Abstract
The intestinal microbiota is considered to be a major reservoir of antibiotic resistance determinants (ARDs) that could potentially be transferred to bacterial pathogens via mobile genetic elements. Yet, this assumption is poorly supported by empirical evidence due to the distant homologies between known ARDs (mostly from culturable bacteria) and ARDs from the intestinal microbiota. Consequently, an accurate census of intestinal ARDs (that is, the intestinal resistome) has not yet been fully determined. For this purpose, we developed and validated an annotation method (called pairwise comparative modelling) on the basis of a three-dimensional structure (homology comparative modelling), leading to the prediction of 6,095 ARDs in a catalogue of 3.9 million proteins from the human intestinal microbiota. We found that the majority of predicted ARDs (pdARDs) were distantly related to known ARDs (mean amino acid identity 29.8%) and found little evidence supporting their transfer between species. According to the composition of their resistome, we were able to cluster subjects from the MetaHIT cohort (n = 663) into six resistotypes that were connected to the previously described enterotypes. Finally, we found that the relative abundance of pdARDs was positively associated with gene richness, but not when subjects were exposed to antibiotics. Altogether, our results indicate that the majority of intestinal microbiota ARDs can be considered intrinsic to the dominant commensal microbiota and that these genes are rarely shared with bacterial pathogens.
Similar content being viewed by others
Main
Antimicrobial resistance is one of the major threats to health identified by the World Health Organization for the coming decades1. The intestinal microbiota plays a pivotal role in this phenomenon as it harbours a vast diversity of bacterial species, some of them possessing antibiotic resistance determinants (ARDs) that may enable their survival under antibiotic exposure. Previous studies attempted to identify ARDs in the intestinal microbiota2,3,4 but were confounded by the distant homologies between known ARDs (mostly from culturable bacteria) and ARDs from the intestinal microbiota (which are generally not cultured)5,6. For these reasons, bioinformatic tools based on sequence comparison (ARG-ANNOT7, CARD8, Resfinder9, DeepARG10) or motif detection (Resfams11) are often unsuccessful in characterizing the diversity of ARDs from metagenomic datasets. Indeed, there is no consensus on an optimal approach to detect ARDs in metagenomic datasets. Consequently, an accurate census of intestinal ARDs (that is, the intestinal resistome12) has not yet been fully determined.
While many bacteria have intrinsic, chromosomally encoded ARDs and the capability of increasing resistance through mutation, they can also enrich their resistance capabilities through the acquisition of exogenous ARDs located on mobile genetic elements (MGEs) such as plasmids, transposons or phages. The intestinal microbiota harbours thousands of bacterial species including well-known pathogens (for example, Enterobacteriaceae and Enterococcus spp.). This unique environment is assumed to be a reservoir of ARDs that can potentially be transferred to bacterial pathogens13. Nonetheless, despite the high selective pressure exerted on the intestinal microbiota by over seven decades of intensive antibiotic usage, a very low number of transfer events from an intestinal commensal to a bacterial pathogen have been observed14,15. This challenges the hypothesis of a mobile resistome and the assumption that the intestinal microbiota serves as a reservoir of ARDs to which pathogenic bacteria have easy access16. In this study, our objective was to perform an extensive characterization of the human gut resistome (including the capacity of ARDs to transfer between species) and to assess its dynamics under various antibiotic exposures.
Prediction of ARDs in the intestinal microbiota
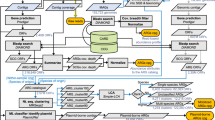
To predict ARDs in the intestinal microbiota, we developed a method based on protein homology modelling (see Methods) that we termed pairwise comparative modelling (PCM). PCM is a generic method using homology modelling to increase the specificity of functional prediction of proteins, especially when they are distantly related to potential homologues. PCM uses a list of reference proteins sequences from a given family, the ARD structures of this family (used as structural templates in protein data bank (PDB) format) and a series of negative references (Fig. 1a and Supplementary Figs. 1–3). Structural models are built using both the ARD reference and negative reference templates. Scores generated from both positive and negative references are used to determine which model performed the best. This is done using a machine-learning algorithm trained on 662 ARDs and 522 negative references. The PCM score equals the number of times the query was classified as an ARD for the bootstraps performed, expressed as a percentage. Candidates with a PCM score ≥50% and an alignment score with the reference template (TM score given by TM-Align) ≥0.517 were predicted as ARDs.
a, Left: class A β-lactamase protein structure (4EWF) obtained from the PDB database. Middle: a candidate protein (MC3.MG12.AS1.GP1.C14.G3 from Faecalibacterium prausnitzii) for class A β-lactamase modelled with a reference class A β-lactamase structural template. This protein had 26.5% amino acid identity with the closest reference class A β-lactamase. Right: the same candidate protein (MC3.MG12.AS1.GP1.C14.G3) for class A β-lactamase this time modelled with a negative reference template. The candidate MC3.MG12.AS1.GP1.C14.G3 was predicted to be a class A β-lactamase with 100% confidence by our model and later found to be functional after gene synthesis. b, Bar plot of the activity of the synthesized pdARDs against antibiotics with respect to the degree of confidence of the prediction (‘reference’ meaning that the protein shares more than ≥95% amino acid identity with a functionally proven ARD, ‘good’ meaning a PCM score over 99% and a TM-Align TM score ≥0.8, ‘fair’ meaning a PCM score between 50 and 80%). c, Number of predictions of ARDs from a 3.9 million gene catalogue of the intestinal microbiota19 using PCM, BLASTP21, ARG-ANNOT7, Resfinder9, DeepARG10, Resfams11 and the HMM-based method published by Berglund et al. for class B1 β-lactamases22. d, Violin plot of the maximal identity observed with a reference ARD for candidates predicted as ARDs (blue violin, n = 6,095) and those not predicted as ARDs (red violin, n = 3,982). The point depicts the median. The width of the violins depicts the distribution of pdARDs according to their maximal identity with a reference ARD. See Supplementary Table 2 for details about candidates sharing at least 40% identity with reference ARDs but that were not predicted as ARDs. Bla, β-lactamase; AAC, aminoglycoside acetylase; ANT, aminoglycoside nucleotidyl transferase; APH, aminoglycoside phosphotransferase; DfrA, type A dihydrofolate reductase; Sul, dihydropteroate synthase; Erm, erythromycin ribosome methylase; Qnr, quinolone resistance; Fos, fosfomycin resistance; Van, D-Ala—D-Lac/Ser ligase (vancomycin resistance).
