Abstract
Emerging viruses have the potential to impose substantial mortality, morbidity and economic burdens on human populations. Tracking the spread of infectious diseases to assist in their control has traditionally relied on the analysis of case data gathered as the outbreak proceeds. Here, we describe how many of the key questions in infectious disease epidemiology, from the initial detection and characterization of outbreak viruses, to transmission chain tracking and outbreak mapping, can now be much more accurately addressed using recent advances in virus sequencing and phylogenetics. We highlight the utility of this approach with the hypothetical outbreak of an unknown pathogen, ‘Disease X’, suggested by the World Health Organization to be a potential cause of a future major epidemic. We also outline the requirements and challenges, including the need for flexible platforms that generate sequence data in real-time, and for these data to be shared as widely and openly as possible.
Similar content being viewed by others
Main
Emerging infectious diseases present one of the greatest public health challenges of the twenty-first century. Among these are zoonotic viruses that originate from reservoir species, often mammals, and jump to humans to cause disease syndromes of varying form and severity. An emerging virus, depending on its ability to transmit among humans, can lead to individual or a few sporadic cases, resulting in a localized outbreak that requires public health intervention or, in the worst scenarios, can develop into a large epidemic or global pandemic. Such emergence events over the past two decades are numerous and varied. They include viruses not previously encountered, such as the SARS and MERS coronaviruses1,2,3, and familiar foes that have reappeared to cause outbreaks, such as swine- and avian-origin influenza4,5, and Ebola6 and Zika7 viruses. Although many outbreaks end naturally or are controlled quickly, questions remain over how best to scientifically respond to these events.
The broad-scale factors responsible for viral emergence have been well documented and include human population growth, increased frequency and reach of travel, changing patterns of land use, changing diets, wars and social upheaval and climate change8,9. These factors increase interactions between humans and reservoir hosts, facilitating exposure to zoonotic viruses and spillover infections in people, and allow emerging viruses to spread more easily through human populations. The interactions between virus genetics, ecology and the host factors that determine virus emergence are so complex that it is impossible to predict what virus will cause the next epidemic, making it essential that our response is scientifically informed, robust and efficient10.
The emergence of virus outbreaks generates a set of common questions, whose answers are central to disease mitigation and control (Table 1), and which at times can only be answered by sequencing of viral genomes. These include what is the virus, is it novel, or does it represent the re-emergence of a known pathogen; what is its mode of transmission; where does the emerging virus come from (in particular, what is its reservoir host and/or geographic source); what ecological factors underpin its emergence; how many introductions into humans have there been; what is the timing of these introduction events, and was there a period of undetected transmission before the first reported case; during flare-ups and future outbreaks, how are they connected to previous events; and what is the nature of virus evolution and is there evidence for local adaptation? In the past, many of these of questions were addressed using case (incidence) data, which led to estimates of key epidemic parameters such as the basic reproductive number (R0, the expected number of secondary cases produced by each case at the start of the outbreak) that were used to inform epidemic control policy. Although still of fundamental importance, case data alone cannot inform public health management with the level of precision necessary for all targeted interventions. Recent advances in virus genome sequencing and phylogenetic analyses, however, mean that we are now in a position to answer such questions with molecular precision, and open new areas of investigations not previously possible based on epidemiological data alone (Table 1).
Virus genomics have been used to investigate infectious disease outbreaks for several decades. This is possible because viruses, particularly those with RNA genomes, generate genetic variation on the same timescale of virus transmission, through a combination of high rates of mutation and replication11,12. Consequently, it is possible to infer epidemiological and emergence dynamics from virus genomes sampled and sequenced over short epidemic timescales. We term the science of using genomics and associated analyses ‘genomic epidemiology’.
Initially, genomic approaches relied on indirect methods (for example, restriction fragment length polymorphisms13) to infer genotypes and differentiate between virus strains. As direct sequencing technologies advanced, there was a transition toward the use of nucleotide sequences from fragments of virus genomes for this purpose14,15,16,17,18,19,20,21. Now, thanks to advances in high-throughput sequencing and decreasing costs22, most virus genomics studies utilize data sets containing tens to thousands of (near) complete virus genomes.
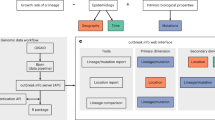
In this Review Article, we will show how our ability to track and understand infectious disease outbreaks have been revolutionized by the addition of virus genomics data. We will highlight the varied uses of virus genomics during the different stages of viral outbreaks, from initial virus detection to understanding the factors contributing towards global spread (Box 1). We will show how genomic epidemiology can be used to track the spread of emerging viruses, where the challenges lie, and establish an agenda for future work. Although we focus on human disease, the genome-based methodologies that we describe can be equally applied to animal and plant infections. Similarly, the increasing ability to rapidly sequence complete genomes of bacterial species means that these technologies offer much to the study of emerging bacterial disease, including those associated with antimicrobial resistance.
Outbreak detection
Most infectious disease outbreaks start with clinicians noticing unusual patterns. Patients may present with patterns of symptoms that are similar to those of more common diseases, but which, after repeated observation and diagnostic testing, may deviate in scale, seasonality or severity. At this very beginning of an outbreak, the most critical task is therefore to identify a causal pathogen. Historically, virus identification has been performed using molecular tools, such as polymerase chain reaction (PCR) and enzyme-linked immunosorbent assay (ELISA), that directly recognize pathogen-derived material (Box 2), or conventional non-molecular techniques, such as microscopy. The advent of untargeted metagenomic sequencing directly from clinical samples, however, means that we are now on the cusp of being able to detect human viruses in a single step, without a priori knowledge of putative causal pathogens (Box 2). The major advantage of sequencing-based approaches is the ability to detect novel viruses—such as the initial appearances of SARS2, MERS3 or Lujo virus23—or unexpected ones, as exemplified by Ebola virus during the 2013–2016 epidemic in West Africa24.
