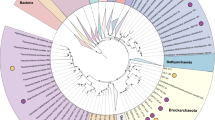
Abstract
Phylogenetic and geological evidence supports the hypothesis that life on Earth originated in thermal environments and conserved energy through methanogenesis or sulfur reduction. Here we describe two populations of the deeply rooted archaeal phylum Korarchaeota, which were retrieved from the metagenome of a circumneutral, suboxic hot spring that contains high levels of sulfate, sulfide, methane, hydrogen and carbon dioxide. One population is closely related to ‘Candidatus Korarchaeum cryptofilum OPF8’, while the more abundant korarchaeote, ‘Candidatus Methanodesulfokores washburnensis’, contains genes that are necessary for anaerobic methane and dissimilatory sulfur metabolisms. Phylogenetic and ancestral reconstruction analyses suggest that methane metabolism originated in the Korarchaeota, whereas genes for dissimilatory sulfite reduction were horizontally transferred to the Korarchaeota from the Firmicutes. Interactions among enzymes involved in both metabolisms could facilitate exergonic, sulfite-dependent, anaerobic oxidation of methane to methanol; alternatively, ‘Ca. M. washburnensis’ could conduct methanogenesis and sulfur reduction independently. Metabolic reconstruction suggests that ‘Ca. M. washburnensis’ is a mixotroph, capable of amino acid uptake, assimilation of methane-derived carbon and/or CO2 fixation by archaeal type III-b RuBisCO for scavenging ribose carbon. Our findings link anaerobic methane metabolism and dissimilatory sulfur reduction within a single deeply rooted archaeal population and have implications for the evolution of these traits throughout the Archaea.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Metagenome sequences used in this study are available on IMG/M (DOE-Joint Genome Institute) under genome identifier 3300005860. Metagenome-assembled genomes are available under NCBI BioProject accession number PRJNA492148. Access to the tSNE-based nucleotide frequency analysis algorithm can be obtained from the Center for Genomics and Bioinformatics at Indiana University. Newick files for three-domain and archaea-only phylogenomic trees are available as Supplementary Data 1–13.
References
Woese, C. R. Bacterial evolution. Microbiol. Rev. 51, 221–271 (1987).
Ueno, Y., Yamada, K., Yoshida, N., Maruyama, S. & Isozaki, Y. Evidence from fluid inclusions for microbial methanogenesis in the early Archaean era. Nature 440, 516–519 (2006).
Shen, Y., Buick, R. & Canfield, D. E. Isotopic evidence for microbial sulphate reduction in the early Archaean era. Nature 410, 77–81 (2001).
Dhillon, A., Goswami, S., Riley, M., Teske, A. & Sogin, M. Domain evolution and functional diversification of sulfite reductases. Astrobiology 5, 18–29 (2005).
Evans, P. N. et al. Methane metabolism in the archaeal phylum Bathyarchaeota revealed by genome-centric metagenomics. Science 350, 434–438 (2015).
Vanwonterghem, I. et al. Methylotrophic methanogenesis discovered in the archaeal phylum Verstraetearchaeota. Nat. Microbiol. 1, 16170 (2016).
Laso-Perez, R. et al. Thermophilic archaea activate butane via alkyl-coenzyme M formation. Nature 539, 396–401 (2016).
Spang, A., Caceres, E. F. & Ettema, T. J. G. Genomic exploration of the diversity, ecology, and evolution of the archaeal domain of life. Science 357, eaaf3883 (2017).
Anantharaman, K. et al. Expanded diversity of microbial groups that shape the dissimilatory sulfur cycle. ISME J. 12, 1715–1728 (2018).
Scheller, S., Goenrich, M., Boecher, R., Thauer, R. K. & Jaun, B. The key nickel enzyme of methanogenesis catalyses the anaerobic oxidation of methane. Nature 465, 606–608 (2010).
Loy, A. et al. Reverse dissimilatory sulfite reductase as phylogenetic marker for a subgroup of sulfur-oxidizing prokaryotes. Environ. Microbiol. 11, 289–299 (2009).
Hinrichs, K. U., Hayes, J. M., Sylva, S. P., Brewer, P. G. & DeLong, E. F. Methane-consuming archaebacteria in marine sediments. Nature 398, 802–805 (1999).
Boetius, A. et al. A marine microbial consortium apparently mediating anaerobic oxidation of methane. Nature 407, 623–626 (2000).
McGlynn, S. E., Chadwick, G. L., Kempes, C. P. & Orphan, V. J. Single cell activity reveals direct electron transfer in methanotrophic consortia. Nature 526, 531–535 (2015).