The performance of PCM to predict ARDs was assessed using in vitro and in silico methods. We synthesized 71 candidate ARDs from 12 families (Table 1) and expressed them in Escherichia coli (see Methods). All 12 predicted ARDs (pdARDs) sharing an amino acid identity >95% with a known ARD had a detectable resistance activity against antibiotics (Fig. 1b). Resistance activity was also detected in 35 out of 41 (85.3%) of the predictions made with a good level of confidence (PCM score >99%, TM score TM-Align >0.9) and in 8 out of 18 (44.4%) of the predictions with a lower level of confidence (PCM score <80%, TM score TM-Align <0.8). The mean amino acid identity of the functional pdARDs (good and fair predictions, n = 43) with known ARDs was 28.6% (range 19.4–82.6%, Supplementary Table 1). We then tested PCM against an experimentally validated functional metagenomics dataset from soils18. In this case, PCM was able to accurately identify 1,374 ARDs out of 1,423 hits (sensitivity 96.6%) (see Methods). Finally, we assessed the performances of PCM with incomplete proteins as inputs, and showed that PCM could correctly predict ARDs when the available amino acid sequence was at least 40% complete (Supplementary Fig. 4). After the in vitro and in silico validation of the method, we used PCM to search for ARDs in a catalogue made of 3,871,657 proteins that was built from the sequencing of faecal samples of 396 human individuals (177 Danes and 219 Spanish) recruited in the MetaHIT project19. In total, we predicted 6,095 ARDs (0.2% of the catalogue) from 20 ARD classes conferring resistance to nine major antibiotic families20: β-lactams (class A, B1–B2, B3, C and D β-lactamases), aminoglycosides (AAC(2’), AAC(3)-I, AAC(3)-II, AAC(6’), ANT, APH, 16S rRNA methylases), tetracyclines (Tet(M), Tet(X)), macrolides (Erm), quinolones (Qnr), sulfonamides (Sul), trimethoprim (DfrA), fosfomycin (Fos) and glycopeptides (Van ligases) (Table 1 and Supplementary Table 1). With the same, extensively curated reference ARDs census as input, only 67 ARDs would have been predicted according to conventional BLASTP21 search with a specific identity threshold (80% over 80% of the reference sequence)3,4. ARG-ANNOT7, Resfinder9 and DeepARG10 were able to predict 54, 50 and 2,139 ARDs, respectively, while Resfams11 predicted a very high number of ARDs (n = 44,105). The HMM-based search for class B1 β-lactamases published by Berglund et al.22 also yielded a high number of hits (n = 3,490) in the 3.9 million protein catalogue (Fig. 1c and Supplementary Fig. 5). Further analysis on a catalogue of dummy, synthetic 3.9 million proteins indeed showed that Resfams, DeepARG and the Berglund et al. HMM-based search lacked specificity (see Supplementary Information). The mean identity shared between predicted (n = 6,095) and reference ARDs was 29.8%; it was significantly higher than candidates not predicted as ARDs (mean 23.0%, Wilcoxon unpaired test P = 2 × 10–16, Fig. 1d). Indeed, most of the pdARDs were distantly related to reference ARDs (Supplementary Figs. 6 and 7). Besides, PCM failed to predict 16 ARDs that shared at least 40% identity with a reference ARD (Supplementary Table 2). The 6,095 pdARDs and their structures are available at http://mgps.eu/Mustard.
Taxonomic distribution of ARDs
A host bacterial phylum could be assigned to 72.3% (4,405 out of 6,095) pdARDs. The majority was identified as from the dominant human intestinal phyla Firmicutes (2,962 out of 4,405, 72.3%) and Bacteroidetes (858 out of 4,405, 19.5%) (Supplementary Fig. 8) with only 5.8% (225 out of 4,405) of pdARDs coming from Proteobacteria. An additional seven pdARDs were predicted to be harboured by Archaea (Methanobrevibacter and Methanoculleus genera), putatively conferring resistance to macrolides, tetracyclines, aminoglycosides, sulfonamides and glycopeptides (Supplementary Table 1). We also predicted ARDs in genera of medical interest where no ARDs had been identified such as Akkermansia23 (ten pdARDs) and Faecalibacterium24 (44 pdARDs). Only 23 out of 6,095 (0.4%) had been previously identified in families and genera that include human pathogens (Enterobacteriaceae, Campylobacter, Enterococcus, Streptococcus and Acinetobacter). The distribution of the families of pdARDs differed according to the phyla (Supplementary Fig. 9): Firmicutes and Proteobacteria were enriched with aminoglycosides-modifying enzymes (spanning APH, ANT and AACs) whereas Bacteroidetes were enriched in Sul and class A β-lactamases. Interestingly, the tigecycline-degrading monooxygenase Tet(X) was frequently found in Bacteroidetes and Proteobacteria, the two phyla between which transfer of the tet(X) gene has been reported14,25. To support these assignments, we sequenced the metagenome of four human faecal samples before and after an overnight culturing using conditions that favoured the growth of oxygen-tolerant bacteria such as Enterobacteriaceae and enterococci (see Methods). The results showed an enrichment of Proteobacteria (over Firmicutes and Bacteroidetes), and a commensurate increase of class C β-lactamases, Fos and Tet(X), along with Van ligases (Supplementary Fig. 10).
Location of the pdARDs and association with MGEs
We investigated the potential for mobility of the pdARDs at different levels. First, we took advantage of the identification of gene clusters based on co-abundance and co-occurrences of genes among the 396 faecal metagenomes used to build the 3.9 million MetaHIT gene catalogue19. A total of 7,381 gene clusters referred to as metagenomic units (MGUs) were identified. Among MGUs, metagenomic species (MGS) are defined as MGUs with ≥700 genes, which are considered to be representative of partial or complete bacterial genomes19. MGUs of <700 genes include MGEs such as plasmids, phages, transposable elements and incomplete chromosomal sequences. The 7,381 MGUs from the 3.9 million gene catalogue of intestinal microbiota gene were queried with the pdARDs. A total of 3,651 (59.9%) pdARDs could be mapped onto an MGU. The distribution of pdARDs as a function of MGU size is shown in Fig. 2a. Most (95.6%, 3,489 out of 3,651) pdARDs mapped onto MGS and the relative abundance of pdARDs correlated strongly with the abundance of their respective MGS (Supplementary Information), supporting their location on the same bacterial host across the 396 individuals. We also searched for pdARDs in MGS pangenomes (MSPs)26 obtained from the 9.9 million intestinal gene catalogue27. Similar to MGS, MSPs are clusters of genes that are co-abundant in a set of sample. In MSPs, genes that are constantly found are referred to as ‘core’, while inconsistently found genes are referred to as ‘accessory’. Besides, ‘shared core’ genes are assumed to be conserved genes shared between phylogroups26. We found 4,912 pdARDs located on MSPs, with the majority being assigned to the core pangenome (83.4%, 4,099 out of 4,912) or shared between core-pangenomes (7.9%, 389 out of 4,912). This was different with MGE-associated genes27 with most not being found in MSPs (Fig. 2b).
a, Distribution of the sizes of the MGU where an ARD was predicted with respect to the co-location of MGE-associated genes. The vertical line depicts the assumed gene size threshold above which MGUs are considered as partial chromosomes referred to as MGS19. b, Bar plot of the categories of MSPs26 assigned to MGE-associated genes27 and pdARDs. c, Proportion of pdARDs co-locating with MGE-associated genes with respect to their phylum. d, Proportion of pdARDs co-locating with MGE-associated genes according to the pdARD family. Of note, the AAC(2’) and 16S rRNA methylases only included three and two pdARDs, respectively, and were accordingly not depicted in this panel.