Once an outbreak has been detected and a causal virus identified, several basic questions can immediately be answered about the virus itself, including: (1) whether it is novel or previously known to infect humans; and (2) if we have the diagnostics, vaccines and therapeutics available to fight it. Importantly, the generation of virus genomics data at this stage will provide deeper insights into these questions by uncovering molecular details not possible with conventional tools. Phylogenetics will also provide an additional level of detail, revealing virus origins, evolutionary characteristics and connections to previous outbreaks in the same region, or to transmissions in other regions6. Given high enough relatedness to other members of a virus family with well-defined reservoir hosts (for example, old-world arenaviruses25), the sequence identification of novel virus species can also be informative about potential reservoirs.
First snapshot of an outbreak
Immediately after a viral outbreak has been identified there exists a ‘fog-of-war’. The extent of the outbreak, the timing and nature of its source, and the contribution of human-to-human transmission will be extremely limited, yet these data are critical to designing effective responses. Genomic epidemiology, if applied quickly and comprehensively, holds the potential to answering these questions24.
To provide an initial snapshot of an outbreak, it is important to understand the diversity of circulating viruses from as many cases as possible. Virus genetic diversity, measured as the average number of nucleotide differences among viruses in the population, will increase as an outbreak progresses due to the accumulation of genetic changes in virus genomes at each round of viral replication6. If this rate of mutational accumulation is relatively constant—that is, it conforms to a ‘molecular clock’ of evolutionary change26—then the rate at which it occurs (referred to as the ‘evolutionary rate’) allows us to estimate when the sequenced viruses last shared a common ancestor. Critically, this provides a lower bound on when an outbreak began, and how long the virus had been circulating prior to discovery5,27,28. If the virus genomes have been sampled over only a limited time-scale, so that only a few mutations have accumulated in the virus population, then evolutionary rates will need to be based on those from prior outbreaks or extrapolations from related viruses29. Later in an epidemic, when viruses have been sequenced over a sufficient period of time to capture mutational accumulation, evolutionary rates can be readily estimated directly from virus genomes sampled during the outbreak30,31,32. Evolutionary rate estimates, however, can be sensitive to model specification over short periods of time33 and depend on the timescale of measurement34. Such issues, as well as the unwarranted implications about changes in transmissibility and virulence that may accompany seemingly inflated evolutionary rates, have been discussed in detail in the context of the 2013–2016 Ebola epidemic in West Africa6.
A common approach to phylogenetic analysis of the genetic diversity of a virus population is to infer a tree from sampled virus genomes with branches measured in units of time (that is, a rooted, time-calibrated tree). This can provide estimates of the date of the last common ancestor at the root of the tree, as well as each individual branching event. As an approximation, these branching events correspond to virus transmission from one case to the next, an insight that offers further key information about the unfolding outbreak35. In addition, models of how the process of virus transmission relates to the shape of phylogenetic trees (Fig. 1) enable important epidemiological inferences. In particular, coalescent models relate the rate at which virus lineages of a phylogenetic tree merge, as common ancestors, to the size of the epidemic. This uses the simple premise that, for a sample of virus genomes, the larger the outbreak is, the further back in time the common ancestor will be found (Fig. 1a–c).
The first three scenarios show a single introduction from a non-human reservoir followed by human-to-human spread. a, A small outbreak from a recent zoonosis with a commensurately short tree, suggesting recent emergence. R0 is greater than 1, indicating the potential to cause a large outbreak. b, A medium-sized outbreak with a deeper tree and internal nodes dispersed. With R0 close to 1, this suggests that emergence into humans was not recent and its transmission potential is just sufficient to persist. The root of the tree is not the index case meaning the zoonosis could be older. c, A large outbreak with R0 greater than 1, and thus exhibiting exponential growth in case numbers. Distinctively for a growing epidemic, internal nodes tend to be towards the root of the tree, suggesting that only a small fraction of the total cases were sampled. d, A scenario of repeated zoonotic jumps with limited human to human transmission. The internal parts of the tree represent the diversity of the virus in the non-human reservoir and the human-to-human transmission cases are closely related. Icons courtesy of S. Knemeyer.
Early in an outbreak, one of the primary concerns is to understand the rate at which the virus may be spreading through the human population. As noted in the introduction, this can be assessed by estimating R0, which is critical for epidemiological projections and for planning public health responses. While R0 can be calculated through epidemiological analyses of case counts, accurate estimates of such data may not be available early in an outbreak, since they require a time-series of cases. As demonstrated during the early spread of the novel influenza A/H1N1 virus in 2009, phylogenetic inference of epidemic growth based on virus genomics can provide estimates of R0 comparable to that inferred from case data36. These calculations can be performed using coalescent models that directly estimate R0, based on classic susceptible–infected–recovered (SIR) models37,38. A similar group of models analyse patterns of lineage birth–death, linking the shape of trees to the rate at which virus lineages split and go extinct, and have recently gained popularity39,40. Both approaches were applied during the 2013–2016 epidemic in West Africa to calculate R0 to assess Ebola virus transmission dynamics, and illuminated the impact of ‘superspreader’ events41,42. All of these methods, however, are beholden to the inherent uncertainty of genome sequence data, especially at the start of an epidemic where such sequences exhibit limited variability and sampling may be biased. Hence, phylogenetic estimates of R0, although probably indicative of broad characteristics such as epidemic growth, may not be precise enough to make critical decisions in the absence of corroborating (epidemiological) information.
The initial snapshot of virus genome sequences can also provide critical insights into the role of a zoonotic transmission during an outbreak (Fig. 1d). Genomic analyses, for example, revealed that Lassa fever virus, which is endemic in West Africa43, primarily spreads via repeated transmission from local rodent reservoirs, as opposed to sustained human-to-human transmission44. This is in contrast to Ebola virus during the 2013–2016 epidemic in West Africa, where genomic epidemiology showed that the outbreak was the result of a single zoonotic spillover, followed by sustained human-to-human transmission45.
