Abstract
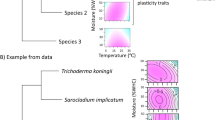
Fungi are the primary agents of terrestrial decomposition, yet our understanding of fungal biogeography lags far behind that of plants, animals and bacteria. Here, we use a trait-based approach to quantify the niches of 23 species of basidiomycete wood decay fungi from across North America, and explore the linkages among functional trait expression, climate and phylogeny. Our analysis reveals a fundamental trade-off between abiotic stress tolerance and competitive ability, whereby fungi with wide thermal and moisture niches exhibit lower displacement ability. The magnitude of this dominance-tolerance trade-off is partially related to the environmental conditions under which the fungi were collected, with thermal niche traits exhibiting the strongest climate relationships. Nevertheless, moisture and thermal dominance-tolerance patterns exhibited contrasting phylogenetic signals, suggesting that these trends are influenced by a combination of niche sorting along taxonomic lines in tandem with acclimation and adaptation at the level of the individual. Collectively, our work reveals key insight into the life history strategies of saprotrophic fungi, demonstrating consistent trait trade-offs across broad spatial scales.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Code availability
All code needed to reproduce these results can be obtained from https://github.com/dsmaynard/fungal_biogeography.
Data availability
Data are archived in a dedicated GitHub repository: https://github.com/dsmaynard/fungal_biogeography. The ITS and LSU sequences were previously deposited in GenBank under accession numbers KX065932–KX065968 and KX065969–KX066002.
References
Treseder, K. K. & Lennon, J. T. Fungal traits that drive ecosystem dynamics on land. Microbiol. Mol. Biol. Rev. 9, 243–262 (2015).
Crowther, T. W. et al. Untangling the fungal niche: the trait-based approach. Front. Microbiol. 5, 579 (2014).
Bradford, M. A. et al. Climate fails to predict wood decomposition at regional scales. Nat. Clim. Change 4, 625–630 (2014).
Ottosson, E. et al. Species associations during the succession of wood-inhabiting fungal communities. Fungal Ecol. 11, 17–28 (2014).
Tedersoo, L. et al. Fungal biogeography. Global diversity and geography of soil fungi. Science 346, 1256688 (2014).
Bahram, M. et al. Structure and function of the global topsoil microbiome. Nature 560, 233–237 (2018).
Glassman, S. I., Wang, I. J. & Bruns, T. D. Environmental filtering by pH and soil nutrients drives community assembly in fungi at fine spatial scales. Mol. Ecol. 26, 6960–6973 (2017).
Boddy, L. Fungal community ecology and wood decomposition processes in angiosperms: from standing tree to complete decay of coarse woody debris. Ecol. Bull. 49, 43–56 (2001).
van der Wal, A. et al. Fungal biomass development in a chronosequence of land abandonment. Soil Biol. Biochem. 38, 51–60 (2006).
Meyer, K. M. et al. Why do microbes exhibit weak biogeographic patterns? ISME J. 12, 1404–1413 (2018).
Meiser, A., Bálint, M. & Schmitt, I. Meta-analysis of deep-sequenced fungal communities indicates limited taxon sharing between studies and the presence of biogeographic patterns. New Phytol. 201, 623–635 (2014).
Kivlin, S. N., Hawkes, C. V. & Treseder, K. K. Global diversity and distribution of arbuscular mycorrhizal fungi. Soil Biol. Biochem. 43, 2294–2303 (2011).
Aguilar-Trigueros, C. A., Powell, J. R., Anderson, I. C., Antonovics, J. & Rillig, M. C. Ecological understanding of root-infecting fungi using trait-based approaches. Trends Plant Sci. 19, 432–438 (2014).
Lennon, J. T., Aanderud, Z. T., Lehmkuhl, B. K. & Schoolmaster, D. R.Jr. Mapping the niche space of soil microorganisms using taxonomy and traits. Ecology 93, 1867–1879 (2012).
Maynard, D. S., Crowther, T. W. & Bradford, M. A. Fungal interactions reduce carbon use efficiency. Ecol. Lett. 20, 1034–1042 (2017).
Maynard, D. S. et al. Diversity begets diversity in competition for space. Nat. Ecol. Evol. 1, 156 (2017).
Maynard, D. S., Crowther, T. W. & Bradford, M. A. Competitive network determines the direction of the diversity-function relationship. Proc. Natl Acad. Sci. USA 114, 11464–11469 (2017).
Crowther, T. W., Boddy, L. & Maynard, D. S. The use of artificial media in fungal ecology. Fungal Ecol. 32, 87–91 (2018).
Crowther, T. W. & Bradford, M. A. Thermal acclimation in widespread heterotrophic soil microbes. Ecol. Lett. 16, 469–477 (2013).
Hochachka, P. W. & Somero, G. N. Biochemical Adaptation: Mechanism and Process in Physiological Evolution (Oxford Univ. Press, New York, 2002).
Crowther, T. W., Boddy, L. & Jones, T. H. Species-specific effects of soil fauna on fungal foraging and decomposition. Oecologia 167, 535–545 (2011).
Boddy, L. Saprotrophic cord-forming fungi: warfare strategies and other ecological aspects. Mycol. Res. 97, 641–655 (1993).
Aguilar-Trigueros, C. A., Rillig, M. C. & Crowther, T. W. Applying allometric theory to fungi. ISME J. 11, 2175–2180 (2017).
Gasch, A. P. Comparative genomics of the environmental stress response in ascomycete fungi. Yeast 24, 961–976 (2007).
Isidorov, V., Tyszkiewicz, Z. & Pirożnikow, E. Fungal succession in relation to volatile organic compounds emissions from Scots pine and Norway spruce leaf litter-decomposing fungi. Atmos. Environ. 131, 301–306 (2016).