Then, we investigated whether genes associated with gene mobility (transposases, conjugative elements and integrons) were present on the same contig than the pdARDs. We found that 7.9% (484 out of 6,095) of pdARDs were co-located with homologues of MGE-associated genes. For pdARDs not found in MGS or in MSPs (n = 974), 876 (89.9%) had no detectable MGE-associated genes in their vicinity.
Finally, we searched for pdARDs homologues (BLASTN >97% identity over >90% of the query length) in the GenBank database (2018 July 11). Only 538 pdARDs homologues were identified, with 49 being located on a plasmid and/or a phage (Supplementary Table 3). Among the 489 remaining pdARDs, 82 (16.8%) were found in multiple species, mainly (73.2%, 60 out of 82) from the same genus (Supplementary Table 4).
The phyla Bacteroidetes, Firmicutes and Tenericutes had the higher proportions of ARDs co-locating with MGEs (Fig. 2c). No ARD family was found to be enriched in MGE, with the exception of the Tet(X) family in which three out of nine (33.3%) predictions (two from Bacteroides fragilis and one from E. coli) were associated with transposases (Fig. 2d).
Distribution of pdARDs in human hosts’ microbiota
In the MetaHIT cohort (663 subjects), we found that subjects carried pdARDs with a median relative abundance of 0.22% (range 0.14–0.38%), with pdARDs from the Tet(M) family being the most abundant (0.07%) and those from class B3 β-lactamases the least (median 0.004%). The average number of unique pdARDs genes detected per metagenome was 1,377 (range 258–2,367). Most pdARDs were shared across multiple subjects, 987 out of 6,095 (16.2%) were found in at least 50% of individuals and only 106 out of 6,095 (1.7%) occurred uniquely in a single individual. All ARD families, with the exception of 16S rRNA methylases and AAC(2’) families, were found in more than 80% of individuals.
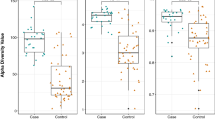
Then, we assessed whether subjects with no recent exposure to antibiotics could cluster according to their intestinal resistome. Based on the pdARDs family patterns, six clusters (that we named ‘resistotypes’ by analogy with the enterotypes28) were detected using Dirichlet multinomial mixture models (Supplementary Fig. 11). The four most frequent resistotypes each represented around 20% of the cohort (the fifth and the sixth representing 8.7 and 7.5%, respectively). The three first resistotypes were characterized by a high abundance of Van ligases (Supplementary Fig. 12). Resistotype 1 was enriched in ANT, while resistotype 3 was driven by Tet(M) and class C β-lactamases. Resistotype 4 was enriched with Tet(X) and class A β-lactamases and resistotype 6 in class B1 β-lactamases and Sul. We observed that resistotypes, as determined by PCM, were highly connected to the composition of the microbiota, and that this effect was more pronounced than resistotypes determined from the results of BLASTP and Resfams (Fig. 3a). The resistotypes of the MetaHIT cohort were found to be associated with enterotypes (chi-square test, P = 5 × 10–4, see Fig. 3b–d and Supplementary Fig. 13). Resistotypes 1 and 3 had higher gene richness and were associated with the Clostridiales-driven enterotype. Resistotype 4 was more prevalent in enterotypes driven by Bacteroides (known to harbour Tet(X) and class A β-lactamases) while resistotype 6 was very specific to the Prevotella enterotype (Fig. 3c,d). The relative abundance of pdARDs was observed to be positively correlated to the gene richness (Fig. 4a, Spearman’s rank correlation test ⍴ = 0.31, P = 5 × 10–16). Conversely, we did not find any link between resistotypes and body mass index, age or gender.
a, Inertia shared between pdARDs profiles and microbiota composition as function of bioinformatics methods. We assessed how gut microbiota β-diversity inertia was connected to the abundance of pdARDs. Co-inertia using the RV coefficient was analysed to detect significant co-structure between datasets69, meaning that different sets of variables (for example, microbial genera abundance and ARDs profiles) were not independent and shared a fraction of inertia. Monte Carlo tests were used to confirm observed relations between different datasets, assuming a P < 0.05. b, Samples proportions for each resistotype depicted as function of enterotypes using the PCM method. c,d, Association between pdARDs gene profile and gut microbiota composition using co-inertia analysis with respect to their enterotypes and pdARDs families (c), and to their resistotypes and MGS relative abundance (d). A taxonomical correspondence for each MGS number can be found in the original paper19. Briefly, all MGS were Firmicutes with the exception of MGS:164 and MGS:445 (both Bacteroidetes).
a, Gene richness and relative abundance of predicted pdARDs in the MetaHIT cohort (n = 663). b, Gene richness and relative abundance of pdARDs in our cohort of subjects with no recent antibiotic exposure (n = 44). c, Gene richness and relative abundance of pdARDs in our cohort of subjects with regards to their antibiotic exposure (n = 102 samples). d,e, Boxplots superimposed by dot plots of the comparisons of the relative abundance of all pdARDs and gene richness, respectively, between the various groups differing by their exposure to antibiotics. Hospitalization: n = 15, Wilcoxon paired test. Short high-dose exposure: n = 10, Wilcoxon paired test. Chronic exposure: n = 31 for patients not exposed to antibiotics, n = 30 for patients chronically exposed to antibiotics, Wilcoxon unpaired test. ATB, antibiotics. The shaded grey area depicts the 95% confidence interval around the blue, linear regression line. For boxplots, the lower, central and upper hinges correspond to the first, second (median) and third quartiles. The upper and lower whiskers, respectively, correspond to the higher and lower values at the 1.5 interquartile range from the hinge (where the interquartile range is the distance between the first and third quartiles).
Dynamics of the pdARDs under various exposures to antibiotics
We investigated the abundances of pdARDs in subjects under various exposures to antibiotics and healthcare environments. Three types of exposure were considered (see Methods for details): hospitalization in a French hospital without receiving antibiotics, n = 15, chronic exposure (Spanish cystic fibrosis patients frequently exposed to antibiotics, n = 30) and short high-dose exposure through selective digestive decontamination (SDD; oral colistin, tobramycin, antifungal amphotericin and parenteral cefotaxime29) at admission in intensive care units in the Netherlands, n = 10). We again confirmed a positive correlation between relative abundance of pdARDs and gene richness among patients unexposed to antibiotics (Fig. 4b, Spearman’s rank correlation test ⍴ = 0.37, P = 0.01, see Methods). However, when all the samples were considered, including those with antibiotic exposure, this relationship was no longer present (Fig. 4c). Instead, the relative abundance of pdARDs was found to be higher in subjects with a chronic exposure than in subjects with no recent exposure (Fig. 4d, Wilcoxon unpaired test P = 1 × 10–10), and gene richness was lower (Fig. 4e, Wilcoxon unpaired test P = 0.006) In particular, subjects with chronic exposure carried more class B1-B2 β-lactamases, AAC(6’), ANT, APH, Erm and DfrA with lower abundance of Sul (Supplementary Fig. 14). At the phylum level, we observed a decrease of Bacteroidetes and Verrucomicrobia and an increase of Firmicutes and Actinobacteria in patients chronically exposed to antibiotics (Supplementary Fig. 15). A total of 74 MGS were found to be differentially abundant among subjects with or without chronic exposure to antibiotics (Supplementary Table 5).