Given availability of virus genomes from potential zoonotic reservoirs, another aim of early virus sequencing from an outbreak is to uncover the identity and geographic location of the reservoir host. The influenza A/H1N1 pandemic that started in 2009 was quickly recognized as being a likely species jump from pigs, as all of the virus genomic segments closely matched those previously seen in swine4,5. Like the 2013–2016 Ebola epidemic in West Africa, the influenza A/H1N1 pandemic probably started as a single introduction into humans that occurred a few months before it was detected5. The initial suspicion, and later confirmation, that the spillover occurred in Mexico, was complicated by a lack of widespread zoonotic genomic surveillance in this region. Retrospective sequencing of samples from Mexican pigs, however, showed that there were close relatives of the human virus circulating in this country at the time of the epidemic, confirming its origin46.
Transmission chain tracking
Beyond the initial characterization of an outbreak, virus genome sequencing offers enormous potential for determining transmission chains to understand networks of ‘who-infected-whom’. The tracking of transmission chains has long been a standard part of public health responses to outbreaks, providing critical information that can be used to interrupt virus spread and reduce the magnitude of an outbreak. This work has traditionally been performed using interview-based contact tracing, which is labour intensive and limited by the availability and openness of patients for interviews. This approach is particularly challenging during large outbreaks characterized by large numbers of co-occurring transmission chains.
Virus genomic-based approaches can provide much more in-depth information compared to traditional non-sequencing based approaches, as the branching patterns of phylogenetic trees approximately correspond to transmission from one case to the next35 (Fig. 1). Virus genome sequences, for example, were used to reconstruct the spread of foot-and-mouth disease virus in the United Kingdom, including the identification of superspreader events47,48,49. Genomic data also played a critical role in understanding flare-ups during the West African Ebola outbreak50,51,52, where phylogenetic analyses showed that most of the flare-ups were linked to persistently infected Ebola survivors (Fig. 2a), thereby demonstrating sexual transmission of the virus50,52. None of these insights would have been possible without virus genomic data.
a, Viral genome sequences were used to distinguish between competing hypotheses for the source of the viruses that triggered the Ebola flare-ups in West Africa. The three main hypotheses and their expected genomic signatures are illustrated here with a hypothetical haplotype network. Genomes from all of the observed flare-ups grouped closely with genomes sequenced from patients in the same country, from earlier in the outbreak (bottom left), consistent with transmission from persistent sources. In contrast, genomes linked to re-introductions from neighbouring countries (right) would be expected to cluster with genomes from a different country and from late in the outbreak. In the case of independent spillovers from a reservoir host (top left, that is, independent sampling from the diversity circulating within the reservoir), the spillover genomes would be linked to the main outbreak by a long branch originating from near the root of the network. GIN, Guinea; LBR, Liberia; SLE, Sierra Leone. b, Expected ‘genomic resolution’ for the inference of transmission chains at the level of individual infections. Resolution is dependent on the serial interval between infections (x-axis; used as a proxy for epidemiological generation time), as well as the genome size and nucleotide substitution rate (y-axis). Icons courtesy of S. Knemeyer.
The utility of virus genomic data for the inference of transmission chains is dependent on several factors, including: (1) the evolutionary rate of the virus; (2) the length of time between the infections of interest; and (3) the proportion of sampled cases; which together determine the resolution of the genetic signal (Fig. 2b). Although RNA viruses exhibit remarkably high evolutionary rates53, their small genome sizes and short epidemiological generation times often result in, on average, less than one substitution per transmission event54,55,56 (Fig. 2b). Hence, virus genomics alone often cannot be expected to perfectly reconstruct transmission chains at the level of individual infections. Combined with epidemiological data, however, virus genomics provides a powerful tool for restricting the number of possible transmission scenarios and for supporting novel modes of transmission47,57,58. In addition, most phylogenetics-based transmission chain analyses have been performed using virus consensus sequences (that is, a single genome per sample/patient that represents the average of the virus population), which may limit resolution. However, as virus infections exhibit diverse intra-host populations (containing intra-host single nucleotide variants (iSNVs)44), newer methods incorporating viral iSNVs may greatly increase the resolution of transmission chain analyses so long as multiple variants are transmitted between hosts59.
Outbreak mapping
As described in the previous sections, genomic epidemiology can be used to detect an outbreak, show its origin and elucidate transmission patterns. Evolutionary inferences from virus genomes, unlike non-sequencing based methods, can also be used to dissect the spatial structure and dynamics of spread, as well as to assess how an epidemic may unfold through time and space.
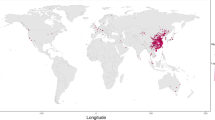
Uncovering the spatial patterns of virus spread during outbreaks is a key objective that has been transformed by genomic epidemiology. Reconstructing a detailed spatial history of virus spread from the origin of an outbreak is generally a task for phylogeographic methods60, which provide location estimates for every ancestral node in a virus phylogeny using simple stochastic (or ‘random walk’) models. Phylogeographic analyses, for example, were used to show how Ebola virus spread across West Africa during the 2013–2016 epidemic61 (Fig. 3). Importantly, virus genome sampling with strong spatiotemporal coverage allowed for the dissection of the entire epidemic into a metapopulation of short- and long-lived transmission chains61. Similar analyses were also used to show that multiple introductions were responsible for sustaining the 2016 Zika outbreak in Florida62. It is important, however, to appreciate the uncertainty of phylogeographic estimates, and to bear in mind that such analyses may only be capable of elucidating partial pictures of outbreak spread. In addition, sampling biases may severely affect these analyses, although the coalescent and birth–death models mentioned above have been extended to account for aspects of virus population structure63,64,65, making the analyses more robust to sampling heterogeneity66.
We illustrate the concept of this approach using the 2013–2016 Ebola epidemic in West Africa. Geographic distances between all pairs of locations, in this case administrative areas in Guinea, Sierra Leone and Liberia, as well as population sizes at the origin and destination of these pairs are combined into a transition rate matrix through a generalized linear model. This matrix parameterizes the phylogenetic process of spread that is being estimated. Each predictor is associated with a coefficient, β, which denotes the strength of contribution with some predictors (for example, population size) positively associated with the intensity of migration whereas others (for example, geographic distance) are negatively associated. A coefficient of 0 implies that the predictor is excluded from the model (represented in the figure by the transparent matrix with β = 0).