El Ariebi, N., Hiscox, J., Scriven, S. A., Müller, C. T. & Boddy, L. Production and effects of volatile organic compounds during interspecific interactions. Fungal Ecol. 20, 144–154 (2016).
Hiscox, J., Baldrian, P., Rogers, H. J. & Boddy, L. Changes in oxidative enzyme activity during interspecific mycelial interactions involving the white-rot fungus Trametes versicolor. Fungal Genet. Biol. 47, 562–571 (2010).
Junninen, K., Similä, M., Kouki, J. & Kotiranta, H. Assemblages of wood-inhabiting fungi along the gradients of succession and naturalness in boreal pine-dominated forests in Fennoscandia. Ecography 29, 75–83 (2006).
Lindahl, B. D. & Finlay, R. D. Activities of chitinolytic enzymes during primary and secondary colonization of wood by basidiomycetous fungi. New Phytol. 169, 389–397 (2006).
Revell, L. J. Phylogenetic signal and linear regression on species data. Methods Ecol. Evol. 1, 319–329 (2010).
Loescher, H., Ayres, E., Duffy, P., Luo, H. & Brunke, M. Spatial variation in soil properties among North American ecosystems and guidelines for sampling designs. PLoS ONE 9, e83216 (2014).
Bradford, M. A. et al. A test of the hierarchical model of litter decomposition. Nat. Ecol. Evol. 1, 1836–1845 (2017).
Ellison, C. E. et al. Population genomics and local adaptation in wild isolates of a model microbial eukaryote. Proc. Natl Acad. Sci. USA 108, 2831–2836 (2011).
Grime, J. P. Vegetation classification by reference to strategies. Nature 250, 26–31 (1974).
Pérez-Harguindeguy, N. et al. New handbook for standardised measurement of plant functional traits worldwide. Aust. J. Bot. 61, 167–234 (2013).
Ramirez, M. L., Chulze, S. N. & Magan, N. Impact of osmotic and matric water stress on germination, growth, mycelial water potentials and endogenous accumulation of sugars and sugar alcohols in Fusarium graminearum. Mycologia 96, 470–478 (2004).
Boddy, L. Effect of temperature and water potential on growth rate of wood-rotting basidiomycetes. Trans. Br. Mycol. Soc. 80, 141–149 (1983).
Jurado, M., Marín, P., Magan, N. & González-Jaén, M. T. Relationship between solute and matric potential stress, temperature, growth, and FUM1 gene expression in two Fusarium verticillioides strains from Spain. Appl. Environ. Microbiol. 74, 2032–2036 (2008).
Molloy, S. Sugar Transport and Water Relations of Agaricus bisporus. PhD thesis, Cranfield Univ. (2004).
Elo, A. E. The Rating of Chess Players, Past and Present (Arco Pub., New York, 1986).
Hijmans, R. J., Cameron, S. E., Parra, J. L., Jones, P. G. & Jarvis, A. Very high resolution interpolated climate surfaces for global land areas. Int. J. Climatol. 25, 1965–1978 (2005).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Series B Stat. Methodol. 57, 289–300 (1995).
Chevan, A. & Sutherland, M. Hierarchical partitioning. Am. Stat. 45, 90–96 (1991).
Lustenhouwer, N., Moran, E. V. & Levine, J. M. Trait correlations equalize spread velocity across plant life histories. Glob. Ecol. Biogeogr. 26, 1398–1407 (2017).
Tuanmu, M. N. & Jetz, W. A global 1-km consensus land-cover product for biodiversity and ecosystem modelling. Glob. Ecol. Biogeogr. 23, 1031–1045 (2014).
Kottek, M., Grieser, J., Beck, C., Rudolf, B. & Rubel, F. World map of the Köppen–Geiger climate classification updated. Meteorol. Z. 15, 259–263 (2006).
Acknowledgements
We thank S. Thomas, A. Neupane and E. Karlsen-Ayala for laboratory assistance, and L. Boddy for discussions throughout this study. This work was supported by grants to T.W.C. from DOB Ecology, Plant-for-the-Planet, the Marie Skłodowska-Curie Actions Fellowship and the German Federal Ministry for Economic Cooperation and Development; and by grants to M.A.B. and D.S.M. from the US National Science Foundation (grant nos. DEB-1457614 and DEB-1601036).
Author information
Authors and Affiliations
Contributions
The study was conceived by T.W.C. The experimental work was designed by D.S.M. and T.W.C. and conducted by T.W.C., D.S.M., K.R.C., D.L. and J.G. The statistical analyses were performed by D.S.M., T.W.C., D.A.T., P.J.T. and D.M.W. The manuscript was written by D.S.M., T.W.C. and M.A.B., with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Materials or Methods, Supplementary References, Supplementary Tables 1–6 and Supplementary Figures 1 and 2.
Rights and permissions
About this article
Cite this article
Maynard, D.S., Bradford, M.A., Covey, K.R. et al. Consistent trade-offs in fungal trait expression across broad spatial scales. Nat Microbiol 4, 846–853 (2019). https://doi.org/10.1038/s41564-019-0361-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0361-5
This article is cited by
-
Above- and belowground fungal biodiversity of Populus trees on a continental scale
Nature Microbiology (2023)
-
Dispersal changes soil bacterial interactions with fungal wood decomposition
ISME Communications (2023)
-
Inter-annual Persistence of Canopy Fungi Driven by Abundance Despite High Spatial Turnover
Microbial Ecology (2023)
-
Eco-physiological Responses of Aquatic Fungi to Three Global Change Stressors Highlight the Importance of Intraspecific Trait Variability
Microbial Ecology (2023)
-
Temperate species underfill their tropical thermal potentials on land
Nature Ecology & Evolution (2023)