This was different with subjects before and after SDD. A drastic loss of gene richness was measured for this group (Fig. 4e): from a mean of 295,919 genes to 95,286 (67.8% reduction, Wilcoxon paired test P = 0.006). Meanwhile, the relative abundance of pdARDs did not change significantly (Fig. 4d, P = 0.4). At the ARD family level, we observed that some families decreased significantly: class C β-lactamases (commonly found in Enterobacteriaceae and Pseudomonadaceae that are specifically targeted by SDD), Fos, Tet(X), APH and ANT (Supplementary Fig. 16). We then analysed the MGS at the phylum level and found that Proteobacteria, Actinobacteria, Firmicutes and Fusobacteria decreased significantly after SDD (Supplementary Fig. 17). A total of 358 MGS were found in this cohort and, despite the small number of subjects (n = 10), we found 133 MGS for which a significant variation was observed (Supplementary Table 6). We tested whether a high abundance of pdARDs could be protective against the antibiotics used in SDD, but found no association: the relative abundance of pdARDs before SDD was not linked to the gene richness after SDD. Hospitalization without antibiotic therapy, that is, potential exposure to antibiotic-resistant nosocomial pathogens without selective pressure, did not affect the gene richness nor the relative abundance of pdARDs (Fig. 4d,e).
Discussion
The results of this study support the concept that the majority of ARDs from the intestinal microbiota is hosted by commensal bacteria, and that their transfer between species (including to opportunistic pathogen) is rare30. We provide several findings to support this assumption: (1) we used a three-dimensional structure-based method to assess the diversity of ARDs in the intestinal microbiota and confirmed that ARDs predicted by PCM in the intestinal microbiota were distantly related to known ARDs; (2) the sensitivity and the specificity of the method was validated by gene synthesis of a subset of predictions and by benchmarking against various datasets (functional metagenomic of the soil microbiota, genomes and random protein catalogue); (3) the majority of pdARDs could be found in clusters of co-abundant genes (MGS and MSPs) in large cohorts of samples, while only a minority was found on plasmids, phages or in the vicinity of MGE-associated genes; (4) we could stratify subjects into resistotypes that were connected to enterotypes; and (5) gene richness, otherwise associated with a healthy status31, was positively correlated to the abundance of ARDs in subjects not exposed to antibiotics.
Our results challenge the paradigm that ARDs of the intestinal microbiota are a threat to public health. As was previously demonstrated for environmental samples18,32, ARDs tend to cluster according to the underlying microbial ecology of the ecosystem, suggesting that the vast majority of ARDs are fixed in their microbial hosts and are not, or are very rarely, transferred. Our results show that the dominant intestinal microbiota is not a major conduit through which opportunistic pathogens can acquire ARDs. Nevertheless, we acknowledge that such transfer events have been reported14,15 and that consequences for public health can be important, as in the case of the vanB vancomycin resistance operon that is shared by Clostridium spp. and enterococci15. Understanding the mechanisms that can lead to the mobilization of ARDs in the intestinal microbiota, as well as a broader census of environmental reservoirs of ARDs (for example, sewage, livestock, the subdominant human intestinal microbiota) will continue to be an important area for future research.
We found that subjects cluster according to the composition of their resistome into six groups named resistotypes (as a reference to the previously described enterotypes28). These resistotypes were indeed connected to the enterotypes. Description of this underlying structure is interesting as one might hypothesize that a particular resistotype, or microbiota enriched with ARDs, might be affected to different degrees by antibiotic therapy. This has previously been observed for β-lactamase-producing Bacteroides that can protect the microbiome against exposure to β-lactams33. In patients undergoing faecal microbiota transplantation, follow-up antibiotic therapy may be adjusted to favour engraftment of the donor microbiota34. Identifying donors with a resilient microbiota, due to a protective resistotype, could open perspectives for the optimization of the clinical implementation of faecal microbiota transplants.
Contrary to initial expectations, some pdARD families decreased in their abundance under antibiotic exposure, especially when patients were exposed to a combination of antibiotics (such as SDD). To resist to a combination of antibiotics, bacteria would need to be intrinsically resistant or to acquire an adequate combination of ARDs. The dynamics of ARDs under antibiotic exposure depend on various parameters: spectrum of the ARD (the level of resistance towards the antibiotic provided by the ARD), the expression level of the ARD and the presence of other resistance mechanisms (intrinsic or acquired). The large number of possible combinations of these factors can explain that, in some situations, a bacterium can be inhibited by antibiotics despite the presence of a putatively compatible ARD. Alternatively, we cannot exclude that changes in pdARDs families could also be explained by simple taxonomic shifts that are not connected to the antibiotics studied.
The limitations of current techniques and of this study leave a number of important questions unresolved. As mentioned earlier, metagenomic sequencing provides information for the dominant fraction of intestinal bacteria, and so ARDs present in subdominant bacteria remain unobserved. Indeed, several ARDs found in opportunistic pathogens among the Enterobacteriaceae (for example, E. coli and Klebsiella pneumoniae) originate from other species in the same Proteobacteria phylum35. A recent study indeed cultured many Proteobacteria species that were not detected in metagenomic sequencing36. We cannot rule out that the subdominant bacteria, which were not probed by metagenomic sequencing, could be an additional reservoir of ARDs. In terms of the clinical samples analysed, we cannot exclude that the differences between patients and controls may be resulting from confounding factors other than the antibiotic exposure.
The method we used to identify distantly related proteins is based on homology modelling and takes advantage of the observation that proteins sharing the same function have more similar structures than amino acid sequences37. Indeed, PCM could identify functional ARDs with amino acid identity below 20% to known ARDs. Notably, PCM can only be used to predict the function of genes that are homologous to known ARDs, and therefore the identification of different classes of ARDs with no homology to known ARDs will still require functional screening. Besides, while PCM was validated in this study, it remains a prediction tool. While similar structures are usually indicative of similar function, this is not always the case and PCM can yield false positive results (as observed in the functional validation of synthesized pdARDs). Due to the scope of our study, gene synthesis validation was not performed for all ARD families, leaving open the possibility that not all pdARDs identified here truly have a role in antibiotic resistance.
In summary, we developed a method, PCM, which could unveil the diversity of ARDs in the intestinal microbiota. Employing this tool, we gathered evidence that the vast majority of the ARDs we predicted showed no sign of mobility and that their abundance was correlated to gene richness. Together with the protective trait of some intestinal bacteria against antibiotics33, our results suggest that the ARDs from the intestinal microbiota might be considered as our ‘resilience allies’38 assuring the preservation of the healthy commensal microbiota under antibiotic exposure.