Phylogeographic inference methods can also be used to provide insights into the factors driving virus spread67 (Fig. 3). Such analyses are enabled by the integration of virus genomics with diverse meta-data sets and are critically dependent on the timeliness of data generation and open sharing. These approaches were initially introduced to confirm the key role of human air transportation in the global circulation of influenza viruses67, but they have also been useful in untangling complex virus transmission dynamics on smaller scales61. To illustrate these methods, in Fig. 3 we show an application of generalized linear modelling to explain Ebola virus migration rates between locations as a function of several potential predictors, to infer virus spread during West African Ebola outbreak (Fig. 3). In this case, geographic distances and population sizes at the location of origin and destination combine into a gravity model of spread, with virus transmission largely occurring within large population centres and geographic spread being more frequent over shorter distances61. These phylodynamic studies illustrate the growing importance of data integration for virus genomic analyses55, which critically depend on accurate metadata (for example, sampling date and sampling location), as well as other data sources that can capture host mobility and geographic, demographic and epidemiological context.
Inter-epidemic evolution and spread
Once outbreaks have been brought under control or (temporarily) resolved, phylogenetic analyses can provide insights into evolutionary patterns during inter-epidemic periods by comparing virus genome sequences sampled across different outbreaks. The most fundamental question is whether the virus in question has been able to persist in human populations between outbreaks, so that each new outbreak has arisen from an endemically circulating lineage (for example, dengue virus), or whether they represent independent zoonotic spillover events from an animal reservoir (for example, Ebola virus). With sufficient sampling of viruses from human and reservoir species, this question can be answered using standard phylogenetic analysis. For example, although both dengue virus and yellow fever virus have transmission cycles that involve mosquitoes and humans (urban transmission) or nonhuman primates (sylvatic transmission), phylogenetic analyses have shown that dengue virus is now an entirely endemic urban virus that does not rely on its sylvatic vectors and hosts to seed new epidemics68. Most human outbreaks of yellow fever, in contrast, have been shown by virus genomics approaches to represent independent emergences of the virus from sylvatic sources, rather than spread via an urban cycle69,70.
Inter-epidemic analyses can also be used to elucidate the nature of virus evolution and spread in reservoir species, which are probably characterized by different evolutionary forces than those seen during human outbreaks71,72. For example, although human outbreaks of Ebola have happened relatively frequently since the 1970s, each outbreak starts as an independent spillover of the virus from an animal (probably bat73) reservoir. Hence, the inter-epidemic evolution of Ebola virus occurs in a species other than humans, such that patterns of genetic divergence among the viruses associated with human epidemics can provide insight into viral replication and transmission within reservoir hosts. For example, there have been suggestions that Ebola virus has spread across Africa in a wave-like manner in its reservoir species74; however, phylogenetic analyses incorporating virus genomic data from recent outbreaks are incompatible with this scenario6. Additionally, while Ebola virus normally evolves according to a relatively constant molecular clock6,45,75,76,77, the phylogenetic branch leading to the viruses sequenced from the small Ebola outbreak that occurred in the Democratic Republic of the Congo in 2014, concurrent with the 2013–2016 epidemic in West Africa, was characterized by a far lower evolutionary rate78. Although the reasons for this reduction in evolutionary tempo are unclear, it is possible that it reflects Ebola virus evolution in a different (unknown) reservoir species that experiences a lower rate of viral replication. Alternatively, this rate disparity may result from the existence of different viral replication states within the same reservoir host, similar to that described during human epidemics, with faster rates observed during continuous human-to-human transmission and slower rates during persistent infections of Ebola survivors79.
Requirements and challenges in genomic epidemiology
Virus genomic methods for outbreak investigation and control are powerful additions to more traditional epidemiological approaches but are critically dependent on well planned and coordinated efforts. The foremost need for genomic epidemiology is timely access to clinical samples and data, which should be built on productive and equitable collaborations with local communities, public health agencies, outbreak responders, local clinics and researchers80. For each clinical sample to be used for virus genomic sequencing, it is essential to obtain a minimal set of metadata related to the infection, including: (1) the date of sample collection and/or onset of symptoms; and (2) the location of sampling. Additional information can greatly increase the utility of genomic epidemiology, including the availability of: (3) travel and contact history; (4) suspected source of infection; and (5) clinical outcome and symptoms. Other factors, including patient history, age, sex and economic status can also help to reveal risk factors underlying infection and transmission. Within ethical constraints, it is important that communication lines remain open so that researchers undertaking data analysis can return actionable results to the public health community.
Other large-scale data resources are essential for investigating the spatio-temporal history and spread of an outbreak. These include the temporal and spatial distribution of cases, ecological conditions, vector abundance, environmental factors and travel patterns. Integration of these other data sources with virus genomic data may reveal new properties of an outbreak, potentially leading to actionable measures55,61,67. Non-genomic data often comes from established networks of collaborations, or from the public domain, highlighting the value of open data and data sharing to outbreak investigations.
An important benefit of genomic epidemiology is that it can directly compare and jointly analyse virus genome sequences obtained during an epidemic, even if those sequences were generated by different laboratories. Consequently, there is an urgent need to make genomic and epidemiological data and analysis tools publically available during ongoing epidemics81. This movement is supported by the World Health Organization (WHO), which has called for data pertaining to public health emergencies to be disseminated openly and immediately following generation, and not withheld until the acceptance or publication of a corresponding scientific paper82. More recently, the WHO has outlined the current and future benefits of virus genome data sharing during outbreaks83. Combined with an acceleration of making manuscripts available via preprint servers such as arXiv and bioRxiv, especially during outbreaks84, there has been a shift towards scientists storing their data and source code on depositories such as GitHub (https://www.github.com), Synapse (https://www.synapse.org) and Data Dryad (https://www.datadryad.org), in close to real-time for others to use. Furthermore, extensive online communities and forums such as Twitter (https://www.twitter.com), Virological (http://virological.org), FluTrackers (https://www.flutrackers.com), ProMED (https://www.promedmail.org), Nextstrain (https://www.nextstrain.org), HealthMap (https://www.healthmap.org) and Microreact (https://www.microreact.org) allow rapid dissemination of unpublished results and analyses. In our experience, not only does the process of open science promote new collaborations and lead to more accurate scientific insights into outbreak research, but it helps in getting relevant information rapidly into the hands of decision makers. Despite these advances, however, the speed, nature and extent of virus genome data sharing is inconsistent, sometimes resulting in confusion over what is, or should be, best practice81,85.
