Abstract
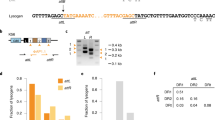
Type III-A CRISPR–Cas systems employ the Cas10–Csm complex to destroy bacteriophages and plasmids, using a guide RNA to locate complementary RNA molecules from the invader and trigger an immune response that eliminates the infecting DNA. In addition, these systems possess the non-specific RNase Csm6, which provides further protection for the host. While the role of Csm6 in immunity during phage infection has been determined, how this RNase is used against plasmids is unclear. Here, we show that Staphylococcus epidermidis Csm6 is required for immunity when transcription across the plasmid target is infrequent, leading to impaired target recognition and inefficient DNA degradation by the Cas10–Csm complex. In these conditions, Csm6 causes growth arrest in the host and prevents further plasmid replication through the indiscriminate degradation of host and plasmid transcripts. In contrast, when plasmid target sequences are efficiently transcribed, Csm6 is dispensable and DNA degradation by Cas10 is sufficient for anti-plasmid immunity. Csm6 therefore provides robustness to the type III-A CRISPR–Cas immune response against difficult targets for the Cas10–Csm complex.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data from this study are available from the authors upon request. The raw data for the RNA-seq experiments can be found at the Sequence Read Archive (NIH) through accession code PRJNA506073. Original gel pictures and northern blots are provided in Supplementary Fig. 6.
References
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007).
Marraffini, L. A. & Sontheimer, E. J. CRISPR interference limits horizontal gene transfer in staphylococci by targeting DNA. Science 322, 1843–1845 (2008).
Gasiunas, G., Barrangou, R., Horvath, P. & Siksnys, V. Cas9–crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc. Natl Acad. Sci. USA 109, E2579–E2586 (2012).
Hale, C. R. et al. RNA-guided RNA cleavage by a CRISPR RNA–Cas protein complex. Cell 139, 945–956 (2009).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Westra, E. R. et al. CRISPR immunity relies on the consecutive binding and degradation of negatively supercoiled invader DNA by Cascade and Cas3. Mol. Cell 46, 595–605 (2012).
Koonin, E. V., Makarova, K. S. & Zhang, F. Diversity, classification and evolution of CRISPR–Cas systems. Curr. Opin. Microbiol. 37, 67–78 (2017).
Pyenson, N. C. & Marraffini, L. A. Type III CRISPR–Cas systems: when DNA cleavage just isn’t enough. Curr. Opin. Microbiol. 37, 150–154 (2017).
Staals, R. H. et al. RNA targeting by the type III-A CRISPR–cas csm complex of Thermus thermophilus. Mol. Cell 56, 518–530 (2014).
Tamulaitis, G. et al. Programmable RNA shredding by the Type III-A CRISPR–Cas system of Streptococcus thermophilus. Mol. Cell 56, 506–517 (2014).
Zhang, J. et al. Structure and mechanism of the CMR complex for CRISPR-mediated antiviral immunity. Mol. Cell 45, 303–313 (2012).
Elmore, J. R. et al. Bipartite recognition of target RNAs activates DNA cleavage by the Type III-B CRISPR–Cas system. Genes Dev. 30, 447–459 (2016).
Estrella, M. A., Kuo, F. T. & Bailey, S. RNA-activated DNA cleavage by the Type III-B CRISPR–Cas effector complex. Genes Dev. 30, 460–470 (2016).
Kazlauskiene, M., Tamulaitis, G., Kostiuk, G., Venclovas, C. & Siksnys, V. Spatiotemporal control of type III-A CRISPR–Cas immunity: coupling DNA degradation with the target RNA recognition. Mol. Cell 62, 295–306 (2016).
Samai, P. et al. Co-transcriptional DNA and RNA cleavage during type III CRISPR–Cas immunity. Cell 161, 1164–1174 (2015).
Makarova, K. S. et al. An updated evolutionary classification of CRISPR–Cas systems. Nat. Rev. Microbiol. 13, 722–736 (2015).
Niewoehner, O. & Jinek, M. Structural basis for the endoribonuclease activity of the type III-A CRISPR-associated protein Csm6. RNA 22, 318–329 (2016).
Burroughs, A. M., Zhang, D., Schaffer, D. E., Iyer, L. M. & Aravind, L. Comparative genomic analyses reveal a vast, novel network of nucleotide-centric systems in biological conflicts, immunity and signaling. Nucleic Acids Res. 43, 10633–10654 (2015).
Kazlauskiene, M., Kostiuk, G., Venclovas, C., Tamulaitis, G. & Siksnys, V. A cyclic oligonucleotide signaling pathway in type III CRISPR–Cas systems. Science 357, 605–609 (2017).
Niewoehner, O. et al. Type III CRISPR–Cas systems produce cyclic oligoadenylate second messengers. Nature 548, 543–548 (2017).
Rouillon, C., Athukoralage, J. S., Graham, S., Gruschow, S. & White, M. F. Control of cyclic oligoadenylate synthesis in a type III CRISPR system. eLife 7, e36734 (2018).
Athukoralage, J. S., Rouillon, C., Graham, S., Gruschow, S. & White, M. F. Ring nucleases deactivate type III CRISPR ribonucleases by degrading cyclic oligoadenylate. Nature 562, 277–280 (2018).
Jiang, W., Samai, P. & Marraffini, L. A. Degradation of phage transcripts by CRISPR-associated RNases enables type III CRISPR–Cas immunity. Cell 164, 710–721 (2016).
Foster, K., Kalter, J., Woodside, W., Terns, R. M. & Terns, M. P. The ribonuclease activity of Csm6 is required for anti-plasmid immunity by Type III-A CRISPR–Cas systems. RNA Biol. https://doi.org/10.1080/15476286.2018.1493334 (2018).