Methods
Constitution of the databases of ARDs
We define as an ARD as in Martínez et al.39: a protein encoded by a gene that confers resistance to antibiotics when it is present or increases susceptibility to antibiotics when it is absent. This definition excluded housekeeping genes in which mutations can confer resistance to some antibiotics (such as topoisomerases in which mutations can lead to fluoroquinolone resistance) and genes involved in the regulation of antibiotic resistance genes. Also, we excluded efflux pumps such as Tet(A) or QepA as very few or no PDBs are available, presumably due to the difficulty to crystallize transmembrane proteins. Amino acid sequences of functionally characterized ARDs from the major antibiotic families used in human medicine (β-lactams, aminoglycosides, tetracyclines, trimethoprim, sulfonamides, macrolides-lincosamides-synergistines, fluoroquinolones, fosfomycin and glycopeptides)20,40 were obtained from the following antibiotic resistance databases: Resfinder9, ARG-ANNOT7, the Lahey Clinic (http://www.lahey.org/studies/), RED-DB (http://www.fibim.unisi.it/REDDB/), Marilyn Roberts’s website for macrolides and tetracycline resistance genes (http://faculty.washington.edu/marilynr/) and from functional metagenomics studies5,6,41. When ARDs were provided as nucleic acids sequences, they were translated into proteins with Prodigal42. Non-redundancy of the reference ARDs was assessed with CD-HIT v4.5.743 (100% identity). The final database was manually curated to remove incomplete sequences and ARDs from families not considered in this work. The cluster of orthologous genes (COG) of each member of the reference dataset was assigned from the v3 eggNOG database44. In total, we collected 1,651 non-redundant amino acid sequences spanning 20 ARD families: class A β-lactamases (Blaa), class B1-B2 β-lactamases (Blab1), class B3 β-lactamases (Blab3), class C β-lactamases (Blac), class D β-lactamases (Blad), aminoglycoside acetyltransferases (AAC) AAC(2’), AAC(3)-I, AAC(3)-II and AAC(6’), aminoglycoside nucleotidyltransferases (ANT), aminoglycoside phosphotransferases (APH), 16S ribosomal RNA (rRNA) methylases, Tet(M), Tet(X), type A dihydrofolate reductases (DfrA), dihydropteroate synthases (Sul), erythromycin ribosome methylases (Erm), quinolone resistance proteins (Qnr), fosfomycin resistance proteins (Fos) and D-Ala–D-Lac/Ser ligases (Van) (Table 1). The recently described plasmid-mediated colistin resistance mcr-1 gene45 could not be included because of the lack of a reliable PDB template obtained by X-ray diffraction at the time of the study.
Interrogation of the catalogue for ARDs
We used a 3,871,657 million proteins catalogue previously published19. This catalogue was built from the metagenomic sequencing of the faeces of 396 subjects from Denmark and Spain. In brief, the 3.9 million gene catalogue results from a non-redundancy filtering at 95% nucleic acid identity and 90% coverage: predicted genes from all samples (45.4 million in total) were clustered using BLAT by single linkage. Any two genes with greater than 95% identity and covering more than 90% of the shorter gene were clustered together. The contigs were originally built using SOAPdenovo (from the MOCAT pipeline46). We selected this catalogue over the more recent 9.9 million gene catalogue that was published during the course of this study27 because MGUs (including the MGS) had been determined only for the 3.9 million gene catalogue. The genes of the catalogue were translated into proteins using Prodigal42 using the –p meta option. For each ARD family, we searched for ARDs using the following three methods: (1) we built a hidden Markov model file for each ARD family and searched the catalogue with Hmmsearch (v3.1)47; (2) we performed a Smith–Waterman alignment with a heuristic seed detection (BLASTP v.2.2.28+)21; and (3) a rigorous Smith–Waterman search (SSearch v.36.3.6)48 with an E-value threshold of 1 × 10–5. Only the hits with a size ranging from 75 to 125% of the mean amino acid size of the ARD family were further considered. All candidates were assigned a COG/NOG from eggNOG v344. When candidates were found in different ARD families (for example, a candidate could be a hit in class B1-B2 and class B3 β-lactamases), the candidate was assigned to the family for which it had the highest amino acid identity with the reference.
Negative references
For each ARD family, COGs/NOGs were attributed to reference ARDs. In parallel, the COGs/NOGs were attributed to the hits obtained during the initial steps of PCM (that is, the hits obtained by the BLASTP/SSearch and Hmmer search). In the list of candidates from a given ARD family, the COGs/NOGs that were not found in the COGs/NOGs attributed to reference ARDs were assumed to be potential COGs/NOGs from false positive hits (Supplementary Fig. 2) as it reproduced the errors of functional assignment that were likely to be generated in sequence-only annotations. The amino acid sequences of the representative proteins from those COGs/NOG groups were obtained from the eggNOG v3 database and were added to the negative reference dataset. A manual curation step was performed to ensure that no references were included in the negative references.
Selection of structural templates
The list of protein structures that could be used as structural templates was downloaded (June 2014 and November 2014) from the PDB library (ref. 49, http://www.rcsb.org/). Using the reference dataset and the negative references described above, Hmmer47, BLASTP21 and SSearch48 were performed on the PDB database with default settings and E values of 1 × 10–5. Results were merged into a non-redundant PDB list. Both lists (references and negative templates) were manually curated to ensure that no references were represented in the negative templates dataset, and vice versa. If more than one PDB shared the same UniProt number (that is, if the structure of a protein has been determined on multiple occasions), we filtered the PDB files to include a unique structure per UniProt number using the following positive criteria: absence of ligand, completeness of the protein and high resolution.
PCM
The concept of PCM is shown in Supplementary Figs. 1–3 and the framework is available at https://github.com/aghozlane/pcm. The concept of leveraging the protein structure in complement to its amino acid sequence was motivated by the fact that proteins sharing common functions would be more conserved in the active site that cannot be observed by the analysis of protein sequence alignments37. Each candidate was subjected to homology modelling with reference templates and negative templates, generating two three-dimensional structures for each candidate (Fig. 1a). The main idea is that if a sequence is truly functionally related to the reference fold, its model must be significantly different from the ones obtained with the negative structural template. Homology modelling was performed by PCM in six main steps (example in Supplementary Fig. 3):
-
(1)
Three structural templates were identified by BLASTP (among the lists produced as described above) that shared the highest amino acid identity with the candidate protein.
-
(2)
A multiple sequence alignment was performed between the candidate and the three templates sequences using Clustalo50.
-
(3)
A prediction of the secondary structure was performed using psipred (v3.5)51. The residues predicted to fold in helix or in beta-sheet conformation with a level of confidence higher or equal to seven were considered to constrain the model.
-
(4)
A comparative modelling was performed with the MODELLER programming interface52. MODELLER automatically calculates a model by satisfaction of spatial restraints such as atomic distance and dihedral angles in the target sequence, extracted from its alignment with the template structures. Stereo-chemical restraints for residues are obtained from the CHARMM-22 molecular force field and statistical preferences obtained from a representative set of known protein structures.