Future perspective
Genomic epidemiology promises much to the study and control of infectious disease outbreaks, particularly if viral genomes can be acquired and analysed in real-time. The accumulated set of these data—together with the rapid development of sophisticated software packages (http://virological.org/c/software)—will provide a valuable resolve for the mitigation and control of future outbreaks. Ultimately, with sufficient genome sequences from individual viral genera and/or families, it may be possible to categorize viruses by their phylogenetic patterns and utilize this information in epidemic preparedness. For example, as well as considering obvious biological features of viruses such as their genome structure and mode of transmission, it may be possible to group viruses according to a series of evolutionary variables such as rate of evolutionary change, extent of antigenic evolution, frequency of recombination, pattern of geographic spread and population dynamics. This information may then help forecast the evolutionary behaviour of any virus, should it re-emerge in human populations, and assist in the selection of future vaccine strains86,87,88. This information will also help counter the alarmist claims that emerging viruses will evolve novel phenotypes, such as airborne transmission in the case of Ebola virus89, that often accompany any major disease outbreak. It is clear, however, that a more fundamental understanding of the genetic and ecological barriers of virus spillover into human populations is needed to better identify risk factors for disease emergence. Long-term capacity building, partnerships with local communities, and commitments to long-term investments on these fronts will go a long way towards better enabling the global community to effectively and rapidly deal with future emerging outbreaks80.
References
Drosten, C. et al. Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N. Engl. J. Med. 348, 1967–1976 (2003).
Ksiazek, T. G. et al. A novel coronavirus associated with severe acute respiratory syndrome. N. Engl. J. Med. 348, 1953–1966 (2003).
Zaki, A. M., van Boheemen, S., Bestebroer, T. M., Osterhaus, A. D. M. E. & Fouchier, R. A. M. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N. Engl. J. Med. 367, 1814–1820 (2012).
Novel Swine-Origin Influenza A (H1N1) Virus Investigation Team et al. Emergence of a novel swine-origin influenza A (H1N1) virus in humans. N. Engl. J. Med. 360, 2605–2615 (2009).
Smith, G. J. D. et al. Origins and evolutionary genomics of the 2009 swine-origin H1N1 influenza A epidemic. Nature 459, 1122–1125 (2009).
Holmes, E. C., Dudas, G., Rambaut, A. & Andersen, K. G. The evolution of Ebola virus: insights from the 2013–2016 epidemic. Nature 538, 193–200 (2016).
Grubaugh, N. D., Faria, N. R., Andersen, K. G. & Pybus, O. G. Genomic insights into Zika virus emergence and spread. Cell 172, 1160–1162 (2018).
Morse, S. S. in Plagues and Politics (ed. Mullan, F.) 8–26 (Palgrave Macmillan, London, 2001).
Wolfe, N. D., Dunavan, C. P. & Diamond, J. Origins of major human infectious diseases. Nature 447, 279–283 (2007).
Holmes, E. C., Rambaut, A. & Andersen, K. G. Pandemics: spend on surveillance, not prediction. Nature 558, 180–182 (2018).
Holland, J. et al. Rapid evolution of RNA genomes. Science 215, 1577–1585 (1982).
Duffy, S., Shackelton, L. A. & Holmes, E. C. Rates of evolutionary change in viruses: patterns and determinants. Nat. Rev. Genet. 9, 267–276 (2008).
Kiko, H., Niggemann, E. & Rüger, W. Physical mapping of the restriction fragments obtained from bacteriophage T4 dC-DNA with the restriction endonucleases SmaI, KpnI and BglII. Mol. Gen. Genet. 172, 303–312 (1979).
Chungue, E., Deubel, V., Cassar, O., Laille, M. & Martin, P. M. Molecular epidemiology of dengue 3 viruses and genetic relatedness among dengue 3 strains isolated from patients with mild or severe form of dengue fever in French Polynesia. J. Gen. Virol. 74, 2765–2770 (1993).
Lanciotti, R. S. et al. Origin of the West Nile virus responsible for an outbreak of encephalitis in the northeastern United States. Science 286, 2333–2337 (1999).
Kinnunen, L., Pöyry, T. & Hovi, T. Generation of virus genetic lineages during an outbreak of poliomyelitis. J. Gen. Virol. 72, 2483–2489 (1991).
McNearney, T. et al. Limited sequence heterogeneity among biologically distinct human immunodeficiency virus type 1 isolates from individuals involved in a clustered infectious outbreak. Proc. Natl Acad. Sci. USA 87, 1917–1921 (1990).
Nichol, S. T. et al. Genetic identification of a hantavirus associated with an outbreak of acute respiratory illness. Science 262, 914–917 (1993).
Ou, C. Y. et al. Molecular epidemiology of HIV transmission in a dental practice. Science 256, 1165–1171 (1992).
Power, J. P. et al. Molecular epidemiology of an outbreak of infection with hepatitis C virus in recipients of anti-D immunoglobulin. Lancet 345, 1211–1213 (1995).
Rossouw, E., Tsilimigras, C. W. & Schoub, B. D. Molecular epidemiology of a coxsackievirus B3 outbreak. J. Med. Virol. 34, 165–171 (1991).
Shendure, J. & Ji, H. Next-generation DNA sequencing. Nat. Biotechnol. 26, 1135–1145 (2008).
Briese, T. et al. Genetic detection and characterization of Lujo virus, a new hemorrhagic fever-associated arenavirus from southern Africa. PLoS Pathog. 5, e1000455 (2009).
Gardy, J. L. & Loman, N. J. Towards a genomics-informed, real-time, global pathogen surveillance system. Nat. Rev. Genet. 19, 9–20 (2018).