Milucka, J. et al. Zero-valent sulphur is a key intermediate in marine methane oxidation. Nature 491, 541–546 (2012).
Wegener, G., Krukenberg, V., Riedel, D., Tegetmeyer, H. E. & Boetius, A. Intercellular wiring enables electron transfer between methanotrophic archaea and bacteria. Nature 526, 587–590 (2015).
Elkins, J. G. et al. A korarchaeal genome reveals insights into the evolution of the Archaea. Proc. Natl Acad. Sci. USA 105, 8102–8107 (2008).
Inskeep, W. P. et al. Phylogenetic and functional analysis of metagenome sequence from high-temperature archaeal habitats demonstrate linkages between metabolic potential and geochemistry. Front. Microbiol. 4, 95 (2013).
McKay, L. J., Hatzenpichler, R., Inskeep, W. P. & Fields, M. W. Occurrence and expression of novel methyl-coenzyme M reductase gene (mcrA) variants in hot spring sediments. Sci. Rep. 7, 7252 (2017).
Barns, S. M., Delwiche, C. F., Palmer, J. D. & Pace, N. R. Perspectives on archaeal diversity, thermophily and monophyly from environmental rRNA sequences. Proc. Natl Acad. Sci. USA 93, 9188–9193 (1996).
Bowers, R. M. et al. Minimum information about a single amplified genome (MISAG) and a metagenome-assembled genome (MIMAG) of bacteria and archaea. Nat. Biotechnol. 35, 725–731 (2017).
van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Dick, G. J. et al. Community-wide analysis of microbial genome sequence signatures. Genome. Biol. 10, R85 (2009).
Slesarev, A. I. et al. The complete genome of hyperthermophile Methanopyrus kandleri AV19 and monophyly of archaeal methanogens. Proc. Natl Acad. Sci. USA 99, 4644–4649 (2002).
Huber, G., Spinnler, C., Gambacorta, A. & Stetter, K. O. Metallosphaera sedula gen, and sp. nov. represents a new genus of aerobic, metal-mobilizing, thermoacidophilic Archaebacteria. Syst. Appl. Microbiol. 12, 38–47 (1989).
Medini, D., Donati, C., Tettelin, H., Masignani, V. & Rappuoli, R. The microbial pan-genome. Curr. Opin. Genet. Dev. 15, 589–594 (2005).
Spang, A. et al. Complex archaea that bridge the gap between prokaryotes and eukaryotes. Nature 521, 173–179 (2015).
Zaremba-Niedzwiedzka, K. et al. Asgard archaea illuminate the origin of eukaryotic cellular complexity. Nature 541, 353–358 (2017).
Jay, Z. J. et al. Marsarchaeota are an aerobic archaeal lineage abundant in geothermal iron oxide microbial mats. Nat. Microbiol. 3, 732–740 (2018).
Da Cunha, V., Gaia, M., Gadelle, D., Nasir, A. & Forterre, P. Lokiarchaea are close relatives of Euryarchaeota, not bridging the gap between prokaryotes and eukaryotes. PLoS Genet. 13, e1006810 (2017).
Dridi, B., Fardeau, M. L., Ollivier, B., Raoult, D. & Drancourt, M. Methanomassiliicoccus luminyensis gen. nov., sp. nov., a methanogenic archaeon isolated from human faeces. Int. J. Syst. Evol. Microbiol. 62, 1902–1907 (2012).
Lever, M. A. & Teske, A. P. Diversity of methane-cycling archaea in hydrothermal sediment investigated by general and group-specific PCR primers. Appl. Environ. Microbiol. 81, 1426–1441 (2015).
Rinke, C. et al. Insights into the phylogeny and coding potential of microbial dark matter. Nature 499, 431–437 (2013).
Hua, Z. S. et al. Genomic inference of the metabolism and evolution of the archaeal phylum Aigarchaeota. Nat. Commun. 9, 2832 (2018).
Müller, A. L., Kjeldsen, K. U., Rattei, T., Pester, M. & Loy, A. Phylogenetic and environmental diversity of DsrAB-type dissimilatory (bi)sulfite reductases. ISME J. 9, 1152–1165 (2015).
Rabus, R. et al. A post-genomic view of the ecophysiology, catabolism and biotechnological relevance of sulphate-reducing prokaryotes. Adv. Microb. Physiol. 66, 55–321 (2015).
Borrel, G. et al. Wide diversity of methane and short-chain alkane metabolisms in uncultured archaea. Nat. Microbiol. https://doi.org/10.1038/s41564-019-0363-3 (2019).