-
(5)
The best model out of a hundred produced by MODELLER (based on the Dope score) was considered for structure assessment analysis using ProQ53 and Prosa-web54. The Dope score (Modeller), z-score (Prosa), MaxSub and Levitt-Gerstein (LG) score (ProQ) are statistical potential variables used to predict the model quality. Both ProQ and Prosa-web are trained on the PDB to determine real protein configuration and they estimate the energetic farvourability of the conformation of each residue in the model.
-
(6)
The best model was aligned with the reference set of structures using TM-Align17 and MAMMOTH55. The r.m.s.d (TM-Align), z-score (MAMMOTH), TM score (MAMMOTH, TM-Align) estimates the degree of superposition of the residue between two structures.
The differences (delta) between the scores determined from each modelling path (with the reference set or the negative set) were calculated and used for the PCM machine-learning program (see below).
For one given candidate, the PCM whole process took an average of 8 CPU-hours (30 min on 16 CPUs).
Taxonomic assignation
The pdARDs were taxonomically assigned by combining the results obtained from BLASTN against the National Center for Biotechnology Information (NCBI) Genomes database (minimal 70% identity and 80% coverage), a BLASTN against the IMOMI in-house database (minimal 85% identity and 90% coverage) and the taxonomy of the metagenomic unit whenever applicable. The lowest taxonomic rank from the results of the three methods was assigned to the pdARD.
Statistical analysis
To discriminate reference proteins from negative references, we used model quality predictors and alignment scores (inferred from the semi-automatic pipeline described above) and developed a custom pipeline in R (R Core Team, 2013, http://www.R-project.org) to perform the classification. The LASSO penalized logistic regression56 implemented in LIBLINEAR57 was used to compute the classifier. Ten-fold stratified cross validation (re-sampled 100 times to obtain more stable accuracy estimates) was used to partition the data into a training and test sets. The LASSO hyper-parameter was optimized for each model in a nested five-fold cross validation on the training dataset using the area under curve as the model selection criterion. From the 100 times re-sampled ten-fold cross validation, receiver operating characteristic analysis was used to evaluate model performance using the median area under curve. Coefficients extracted for each modelling or alignment score were also evaluated for their stability throughout the computed models. The PCM score was the ratio (expressed as a percentage) between the numbers of time a candidate was classified as a reference and the number of bootstraps. Predicted ARDs were candidates with a PCM score ≥50% and a TM score given by TM-Align ≥0.517. To control how structural modelling brought additional information compared to amino acid sequence alignment only, we built a logistic regression model based on T-coffee alignment score (R glm, ten-fold stratification, re-sampled 100 times). We then compared the two classifier models used for PCM and for T-coffee alignment based on the reference set (see Supplementary Information).
Validation of the method with a functional metagenomic dataset
The performance of PCM was assessed by analysing the data in Forsberg et al., where the ARD content of different North American soils was analysed using functional metagenomics18. The screening of the clones was performed on aztreonam, chloramphenicol, ciprofloxacin, colistin, cefepime, cefotaxime, cefoxitin, D-cycloserine, ceftazidime, gentamicin, meropenem, penicillin, piperacillin, piperacillin-tazobactam, tetracycline, tigecycline, trimethoprim and trimethoprim-sulfamethoxazole (cotrimoxazole). Here, we collected the nucleotide sequences of the inserts deposited on GenBank (KJ691878–KJ696532). The sequence translation of the open reading frames was performed by Prodigal (using default parameters)42. A total of 4,654 insert sequences were collected, in which 12,904 amino acid sequences were predicted. We then searched for ARDs belonging to the relevant ARD families according to the antibiotics used for the screening of the clones: β-lactamases (all classes), APH, ANT, AAC(2’), AAC(3)-I, AAC(3)-II, AAC(6’), 16S rRNA methylases, Tet(M), Tet(X), Qnr, Sul and DfrA, using the Supplementary Table 2 of the Forsberg et al. paper. Inserts with no putative ARDs (according to the annotation of the gene) were removed (n = 269). Inserts selected on cycloserine (n = 868) and chloramphenicol (n = 129) were not considered here because they were not included in the 20 ARD families in this work. Fourteen inserts that contained more than one putative ARD that could be identified to confer resistance to the antibiotic used for the screening (for example, two β-lactamases) were not considered in this analysis. An additional 1,658 inserts containing no putative ARDs or a putative ARDs that did not confer resistance to the antibiotic used for selection were discarded and so were 294 inserts containing efflux pumps, as these were not considered in this study. The resulting validation set contained 1,423 inserts (with resistance genes) for a total of 3,778 genes. To compare the outcome of PCM with other tools, the results for class B1-B2 and B3 β-lactamases generated by PCM were merged into one class B β-lactamases group as other tools do not separately consider the different class B β-lactamases.
In total, 1,390 unique hits were found during the initial screen of PCM, of which 1,374 were predicted as ARDs (Supplementary Table 7). Among the 33 ARDs not included for PCM, 12 were not considered because they were undersized and ten because they were oversized. No hits for AAC(2’), ANT, Qnr or Sul were found. The mean identity shared with reference ARDs was 37.6% (range 18.8–94.5). Overall, the sensitivity was 96.6%, with no false negative. In comparison, only eight ARDs would have been identified by a conventional method (combination of Hmmsearch, BLASTP and SSearch with both a minimal identity with a reference ARD and coverage over or equal to 80%). Conversely, Resfams11 that was specifically designed to identify ARDs from functional metagenomic datasets showed a similar sensitivity to PCM with the identification of 1,346 ARDs out of 1,423 (94.6% sensitivity).
Validation of the method for incomplete genes
The 3.9 million gene catalogue harbours 41.4% of genes that are predicted to be incomplete either on the 5’, the 3’ or both extremities19. As the size parameter is crucial for homology modelling, we tested to what extent the prediction of incomplete ARDs by PCM could remain valid. We selected 12 reference class A β-lactamases (BlaZ, CblA-1, CepA-29, CfxA2, CfxA6, CTX-M-8, KPC-10, OXY-1, PER-1, SHV-100, TEM-101 and VEB-1) and we then iteratively removed 5% of the amino acid sequence at both edges to obtain 16 bi-directionally trimmed candidates (from 100 to 25%) per reference ARD. Candidate genes were chosen to span the diversity of known β-lactamases, but the main representative β-lactamase of the subfamily (for example, TEM-1 for TEM β-lactamase) was not necessarily chosen. Note that SHV-100 has a slightly longer sequence (13 amino acid duplication) than other SHV. A total of 192 PCM experiments were performed: we observed that the 12 references were correctly predicted as ARDs when at least 40% of the protein remained (that is, 30% trim from each extremity, Supplementary Fig. 4). Thus, we are confident that with the 75% size threshold used in this study (a maximum of 25% removed from one edge), no misclassification due to an incomplete gene would be expected.