Salazar-Bravo, J., Ruedas, L. A. & Yates, T. L. Mammalian reservoirs of arenaviruses. Curr. Top. Microbiol. Immunol. 262, 25–63 (2002).
dos Reis, M., Donoghue, P. C. J. & Yang, Z. Bayesian molecular clock dating of species divergences in the genomics era. Nat. Rev. Genet. 17, 71–80 (2016).
Rambaut, A. & Holmes, E. The early molecular epidemiology of the swine-origin A/H1N1 human influenza pandemic. PLoS Curr. 1, RRN1003 (2009).
Korber, B. Timing the ancestor of the HIV-1 pandemic strains. Science 288, 1789–1796 (2000).
Cotten, M. et al. Full-genome deep sequencing and phylogenetic analysis of novel human betacoronavirus. Emerg. Infect. Dis. 19, 736–42B (2013).
Rambaut, A. Estimating the rate of molecular evolution: incorporating non-contemporaneous sequences into maximum likelihood phylogenies. Bioinformatics 16, 395–399 (2000).
Drummond, A., Pybus, O. G. & Rambaut, A. Inference of viral evolutionary rates from molecular sequences. Adv. Parasitol. 54, 331–358 (2003).
Drummond, A. J., Nicholls, G. K., Rodrigo, A. G. & Solomon, W. Estimating mutation parameters, population history and genealogy simultaneously from temporally spaced sequence data. Genetics 161, 1307–1320 (2002).
Möller, S., du Plessis, L. & Stadler, T. Impact of the tree prior on estimating clock rates during epidemic outbreaks. Proc. Natl Acad. Sci. USA 115, 4200–4205 (2018).
Duchêne, S., Holmes, E. C. & Ho, S. Y. W. Analyses of evolutionary dynamics in viruses are hindered by a time-dependent bias in rate estimates. Proc. Biol. Sci. 281, 20140732 (2014).
Hall, M. D., Woolhouse, M. E. J. & Rambaut, A. Using genomics data to reconstruct transmission trees during disease outbreaks. Rev. Sci. Tech. 35, 287–296 (2016).
Fraser, C. et al. Pandemic potential of a strain of influenza A (H1N1): early findings. Science 324, 1557–1561 (2009).
Volz, E. M., Kosakovsky Pond, S. L., Ward, M. J., Leigh Brown, A. J. & Frost, S. D. W. Phylodynamics of infectious disease epidemics. Genetics 183, 1421–1430 (2009).
Rasmussen, D. A., Ratmann, O. & Koelle, K. Inference for nonlinear epidemiological models using genealogies and time series. PLoS Comput. Biol. 7, e1002136 (2011).
Stadler, T. et al. Estimating the basic reproductive number from viral sequence data. Mol. Biol. Evol. 29, 347–357 (2012).
Kühnert, D., Stadler, T., Vaughan, T. G. & Drummond, A. J. Simultaneous reconstruction of evolutionary history and epidemiological dynamics from viral sequences with the birth-death SIR model. J. R. Soc. Interface 11, 20131106 (2014).
Stadler, T., Kühnert, D., Rasmussen, D. A. & du Plessis, L. Insights into the early epidemic spread of Ebola in Sierra Leone provided by viral sequence data. PLoS Curr. https://doi.org/10.1371/currents.outbreaks.02bc6d927ecee7bbd33532ec8ba6a25f (2014).
Volz, E. & Pond, S. Phylodynamic analysis of Ebola virus in the 2014 Sierra Leone epidemic. PLoS Curr. https://doi.org/10.1371/currents.outbreaks.6f7025f1271821d4c815385b08f5f80e (2014).
McCormick, J. B. & Fisher-Hoch, S. P. Lassa fever. Curr. Top. Microbiol. Immunol. 262, 75–109 (2002).
Andersen, K. G. et al. Clinical sequencing uncovers origins and evolution of Lassa virus. Cell 162, 738–750 (2015).
Gire, S. K. et al. Genomic surveillance elucidates Ebola virus origin and transmission during the 2014 outbreak. Science 345, 1369–1372 (2014).
Mena, I. et al. Origins of the 2009 H1N1 influenza pandemic in swine in Mexico. eLife 5, e16777 (2016).
Morelli, M. J. et al. A Bayesian inference framework to reconstruct transmission trees using epidemiological and genetic data. PLoS Comput. Biol. 8, e1002768 (2012).
Cottam, E. M. et al. Integrating genetic and epidemiological data to determine transmission pathways of foot-and-mouth disease virus. Proc. Biol. Sci. 275, 887–895 (2008).
Cottam, E. M. et al. Molecular epidemiology of the foot-and-mouth disease virus outbreak in the United Kingdom in 2001. J. Virol. 80, 11274–11282 (2006).
Mate, S. E. et al. Molecular evidence of sexual transmission of Ebola virus. N. Engl. J. Med. 373, 2448–2454 (2015).
Blackley, D. J. et al. Reduced evolutionary rate in re-emerged Ebola virus transmission chains. Sci. Adv. 2, e1600378 (2016).
Diallo, B. et al. Resurgence of Ebola virus disease in Guinea linked to a survivor with virus persistence in seminal fluid for more than 500 days. Clin. Infect. Dis. 63, 1353–1356 (2016).
Duffy, S., Shackelton, L. A. & Holmes, E. C. Rates of evolutionary change in viruses: patterns and determinants. Nat. Rev. Genet. 9, 267–276 (2008).
Biek, R., Pybus, O. G., Lloyd-Smith, J. O. & Didelot, X. Measurably evolving pathogens in the genomic era. Trends Ecol. Evol. 30, 306–313 (2015).
Baele, G., Suchard, M. A., Rambaut, A. & Lemey, P. Emerging concepts of data integration in pathogen phylodynamics. Syst. Biol. 66, e47–e65 (2017).
Campbell, F., Strang, C., Ferguson, N., Cori, A. & Jombart, T. When are pathogen genome sequences informative of transmission events? PLoS Pathog. 14, e1006885 (2018).
Mate, S. E. et al. Molecular evidence of sexual transmission of Ebola virus. N. Engl. J. Med. 373, 2448–2454 (2015).
