; X.C. and L.Z. assisted with the mouse experiments, tissue processing and analyses; L.J. assisted with the RT-qPCR; H.Z supervised the project; H.Z. and T.G. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–9.
Rights and permissions
About this article
Cite this article
Guo, T., Zuo, Y., Qian, L. et al. ADP-ribosyltransferase PARP11 modulates the interferon antiviral response by mono-ADP-ribosylating the ubiquitin E3 ligase β-TrCP. Nat Microbiol 4, 1872–1884 (2019). https://doi.org/10.1038/s41564-019-0428-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0428-3
This article is cited by
-
PARP11 interfer(on)es with CAR T cell efficacy
Nature Cancer (2022)
-
Depression compromises antiviral innate immunity via the AVP-AHI1-Tyk2 axis
Cell Research (2022)
-
Intracellular mono-ADP-ribosyltransferases at the host–virus interphase
Cellular and Molecular Life Sciences (2022)
-
Targeting PARP11 to avert immunosuppression and improve CAR T therapy in solid tumors
Nature Cancer (2022)
-
ADP-ribosyltransferase PARP11 suppresses Zika virus in synergy with PARP12
Cell & Bioscience (2021)