Gene synthesis
We selected 71 pdARDs from 12 ARD families: 14 from class A β-lactamases, eight from class B1-B2 β-lactamases, seven from class B3 β-lactamases, four from class C β-lactamases, two from class D β-lactamases, 2 AAC(3)-I, 5 AAC(3)-II, 8 AAC(6’), 3 ANT, 4 APH, 13 Tet(M) and 1 Tet(X)) for gene synthesis and sub-cloning into E. coli to test the decrease of susceptibility to antibiotics. For β-lactamases, a chromogenic test (nitrocefin) was used to detect function. Minimal inhibitory concentrations (MIC) were determined by E-Test strips (bioMérieux, Marcy-l’Etoile, France) in duplicate. A pdARD was considered to have an activity against an antibiotic (tobramycin for AAC(3)-I, AAC(3)-II, AAC(6’) and ANT; kanamycin for APH and tetracycline for Tet(M)) when the MIC of the clone was above the MIC of a clone harbouring the plasmid without a synthesized gene or when the colour of the broth containing nitrocefin turned red, in the case of β-lactamases. We used the plasmid vector pET-22b+ (embedding a β-lactamase-encoding gene) for pdARDs hypothesized to confer resistance to aminoglycosides and the pET-26b (embedding a gene conferring resistance to kanamycin) for the other pdARDs. The selection of the pdARDs for synthesis was performed as follows: references (n = 12), pdARDs that shared a high identity with known ARDs (≥95% amino acid identity and ≥80% coverage with a reference ARD); good predictions (n = 41), pdARDs with the highest degree of confidence for the prediction (PCM score >99%, TM score TM-Align >0.9 and <70% amino acid identity with a reference ARD); fair predictions (n = 18): pdARDs with the lowest degree of confidence for the prediction (PCM score <80%, TM score TM-Align <0.8 and <70% amino acid identity with a reference ARD).
Signatures of MGEs nearby the predictions of ARDs
We searched for MGE-associated proteins encoded by genes located in the same contigs as pdARDs. The 3.9 million gene catalogue results from a non-redundancy filtering at 95% for the genes19, but to identify the contigs on which pdARDs were identified, we needed to return to the redundant catalogue (that is, the non-dereplicated catalogue of genes) and identified homologues sharing 95% nucleic acid identity with the pdARDs. By doing so, we could identify contigs (n = 16,955) carrying at least one pdARD. The mean size of the contigs was 19,711 base pairs (minimum 500, maximum 461,981, median 8,513). In total, the 16,955 contigs contained a total of 908,888 genes after the subtraction of pdARDs. The 908,888 genes were then translated into proteins with Prodigal42 and queried for IS elements using BLASTP (query size threshold, 150 amino acids; E value, 1 × 10–30; identity threshold, 40%) against the ISfinder database58. Conjugative elements were queried among the same gene set (n = 908,888) with Conjscan59, using the default parameters and the filters recommended by the authors (best E < 0.001 and sequence coverage of at least 50%). Most proteins belonging to the type IV secretion systems (T4SS), which are involved in conjugation, are ubiquitous in that they have numerous homologues. Hence, when searching for conjugation proteins in a 3.9 million protein catalogue, there would be a high risk of false positives. Accordingly, the co-location of hits was deemed crucial. A conjugative T4SS is made from:
-
a protease (VirB4)
-
a second coupling protein protease (t4cp)
-
a relaxase (MOB)
-
a proteic complex (MPF) composed of at least ten proteins
To identify a T4SS on a contig, we required presence of at least one virB4 hit, a t4cp1 or t4cp2 hit, a MOB hit and a certain number of MPF hits. All hits must co-localize. A MOB element alone can mobilize a neighbouring gene (such as an ARD-encoding gene) via other T4SSs. However, in our dataset the short length of contigs led us to adapt those parameters (following the recommendations of the developers of the Conjscan software). Besides the MOB element, we considered that the presence of two hits from the same family (for example, T_virB6 and T_virB8, or B_traF and B_traH) or virB4+ any hit from another family on the same contig as a pdARD was a strong indication of the presence of mobility associated elements. Integrons were identified using IntegronFinder60 on the 16,955 contigs using default parameters.
We also searched for pdARDs in MSPs26 obtained from the 9.9 million intestinal gene catalogue27 using BLASTN with a 95% identity threshold over 90% of the query.
Finally, we searched for homologues of pdARDs in GenBank with 97% identity threshold over 90% of the query. We found 820 out of 6095 pdARDs (13.5%) that aligned against 139,413 GenBank entries. We filtered hits corresponding to a virus, a plasmid or a vague taxonomic affiliation by considering the following terms: uncultured bacterium, artificial, unidentified, uncultured organism, environmental samples and metagenome.
Distribution of the pdARDs in the MetaHIT cohort (n = 663 subjects)
pdARDs profiles were obtained from the abundance matrix of the 3.9 million genes as described in Nielsen et al19. The ‘reads per kilobase per million mapped read’ method was used to normalize the mapping counts. After summing the relative abundances of pdARDs genes belonging to the same family, Dirichlet multinomial mixture models were used to find ARD clusters (that is, resistotypes) using the Dirichlet multinomial R package. The same method was applied to detect gut microbiota clusters (that is, enterotypes)61. The Laplace criterion was used to define optimal number of clusters as described on oral and faecal microbial dataset62. By analogy with the term enterotype, we chose to name a cluster of subjects on the basis of their similarity of their faecal relative abundance of pdARDs a resistotype. The chi-squared test was used to assess the associations between resistotypes and enterotypes. Rarefaction analysis at 1 million reads was done to determine the gene richness per samples. RLQ analysis63 was conducted to assess the associations between the relative abundances of pdARDs, their characteristics (family, size of the cluster of associated genes) and those of subjects (enterotypes, resistotypes, gender, body mass index, age). Of note, we excluded the patients suffering from inflammatory bowel disorders from this analysis. Co-inertia analysis was conducted to assess the associations between microbiota β-diversity and pdARDs profiles. Microbiota composition was assessed using MGS (see below) relative abundance and β-diversity by square root Jensen–Shannon Divergence. A principal coordinate analysis was done on Jensen–Shannon Divergence distance matrix and a principal component analysis was done on ARD profiles. Both analyses were then subjected to co-inertia analysis and Monte Carlo permutation was done to assess to robustness of shared inertia.
Constitution of cohorts of patients with various antibiotic exposures
We included three cohorts of patients with various exposures to antibiotics.
Hospitalization without antibiotics
A total of 31 patients with no exposure to antibiotics or hospitalization during the three preceding months and admitted to the medicine ward of the Beaujon University Teaching Hospital (Clichy, France) were included and provided a faecal sample at admission. Among them, 16 also provided a stool sample at discharge. One patient received antibiotics between admission and discharge and was not further considered for the analysis. In total, 15 patients could provide a stool sample soon after admission (T0) and at discharge (T1). The mean time between T0 and T1 samples was 10.7 days. The mean age of patients was 67.8 years old and the gender ratio (M/F) was 1.3. All patients gave informed consent. This work was approved by the French National Institutional Review Board (IRB 00008522) and registered at clinicaltrials.gov (NCT02031588).