Resik, S. et al. Limitations to contact tracing and phylogenetic analysis in establishing HIV type 1 transmission networks in Cuba. AIDS Res. Hum. Retroviruses 23, 347–356 (2007).
Worby, C. J., Lipsitch, M. & Hanage, W. P. Shared genomic variants: identification of transmission routes using pathogen deep-sequence data. Am. J. Epidemiol. 186, 1209–1216 (2017).
Faria, N. R., Suchard, M. A., Rambaut, A. & Lemey, P. Toward a quantitative understanding of viral phylogeography. Curr. Opin. Virol. 1, 423–429 (2011).
Dudas, G. et al. Virus genomes reveal factors that spread and sustained the Ebola epidemic. Nature 544, 309–315 (2017).
Grubaugh, N. D. et al. Genomic epidemiology reveals multiple introductions of Zika virus into the United States. Nature 90, 4864 (2017).
Vaughan, T. G., Kühnert, D., Popinga, A., Welch, D. & Drummond, A. J. Efficient Bayesian inference under the structured coalescent. Bioinformatics 30, 2272–2279 (2014).
Müller, N. F., Rasmussen, D. A. & Stadler, T. The structured coalescent and its approximations. Mol. Biol. Evol. 34, 2970–2981 (2017).
Kühnert, D., Stadler, T., Vaughan, T. G. & Drummond, A. J. Phylodynamics with migration: a computational framework to quantify population structure from genomic data. Mol. Biol. Evol. 33, 2102–2116 (2016).
De Maio, N., Wu, C.-H., O’Reilly, K. M. & Wilson, D. New routes to phylogeography: a bayesian structured coalescent approximation. PLoS Genet. 11, e1005421 (2015).
Lemey, P. et al. Unifying viral genetics and human transportation data to predict the global transmission dynamics of human influenza H3N2. PLoS Pathog. 10, e1003932 (2014).
Wang, E. et al. Evolutionary relationships of endemic/epidemic and sylvatic dengue viruses. J. Virol. 74, 3227–3234 (2000).
Cardoso, J. & da, C. et al. Yellow fever virus in Haemagogus leucocelaenus and Aedes serratus mosquitoes, southern Brazil, 2008. Emerg. Infect. Dis. 16, 1918–1924 (2010).
Faria, N. R. et al. Genomic and epidemiological monitoring of yellow fever virus transmission potential. Science 361, 894–899 (2018).
Diehl, W. E. et al. Ebola virus glycoprotein with increased infectivity dominated the 2013–2016 epidemic. Cell 167, 1088–1097 (2016).
Urbanowicz, R. A. et al. Human adaptation of Ebola virus during the West African outbreak. Cell 167, 1079–1085 (2016).
Leroy, E. M. et al. Fruit bats as reservoirs of Ebola virus. Nature 438, 575–576 (2005).
Walsh, P. D., Biek, R. & Real, L. A. Wave-like spread of Ebola Zaire. PLoS Biol. 3, e371 (2005).
Carroll, S. A. et al. Molecular evolution of viruses of the family Filoviridae based on 97 whole-genome sequences. J. Virol. 87, 2608–2616 (2013).
Dudas, G. & Rambaut, A. Phylogenetic analysis of Guinea 2014 EBOV Ebolavirus outbreak. PLoS Curr. https://doi.org/10.1371/currents.outbreaks.84eefe5ce43ec9dc0bf0670f7b8b417d (2014).
Rambaut, A. et al. Comment on ‘Mutation rate and genotype variation of Ebola virus from Mali case sequences’. Science 353, 658 (2016).
Lam, T. T.-Y., Zhu, H., Chong, Y. L., Holmes, E. C. & Guan, Y. Puzzling origins of the Ebola outbreak in the Democratic Republic of the Congo, 2014. J. Virol. 89, 10130–10132 (2015).
Blackley, D. J. et al. Reduced evolutionary rate in reemerged Ebola virus transmission chains. Sci. Adv. 2, e1600378 (2016).
Yozwiak, N. L. et al. Roots, not parachutes: research collaborations combat outbreaks. Cell 166, 5–8 (2016).
Yozwiak, N. L., Schaffner, S. F. & Sabeti, P. C. Data sharing: make outbreak research open access. Nature 518, 477–479 (2015).
WHO. Policy statement on data sharing by WHO in the context of public health emergencies (as of 13 April 2016). Wkly. Epidemiol. Rec. 91, 237–240 (2016).
WHO R&D Blueprint Meeting on Pathogen Genetic Sequence Data (GSD) Sharing in the Context of Public Health Emergencies, 28-29 September 2017 (WHO, 2017).
Johansson, M. A., Reich, N. G., Meyers, L. A. & Lipsitch, M. Preprints: an underutilized mechanism to accelerate outbreak science. PLoS Med. 15, e1002549 (2018).
Callaway, E. Zika-microcephaly paper sparks data-sharing confusion. Nature News https://doi.org/10.1038/nature.2016.19367 (2016).
Luksza, M. & Lässig, M. A predictive fitness model for influenza. Nature 507, 57–61 (2014).
Smith, D. J. et al. Mapping the antigenic and genetic evolution of influenza virus. Science 305, 371–376 (2004).
Neher, R. A., Bedford, T., Daniels, R. S., Russell, C. A. & Shraiman, B. I. Prediction, dynamics, and visualization of antigenic phenotypes of seasonal influenza viruses. Proc. Natl Acad. Sci. USA 113, E1701–9 (2016).
Osterholm, M. T. et al. Transmission of Ebola viruses: what we know and what we do not know. mBio 6, e00137 (2015).
Sabir, J. S. M. et al. Co-circulation of three camel coronavirus species and recombination of MERS-CoVs in Saudi Arabia. Science 351, 81–84 (2016).
Dudas, G., Carvalho, L. M., Rambaut, A. & Bedford, T. MERS-CoV spillover at the camel-human interface. eLife 7, (2018).
Faria, N. R. et al. Zika virus in the Americas: early epidemiological and genetic findings. Science 352, 345–349 (2016).
Faria, N. R. et al. Establishment and cryptic transmission of Zika virus in Brazil and the Americas. Nature 546, 406–410 (2017).