Liu, Y. & Whitman, W. B. Metabolic, phylogenetic, and ecological diversity of the methanogenic archaea. Ann. NY Acad. Sci. 1125, 171–189 (2008).
Thauer, R. K., Kaster, A. K., Seedorf, H., Buckel, W. & Hedderich, R. Methanogenic archaea: ecologically relevant differences in energy conservation. Nat. Rev. Microbiol. 6, 579–591 (2008).
Wagner, T., Koch, J., Ermler, U. & Shima, S. Methanogenic heterodisulfide reductase (HdrABC-MvhAGD) uses two noncubane [4Fe-4S] clusters for reduction. Science 357, 699–703 (2017).
Lang, K. et al. New mode of energy metabolism in the seventh order of methanogens as revealed by comparative genome analysis of “Candidatus methanoplasma termitum”. Appl. Environ. Microbiol. 81, 1338–1352 (2015).
Amend, J. P. & Shock, E. L. Energetics of overall metabolic reactions of thermophilic and hyperthermophilic Archaea and Bacteria. FEMS Microbiol. Rev. 25, 175–243 (2001).
Hedderich, R. et al. Anaerobic respiration with elemental sulfur and with disulfides. FEMS Microbiol. Rev. 22, 353–381 (1998).
Pereira, I. A. et al. A comparative genomic analysis of energy metabolism in sulfate reducing bacteria and archaea. Front. Microbiol. 2, 69 (2011).
Dong, M. et al. In vitro methanol production from methyl coenzyme M using the Methanosarcina barkeri MtaABC protein complex. Biotechnol. Prog. 33, 1243–1249 (2017).
Haroon, M. F. et al. Anaerobic oxidation of methane coupled to nitrate reduction in a novel archaeal lineage. Nature 500, 567–570 (2013).
Welander, P. V. & Metcalf, W. W. Mutagenesis of the C1 oxidation pathway in Methanosarcina barkeri: new insights into the Mtr/Mer bypass pathway. J. Bacteriol. 190, 1928–1936 (2008).
Meyerdierks, A. et al. Metagenome and mRNA expression analyses of anaerobic methanotrophic archaea of the ANME-1 group. Environ. Microbiol. 12, 422–439 (2010).
Kono, T. et al. A RuBisCO-mediated carbon metabolic pathway in methanogenic archaea. Nat. Commun. 8, 14007 (2017).
Sato, T., Atomi, H. & Imanaka, T. Archaeal type III RuBisCOs function in a pathway for AMP metabolism. Science 315, 1003–1006 (2007).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Li, D., Liu, C. M., Luo, R., Sadakane, K. & Lam, T. W. MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31, 1674–1676 (2015).
Eren, A. M. et al. Anvi’o: an advanced analysis and visualization platform for ‘omics data. PeerJ 3, e1319 (2015).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11, 119 (2010).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Tatusov, R. L., Galperin, M. Y., Natale, D. A. & Koonin, E. V. The COG database: a tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res. 28, 33–36 (2000).
Delmont, T. O. & Eren, A. M. Linking pangenomes and metagenomes: the Prochlorococcus metapangenome. PeerJ 6, e4320 (2018).
van Dongen, S. & Abreu-Goodger, C. Using MCL to extract clusters from networks. Methods Mol. Biol. 804, 281–295 (2012).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44, D457–D462 (2016).
Finn, R. D. et al. Pfam: the protein families database. Nucleic Acids Res. 42, D222–D230 (2014).
Richter, M. & Rosselló-Móra, R. Shifting the genomic gold standard for the prokaryotic species definition. Proc. Natl Acad. Sci. USA 106, 19126–19131 (2009).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012).
Abascal, F., Zardoya, R. & Posada, D. ProtTest: selection of best-fit models of protein evolution. Bioinformatics 21, 2104–2105 (2005).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Nguyen, L. T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Pruesse, E., Peplies, J. & Glockner, F. O. SINA: accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 28, 1823–1829 (2012).
Ludwig, W. et al. ARB: a software environment for sequence data. Nucleic Acids Res. 32, 1363–1371 (2004).