Hatoum-Aslan, A., Maniv, I., Samai, P. & Marraffini, L. A. Genetic characterization of antiplasmid immunity through a type III-A CRISPR–CAS system. J. Bacteriol. 196, 310–317 (2014).
Anantharaman, V., Iyer, L. M. & Aravind, L. Presence of a classical RRM-fold palm domain in Thg1-type 3′–5′nucleic acid polymerases and the origin of the GGDEF and CRISPR polymerase domains. Biol. Direct. 5, 43 (2010).
Marraffini, L. A. & Sontheimer, E. J. Self versus non-self discrimination during CRISPR RNA-directed immunity. Nature 463, 568–571 (2010).
Deng, L., Garrett, R. A., Shah, S. A., Peng, X. & She, Q. A novel interference mechanism by a type IIIB CRISPR-Cmr module in Sulfolobus. Mol. Microbiol. 87, 1088–1099 (2013).
Goldberg, G. W., Jiang, W., Bikard, D. & Marraffini, L. A. Conditional tolerance of temperate phages via transcription-dependent CRISPR–Cas targeting. Nature 514, 633–637 (2014).
Kreiswirth, B. N. et al. The toxic shock syndrome exotoxin structural gene is not detectably transmitted by a prophage. Nature 305, 709–712 (1983).
Helle, L. et al. Vectors for improved Tet repressor-dependent gradual gene induction or silencing in Staphylococcus aureus. Microbiology 157, 3314–3323 (2011).
Liu, T. Y., Iavarone, A. T. & Doudna, J. A. RNA and DNA targeting by a reconstituted Thermus thermophilus Type III-A CRISPR–Cas system. PLoS ONE 12, e0170552 (2017).
Ruiz-Maso, J. A. et al. Plasmid rolling-circle replication. Microbiol Spectr. 3, PLAS-0035-2014 (2015).
Sheppard, N. F., Glover, C. V. 3rd, Terns, R. M. & Terns, M. P. The CRISPR-associated Csx1 protein of Pyrococcus furiosus is an adenosine-specific endoribonuclease. RNA 22, 216–224 (2016).
Anantharaman, V., Makarova, K. S., Burroughs, A. M., Koonin, E. V. & Aravind, L. Comprehensive analysis of the HEPN superfamily: identification of novel roles in intra-genomic conflicts, defense, pathogenesis and RNA processing. Biol. Direct. 8, 15 (2013).
Upton, J. W. & Chan, F. K. Staying alive: cell death in antiviral immunity. Mol. Cell 54, 273–280 (2014).
Labrie, S. J., Samson, J. E. & Moineau, S. Bacteriophage resistance mechanisms. Nat. Rev. Microbiol. 8, 317–327 (2010).
Horinouchi, S. & Weisblum, B. Nucleotide sequence and functional map of pE194, a plasmid that specifies inducible resistance to macrolide, lincosamide, and streptogramin type B antibodies. J. Bacteriol. 150, 804–814 (1982).
Horinouchi, S. & Weisblum, B. Nucleotide sequence and functional map of pC194, a plasmid that specifies inducible chloramphenicol resistance. J. Bacteriol. 150, 815–825 (1982).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Ray, M. D., Boundy, S. & Archer, G. L. Transfer of the methicillin resistance genomic island among staphylococci by conjugation. Mol. Microbiol. 100, 675–685 (2016).
Lamberte, L. E. et al. Horizontally acquired AT-rich genes in Escherichia coli cause toxicity by sequestering RNA polymerase. Nat. Microbiol. 2, 16249 (2017).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Acknowledgements
The authors would like to thank A. Meeske and C. Mo for critical reading of the manuscript. They also thank the following: G. Goldberg for plasmid pGG25; C. Kenney and W. Jiang for sharing their insights on spc1-flip conjugation; the Rockefeller University Genomics Resource Center for performing the Csm6-targeting Nextseq RNA-seq experiment; and T. Carroll (of the Rockefeller University Bioinformatics Resource Center) and E. Stoyanova for helpful discussions on the bioinformatic analysis. J.T.R. was supported by a Boehringer Ingelheim Fonds Ph.D. fellowship. L.M. is supported by a Burroughs Wellcome Fund PATH Award and a NIH Director’s Pioneer Award (DP1GM128184).
Author information
Authors and Affiliations
Contributions
J.T.R. and L.M. designed the study. J.T.R performed all experiments and analysed the next-generation sequencing data. J.T.R. and L.M. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
L.M. is a cofounder and Scientific Advisory Board member of Intellia Therapeutics and a cofounder of Eligo Biosciences. J.T.R. declares no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–6, Supplementary Tables 1–3 and Supplementary References.
Rights and permissions
About this article
Cite this article
Rostøl, J.T., Marraffini, L.A. Non-specific degradation of transcripts promotes plasmid clearance during type III-A CRISPR–Cas immunity. Nat Microbiol 4, 656–662 (2019). https://doi.org/10.1038/s41564-018-0353-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-018-0353-x
This article is cited by
-
DNA-targeting short Argonautes complex with effector proteins for collateral nuclease activity and bacterial population immunity
Nature Microbiology (2024)
-
Bacteriophages avoid autoimmunity from cognate immune systems as an intrinsic part of their life cycles
Nature Microbiology (2024)
-
The CRISPR effector Cam1 mediates membrane depolarization for phage defence
Nature (2024)
-
Molecular mechanism of allosteric activation of the CRISPR ribonuclease Csm6 by cyclic tetra-adenylate
The EMBO Journal (2023)
-
Precise transcript targeting by CRISPR-Csm complexes
Nature Biotechnology (2023)