Abstract
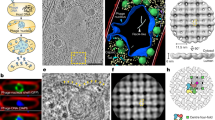
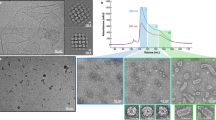
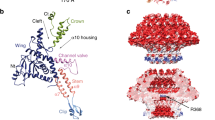
For successful infection, bacteriophages must overcome multiple barriers to transport their genome and proteins across the bacterial cell envelope. We use cryo-electron tomography to study the infection initiation of phage P22 in Salmonella enterica serovar Typhimurium, revealing how a channel forms to allow genome translocation into the cytoplasm. Our results show free phages that initially attach obliquely to the cell through interactions between the O antigen and two of the six tailspikes; the tail needle also abuts the cell surface. The virion then orients perpendicularly and the needle penetrates the outer membrane. The needle is released and the internal head protein gp7* is ejected and assembles into an extracellular channel that extends from the gp10 baseplate to the cell surface. A second protein, gp20, is ejected and assembles into a structure that extends the extracellular channel across the outer membrane into the periplasm. Insertion of the third ejected protein, gp16, into the cytoplasmic membrane probably completes the overall trans-envelope channel into the cytoplasm. Construction of a trans-envelope channel is an essential step during infection of Gram-negative bacteria by all short-tailed phages, because such virions cannot directly deliver their genome into the cell cytoplasm.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ackermann, H. W. Bacteriophage observations and evolution. Res. Microbiol. 154, 245–251 (2003).
Casjens, S. R. & Molineux, I. J. Short noncontractile tail machines: adsorption and DNA delivery by podoviruses. Adv. Exp. Med. Biol. 726, 143–179 (2012).
Davidson, A. R., Cardarelli, L., Pell, L. G., Radford, D. R. & Maxwell, K. L. Long noncontractile tail machines of bacteriophages. Adv. Exp. Med. Biol. 726, 115–142 (2012).
Leiman, P. G. & Shneider, M. M. Contractile tail machines of bacteriophages. Adv. Exp. Med. Biol. 726, 93–114 (2012).
Hu, B., Margolin, W., Molineux, I. J. & Liu, J. Structural remodeling of bacteriophage T4 and host membranes during infection initiation. Proc. Natl Acad. Sci. USA 112, E4919–E4928 (2015).
Taylor, N. M. I. et al. Structure of the T4 baseplate and its function in triggering sheath contraction. Nature 533, 346–352 (2016).
Farley, M. M., Tu, J., Kearns, D. B., Molineux, I. J. & Liu, J. Ultrastructural analysis of bacteriophage Phi29 during infection of Bacillus subtilis. J. Struct. Biol. 197, 163–171 (2017).
Xu, J., Gui, M., Wang, D. & Xiang, Y. The bacteriophage ϕ29 tail possesses a pore-forming loop for cell membrane penetration. Nature 534, 544–547 (2016).
Kemp, P., Garcia, L. R. & Molineux, I. J. Changes in bacteriophage T7 virion structure at the initiation of infection. Virology 340, 307–317 (2005).
Kemp, P., Gupta, M. & Molineux, I. J. Bacteriophage T7 DNA ejection into cells is initiated by an enzyme-like mechanism. Mol. Microbiol. 53, 1251–1265 (2004).
Chang, C. Y., Kemp, P. & Molineux, I. J. Gp15 and gp16 cooperate in translocating bacteriophage T7 DNA into the infected cell. Virology 398, 176–186 (2010).
Hu, B., Margolin, W., Molineux, I. J. & Liu, J. The bacteriophage T7 virion undergoes extensive structural remodeling during infection. Science 339, 576–579 (2013).
Israel, V. E proteins of bacteriophage P22. I. Identification and ejection from wild-type and defective particles. J. Virol. 23, 91–97 (1977).
Molineux, I. J. & Panja, D. Popping the cork: mechanisms of phage genome ejection. Nat. Rev. Microbiol. 11, 194–204 (2013).
Susskind, M. M. & Botstein, D. Molecular genetics of bacteriophage P22. Microbiol. Rev. 42, 385–413 (1978).
Hryc, C. F. et al. Accurate model annotation of a near-atomic resolution cryo-EM map. Proc. Natl Acad. Sci. USA 114, 3103–3108 (2017).
Lander, G. C. et al. The P22 tail machine at subnanometer resolution reveals the architecture of an infection conduit. Structure 17, 789–799 (2009).