Acknowledgements
The authors appreciate support from the NASA Postdoctoral Program through the NASA Astrobiology Institute (L.J.M.), the Montana Agricultural Experiment Station (project 911300; W.P.I.), the National Institutes of Health IDeA Program (COBRE grant GM110732; M.D.), the NSF Integrative Graduate Education and Research Traineeship Program (NSF DGE 0654336; Z.J.J.), the W. M. Keck Foundation (L.J.M., M.W.F., K.B.K. and W.P.I.), NSF 1736255 (L.J.M. and M.W.F.) and by the US Department of Energy—Ecosystems and Networks Integrated with Genes and Molecular Assemblies (contract number DE-AC02–05CH11231; M.W.F.). Metagenome sequencing of DNA from Washburn Hot Springs was conducted at the DOE-Joint Genome Institute under the Community Sequencing Program (CSP 701; W.P.I.). Computations were performed on the Hyalite High-Performance Computing System, operated and supported by MSU’s Information Technology Center. We thank the NSF EarthCube ECOGEO RCN (1440066) for metagenomics analysis support and individuals involved in the 2016 ‘Omics Workshop’ at the University of Hawaii; V. Krukenberg, G. Borrel, S. Gribaldo, R. Hatzenpichler, M. Lever and A. Teske for helpful discussions and J. Beam for assistance with sample collection and processing.
Author information
Authors and Affiliations
Contributions
W.P.I. and L.J.M. designed the investigation. W.P.I. and Z.J.J. collected samples and performed initial metagenome processing. L.J.M. and M.D. performed clustering, coverage analyses and phylogenetic analyses. L.J.M., M.W.F. and W.P.I. built metabolic reconstructions. M.D. performed additional metagenome assemblies, phylogenomic, emergent self-organizing maps and sequence reconstruction analyses. T.O.D. and A.M.E. performed pangenomics and microdiversity analyses. M.D., K.B.K. and L.J.M. performed phylogenetic analyses of McrA and DsrAB protein sequences. K.B.K. and L.J.M. performed phylogenetic analysis of mcrA genes. D.B.R. and M.D. assisted with nucleotide frequency analyses. L.J.M., W.P.I. and M.D. wrote the manuscript and responses to reviewer comments. All authors contributed to manuscript editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2, Supplementary Table 5, Supplementary Table 7, Supplementary Figures 1–12, Supplementary Discussion, Supplementary Data Legends and Supplementary References.
Supplementary Table 3
Gene clusters, clusters of orthologous genes identification numbers and functions corresponding to each of the Korarchaeota displayed in Fig. 1.
Supplementary Table 4
List of archaeal clusters of orthologous genes used in phylogenomic analysis.
Supplementary Table 6
Protein BLAST comparisons of ancestral dissimilatory sulfur reductase sequences to Candidatus Methanodesulfokores washburnensis and the NCBI protein database.
Supplementary Table 8
List of abbreviations used in metabolic reconstruction (Fig. 4).
Dataset 1
Newick tree files corresponding to Fig. 2a.
Dataset 2
Newick tree files corresponding to Supplementary Figure 5a.
Dataset 3
Newick tree files corresponding to Supplementary Figure 5b.
Dataset 4
Newick tree files corresponding to Supplementary Figure 5c.
Dataset 5
Newick tree files corresponding to Supplementary Figure 5d.
Dataset 6
Newick tree files corresponding to Supplementary Figure 5e.
Dataset 7
Newick tree files corresponding to Supplementary Figure 5f.
Dataset 8
Newick tree files corresponding to Supplementary Figure 5g.
Dataset 9
Newick tree files corresponding to Supplementary Figure 5h.
Dataset 10
Newick tree files corresponding to Supplementary Figure 5i.
Dataset 11
Newick tree files corresponding to Supplementary Figure 5j.
Dataset 12
Newick tree files corresponding to Supplementary Figure 5k.
Dataset 13
Concatenated alignment file (.afa) of 56 conserved proteins from three domains. File size = 2 MB.
Rights and permissions
About this article
Cite this article
McKay, L.J., Dlakić, M., Fields, M.W. et al. Co-occurring genomic capacity for anaerobic methane and dissimilatory sulfur metabolisms discovered in the Korarchaeota. Nat Microbiol 4, 614–622 (2019). https://doi.org/10.1038/s41564-019-0362-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0362-4
This article is cited by
-
A compendium of viruses from methanogenic archaea reveals their diversity and adaptations to the gut environment
Nature Microbiology (2023)
-
Diversity and function of methyl-coenzyme M reductase-encoding archaea in Yellowstone hot springs revealed by metagenomics and mesocosm experiments
ISME Communications (2023)
-
Discovery of a novel bacterial class with the capacity to drive sulfur cycling and microbiome structure in a paleo-ocean analog
ISME Communications (2023)
-
Conserved and lineage-specific hypothetical proteins may have played a central role in the rise and diversification of major archaeal groups
BMC Biology (2022)
-
Expanding the phylogenetic distribution of cytochrome b-containing methanogenic archaea sheds light on the evolution of methanogenesis
The ISME Journal (2022)