Metsky, H. C. et al. Zika virus evolution and spread in the Americas. Nature 66, 366 (2017).
Christie, A. et al. Possible sexual transmission of Ebola virus — Liberia, 2015. MMWR Morb. Mortal. Wkly. Rep. 64, 479–481 (2015).
Whitmer, S. L. M. et al. Active Ebola virus replication and heterogeneous evolutionary rates in EVD survivors. Cell Rep. 22, 1159–1168 (2018).
Dietzel, E., Schudt, G., Krähling, V., Matrosovich, M. & Becker, S. Functional characterization of adaptive mutations during the West African Ebola virus outbreak. J. Virol. 91, e01913–16 (2017).
List of Blueprint Priority Diseases (WHO, 2018); https://www.who.int/blueprint/priority-diseases/en/
Boisen, M. L. et al. Field validation of the ReEBOV antigen rapid test for point-of-care diagnosis of Ebola virus infection. J. Infect. Dis. 214, S203–S209 (2016).
Broadhurst, M. J. et al. ReEBOV antigen rapid test kit for point-of-care and laboratory-based testing for Ebola virus disease: a field validation study. Lancet 386, 867–874 (2015).
Chotiwan, N. et al. Rapid and specific detection of Asian- and African-lineage Zika viruses. Sci. Transl. Med. 9, eaag0538 (2017).
Imai, M. et al. Development of H5-RT-LAMP (loop-mediated isothermal amplification) system for rapid diagnosis of H5 avian influenza virus infection. Vaccine 24, 6679–6682 (2006).
Hong, T. C. T. et al. Development and evaluation of a novel loop-mediated isothermal amplification method for rapid detection of severe acute respiratory syndrome coronavirus. J. Clin. Microbiol. 42, 1956–1961 (2004).
Hattersley, S. M., Greenman, J. & Haswell, S. J. The application of microfluidic devices for viral diagnosis in developing countries. Methods Mol. Biol. 949, 285–303 (2013).
Patolsky, F. et al. Electrical detection of single viruses. Proc. Natl Acad. Sci. USA 101, 14017–14022 (2004).
Chen, Y. et al. Field-effect transistor biosensor for rapid detection of Ebola antigen. Sci. Rep. 7, 10974 (2017).
Afsahi, S. et al. Novel graphene-based biosensor for early detection of Zika virus infection. Biosens. Bioelectron. 100, 85–88 (2018).
Pardee, K. et al. Paper-based synthetic gene networks. Cell 159, 940–954 (2014).
Gootenberg, J. S. et al. Nucleic acid detection with CRISPR-Cas13a/C2c2. Science 356, 438–442 (2017).
Myhrvold, C. et al. Field-deployable viral diagnostics using CRISPR-Cas13. Science 360, 444–448 (2018).
Gootenberg, J. S. et al. Multiplexed and portable nucleic acid detection platform with Cas13, Cas12a, and Csm6. Science 360, 439–444 (2018).
Chen, J. S. et al. CRISPR-Cas12a target binding unleashes indiscriminate single-stranded DNase activity. Science 360, 436–439 (2018).
Gu, W. et al. Depletion of Abundant Sequences by Hybridization (DASH): using Cas9 to remove unwanted high-abundance species in sequencing libraries and molecular counting applications. Genome Biol. 17, 41 (2016).
Siddle, K. J. et al. Capturing diverse microbial sequence with comprehensive and scalable probe design. bioRxiv https://doi.org/10.1101/279570 (2018).
Matranga, C. B. et al. Enhanced methods for unbiased deep sequencing of Lassa and Ebola RNA viruses from clinical and biological samples. Genome Biol. 15, 519 (2014).
Quick, J. et al. Multiplex PCR method for MinION and Illumina sequencing of Zika and other virus genomes directly from clinical samples. Nat. Protoc. 12, 1261–1276 (2017).
Quick, J. et al. Real-time, portable genome sequencing for Ebola surveillance. Nature 530, 228–232 (2016).
Acknowledgements
We thank G. Dudas and S. Knemeyer for help with figure creation. N.D.G. is supported by NIH training grant 5T32AI007244-33. P.L. and A.R. acknowledge funding from the European Research Council under the European Union’s Horizon 2020 research and innovation programme (grant agreement no. 725422-ReservoirDOCS) and from the Wellcome Trust Collaborative Award (grant number 206298/Z/17/Z—ARTICnetwork). P.L. acknowledges support by the Research Foundation—Flanders (‘Fonds voor Wetenschappelijk Onderzoek - Vlaanderen’, G066215N, G0D5117N and G0B9317N). O.G.P. is supported by the European Union’s Seventh Framework Programme (FP7/2007-2013)/European Research Council (614725-PATHPHYLODYN) and by the Oxford Martin School. E.C.H. is supported by an ARC Australian Laureate Fellowship (FL170100022). K.G.A. is a Pew Biomedical Scholar, and is supported by NIH NCATS CTSA UL1TR002550, NIAID contract HHSN272201400048C, NIAID R21AI137690, NIAID U19AI135995, and The Ray Thomas Foundation.
Author information
Authors and Affiliations
Contributions
All listed authors have contributed to the conceptualization, writing and preparation of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Grubaugh, N.D., Ladner, J.T., Lemey, P. et al. Tracking virus outbreaks in the twenty-first century. Nat Microbiol 4, 10–19 (2019). https://doi.org/10.1038/s41564-018-0296-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-018-0296-2
This article is cited by
-
Emerging and re-emerging pediatric viral diseases: a continuing global challenge
Pediatric Research (2024)
-
Serological evidence of arenavirus circulation in wild rodents from central-west, southeast, and south regions of Brazil, 2002–2006
Brazilian Journal of Microbiology (2023)
-
Increased interregional virus exchange and nucleotide diversity outline the expansion of chikungunya virus in Brazil
Nature Communications (2023)
-
Mapping the viruses belonging to the order Bunyavirales in China
Infectious Diseases of Poverty (2022)
-
Metagenomics-enabled microbial surveillance
Nature Microbiology (2022)