Chang, J., Weigele, P., King, J., Chiu, W. & Jiang, W. Cryo-EM asymmetric reconstruction of bacteriophage P22 reveals organization of its DNA packaging and infecting machinery. Structure 14, 1073–1082 (2006).
Lander, G. C. et al. The structure of an infectious P22 virion shows the signal for headful DNA packaging. Science 312, 1791–1795 (2006).
Tang, J. et al. Peering down the barrel of a bacteriophage portal: the genome packaging and release valve in p22. Structure 19, 496–502 (2011).
Olia, A. S., Prevelige, P. E. Jr, Johnson, J. E. & Cingolani, G. Three-dimensional structure of a viral genome-delivery portal vertex. Nat. Struct. Mol. Biol. 18, 597–603 (2011).
Steinbacher, S. et al. Crystal structure of P22 tailspike protein: interdigitated subunits in a thermostable trimer. Science 265, 383–386 (1994).
Steinbacher, S. et al. Crystal structure of phage P22 tailspike protein complexed with Salmonella sp. O-antigen receptors. Proc. Natl Acad. Sci. USA 93, 10584–10588 (1996).
Steinbacher, S. et al. Phage P22 tailspike protein: crystal structure of the head-binding domain at 2.3 A, fully refined structure of the endorhamnosidase at 1.56 A resolution, and the molecular basis of O-antigen recognition and cleavage. J. Mol. Biol. 267, 865–880 (1997).
Olia, A. S., Casjens, S. & Cingolani, G. Structure of phage P22 cell envelope-penetrating needle. Nat. Struct. Mol. Biol. 14, 1221–1226 (2007).
Pintilie, G., Chen, D. H., Haase-Pettingell, C. A., King, J. A. & Chiu, W. Resolution and probabilistic models of components in cryoEM maps of mature P22 bacteriophage. Biophys. J. 110, 827–839 (2016).
Andres, D. et al. Tailspike interactions with lipopolysaccharide effect DNA ejection from phage P22 particles in vitro. J. Biol. Chem. 285, 36768–36775 (2010).
Berget, P. B. & Poteete, A. R. Structure and functions of the bacteriophage P22 tail protein. J. Virol. 34, 234–243 (1980).
King, J., Lenk, E. V. & Botstein, D. Mechanism of head assembly and DNA encapsulation in Salmonella phage P22. II. Morphogenetic pathway. J. Mol. Biol. 80, 697–731 (1973).
Lenk, E., Casjens, S., Weeks, J. & King, J. Intracellular visualization of precursor capsids in phage P22 mutant infected cells. Virology 68, 182–199 (1975).
Botstein, D., Waddell, C. H. & King, J. Mechanism of head assembly and DNA encapsulation in Salmonella phage p22. I. Genes, proteins, structures and DNA maturation. J. Mol. Biol. 80, 669–695 (1973).
Strauss, H. & King, J. Steps in the stabilization of newly packaged DNA during phage P22 morphogenesis. J. Mol. Biol. 172, 523–543 (1984).
Conlin, C. A., Trun, N. J., Silhavy, T. J. & Miller, C. G. Escherichia coli prlC encodes an endopeptidase and is homologous to the Salmonella Typhimurium opdA gene. J. Bacteriol. 174, 5881–5887 (1992).
Conlin, C. A., Vimr, E. R. & Miller, C. G. Oligopeptidase A is required for normal phage P22 development. J. Bacteriol. 174, 5869–5880 (1992).
Jin, Y. et al. Bacteriophage P22 ejects all of its internal proteins before its genome. Virology 485, 128–134 (2015).
Kastowsky, M., Gutberlet, T. & Bradaczek, H. Molecular modelling of the three-dimensional structure and conformational flexibility of bacterial lipopolysaccharide. J. Bacteriol. 174, 4798–4806 (1992).
Goldman, R. C. & Leive, L. Heterogeneity of antigenic-side-chain length in lipopolysaccharide from Escherichia coli 0111 and Salmonella Typhimurium LT2. Eur. J. Biochem. 107, 145–153 (1980).
Palva, E. T. & Makela, P. H. Lipopolysaccharide heterogeneity in Salmonella typhimurium analyzed by sodium dodecyl sulfate polyacrylamide gel electrophoresis. Eur. J. Biochem. 107, 137–143 (1980).
Baxa, U. et al. Interactions of phage P22 tails with their cellular receptor, Salmonella O-antigen polysaccharide. Biophys. J. 71, 2040–2048 (1996).
Israel, V. A model for the adsorption of phage P22 to Salmonella typhimurium. J. Gen. Virol. 40, 669–673 (1978).
Israel, V., Rosen, H. & Levine, M. Binding of bacteriophage P22 tail parts to cells. J. Virol. 10, 1152–1158 (1972).
Tu, J. G. et al. Dual host specificity of phage SP6 is facilitated by tailspike rotation. Virology 507, 206–215 (2017).
Leavitt, J. C. et al. The tip of the tail needle affects the rate of DNA delivery by bacteriophage P22. PLoS ONE 8, e70936 (2013).
Israel, V. Role of the bacteriophage P22 tail in the early stages of infection. J. Virol. 18, 361–364 (1976).
Perez, G. L., Huynh, B., Slater, M. & Maloy, S. Transport of phage P22 DNA across the cytoplasmic membrane. J. Bacteriol. 191, 135–140 (2009).
Zhao, H. et al. Structure of a bacterial virus DNA-injection protein complex reveals a decameric assembly with a constricted molecular channel. PLoS ONE 11, e0149337 (2016).
Botstein, D., Chan, R. K. & Waddell, C. H. Genetics of bacteriophage P22. II. Gene order and gene function. Virology 49, 268–282 (1972).
Poteete, A. R. & King, J. Functions of two new genes in Salmonella phage P22 assembly. Virology 76, 725–739 (1977).
Farley, M. M., Hu, B., Margolin, W. & Liu, J. Minicells, back in fashion. J. Bacteriol. 198, 1186–1195 (2016).
Reeve, J. N. Bacteriophage infection of minicells: a general method for identification of “in vivo” bacteriophage directed polypeptide biosynthesis. Mol. Gen. Genet. 158, 73–79 (1977).
Roozen, K. J., Fenwick, R. G. Jr. & Curtiss, R.3rd. Synthesis of ribonucleic acid and protein in plasmid-containing minicells of Escherichia coli K-12. J. Bacteriol. 107, 21–33 (1971).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Mastronarde, D. N. & Held, S. R. Automated tilt series alignment and tomographic reconstruction in IMOD. J. Struct. Biol. 197, 102–113 (2017).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Agulleiro, J. I. & Fernandez, J. J. Tomo3D 2.0—exploitation of advanced vector extensions (AVX) for 3D reconstruction. J. Struct. Biol. 189, 147–152 (2015).
Hrabe, T. et al. PyTom: a python-based toolbox for localization of macromolecules in cryo-electron tomograms and subtomogram analysis. J. Struct. Biol. 178, 177–188 (2012).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Acknowledgements
We thank S. Casjens for providing phage strains and amber mutant sequence data. We also thank M. M. Susskind and A. R. Poteete for phage and bacterial strains. This work was supported by grant nos. GM124378 and GM110243 to I.J.M. and J.L. C.W., J.T. and J.L. were also supported in part by grant no. AI087946 from the NIAID and grant no. AU-1714 from the Welch Foundation.
Author information
Authors and Affiliations
Contributions
J.L. and I.J.M. designed the research. C.W., J.T., I.J.M. and J.L. prepared the samples and collected and analysed the data. C.W., J.T., J.L. and I.J.M. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–4 and Supplementary Table 1.
Rights and permissions
About this article
Cite this article
Wang, C., Tu, J., Liu, J. et al. Structural dynamics of bacteriophage P22 infection initiation revealed by cryo-electron tomography. Nat Microbiol 4, 1049–1056 (2019). https://doi.org/10.1038/s41564-019-0403-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0403-z
This article is cited by
-
Nearly complete structure of bacteriophage DT57C reveals architecture of head-to-tail interface and lateral tail fibers
Nature Communications (2023)
-
Genomic analysis of Anderson typing phages of Salmonella Typhimrium: towards understanding the basis of bacteria-phage interaction
Scientific Reports (2023)
-
Phage diversity, genomics and phylogeny
Nature Reviews Microbiology (2020)
-
Translation of the long-term fundamental studies on viral DNA packaging motors into nanotechnology and nanomedicine
Science China Life Sciences (2020)
-
Structures of T7 bacteriophage portal and tail suggest a viral DNA retention and ejection mechanism
Nature Communications (2019)