Chronic exposure
Thirty cystic fibrosis patients were enroled at the Cystic Fibrosis Unit of the Ramón y Cajal Hospital in Madrid. One faecal sample was collected at the occasion of a consultation. All subjects for this study were provided a consent form describing the study and providing sufficient information for subjects to make an informed decision about their participation as faecal donors in this study. Cystic fibrosis is a genetic disease that leads to an impairment of the lung function through an uncontrolled production of mucus. The consequence is chronic bacterial colonization, resulting in deleterious reactive fibrosis of the lung. Bacterial load is controlled by chronic exposure to antibiotics (home-therapy, mostly oral and inhaled in our cohort), which has resulted in significant life prolongation, and the near-absence of hospital care. Hence, the cystic fibrosis patients had been exposed to various antibiotics during the five years before the faecal sample was collected:
-
β-lactams (ampicilln, amoxycillin, cloxacillin, piperacillin-tazobactam, cefepime, ceftriaxone, ceftazidime, cefditoren, meropenem): 25 out of 30
-
Macrolides (azithromycin, clarithromcyin): 17 out of 30
-
Colistin: 21 out of 30
-
Fluoroquinolones (ciprofloxacin, levofloxacin, moxifloxacin): 26 out of 30
-
Cotrimoxazole: 14 out of 30
-
Glycopeptides (vancomycin): 1 out of 30
-
Aminoglycosides (amikacin, tobramycin): 12 out of 30
-
Tetracyclines (doxycycline, minocycline): 2 out of 30
-
Linezolid: 3 out of 30
-
Rifampin: 1 out of 30
-
Fosfomycin: 5 out of 30
On average, cystic fibrosis patients had been exposed to 5.9 different antibiotics and had an average of 12.2 antibiotic courses during the five years before the sample was taken. The mean age was 36.3 years old and the gender ratio (M/F) was 1.3. The consent form was obtained before that subject provided any faecal sample for the study and was signed by the subject or legally acceptable surrogate, and the investigator-designated research professional obtaining the consent. According to the National Spanish laws the study did not require the approval of the Ethics Committee. Nonetheless, the Ethics Committee of the Hospital Ramón y Cajal guaranteed that the study was performed done according to the good clinical practices guidelines.
Short high-dose exposure
Short high-dose exposure consists of administering a mixture of topical and parenteral antibiotics and antifungal agents to a patient at admission to eliminate potential bacterial and fungal pathogens. SDD has been shown to significantly reduce mortality in the intensive care unit29 and is now part of standard care for intensive care patients in the Netherlands. To assess the effect of SDD on the intestinal microbiota, we analysed the faecal samples from 13 patients admitted to the intensive care unit of the University Medical Centre of Utrecht (Netherlands). The samples were collected at admission (T0, first sample passed after admission) and after SDD (T1). Among the 13 patients for whom a faecal sample could be obtained at T0, 10 could provide a faecal sample at T1. The mean age was 59.9 years old and the gender ratio (M/F) was 0.5. SDD consisted of 4 days of intravenous cefotaxime and topical application of tobramycin, colistin and amphotericin B. Additionally, a subset of samples (n = 4) from this cohort was cultured in a brain-heart infusion broth overnight in ambient atmosphere at 37 °C. The protocol for the collection of stool samples was reviewed and approved by the institutional review board of the University Medical Centre of Utrecht (Netherlands) under number 10/0225. Informed consent for faecal sampling during hospitalization was waived. Written consent was obtained for the collection of faecal samples after hospitalization.
Metagenomic sequencing and mapping
Total faecal DNA was extracted64,65 and sequenced using SOLiD 5500 wildfire (Life Technologies) resulting in a mean of 68.5 million sequences of 35-base-long single-end reads. High-quality reads were generated with quality score cut-off >20. Reads with a positive match with human, plant, cow or SOLiD adaptor sequences were removed.
Filtered high-quality reads were mapped to the MetaHIT 3.9 million gene catalogue19 using the METEOR software66. The read alignments were performed in colourspace with Bowtie software (version 1.1.0)67. Uniquely mapped reads (reads mapping to a single gene from the catalogue) were attributed to the corresponding genes. Shared reads (mapping different genes of the catalogue) were attributed according to the ratio of their unique mapping counts, as following: as a read can map on different genes of the catalogue, the abundance of a gene G(Ag) depends on the abundance of uniquely mapped reads (Au), that is, reads that map only to the gene G and on the abundance of N shared reads (As) that aligned with M genes in addition to the gene G:
where
For each shared read, the gain of abundance corresponds to a coefficient Co that takes in account the total number of uniquely mapped reads on the M genes:
For instance, if a gene G is mapped by ten reads that only map to it (unique reads), but also with one read that also align on a gene M that was mapped by five unique reads, then:
To decrease technical biases due to different sequencing depth, samples with at least 5 million mapped reads were downsized to 5 million mapped reads (random sampling of 5 million mapped reads without replacement) using R package momr31. The abundance of each gene in a sample was then normalized by dividing the number of reads that mapped to the gene (Ag) by the gene nucleotide length and by the total number of reads from the sample. The resulting set of gene abundances, termed a microbial gene profile, was used to estimate the abundance of MGS19.
Gene richness analysis
Microbial gene richness was calculated by counting the number of genes mapped at least once for a given sample. Gene richness was calculated using R package momr for samples where 5 million or more reads had been mapped to the 3.9 million gene catalogue.
MGS
MGS are co-abundant gene groups with more than 700 genes and can be considered part of complete bacterial species genomes. 741 MGS were delineated from 396 human gut microbiome samples19. In this study, the relative abundance of MGS was determined as the median abundance of 90% of the genes composing each cluster, meaning that the 10% genes with the lowest abundance for each MGS were not considered for the calculation of the abundance of the MGS. Typically, these genes correspond to genes with zero count, to accessory genes (hence their detection is not constant) or to genes that are not detected because of insufficient sequencing depth. The MGS taxonomical annotation was updated by sequence similarity using NCBI BLASTN, when more than 50% of the genes matched the same reference of NCBI database (December 2014 version) at a threshold of 95% of identity and 90% of gene length coverage to get the species annotation19.
Statistical analysis for the distribution of pdARDs and MGS between groups
Statistical analyses for the differential abundances of pdARDs and MGS were performed using the application SHAMAN68(http://shaman.pasteur.fr/). Data are available at (https://github.com/aghozlane/evotar), with the graphical representations using the abundances from the matrix rarefied at 5 million reads. The relationship between richness and the abundance of ARDs was assessed by the Spearman correlation test. The statistical threshold for significance was set at a P value of 0.05.
Code availability
The PCM code can be found at https://github.com/aghozlane/pcm.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The 6,095 pdARDs PDB files, nucleotide and amino acid sequences can be downloaded from http://mgps.eu/Mustard/. The 3.9 million gene catalogue and the MGS database are accessible at https://www.cbs.dtu.dk/projects/CAG/. The reads from the clinical samples generated in this study are available under the accession number PRJEB27799 at the European Nucleotide Archive.
References
United Nations High-Level Meeting on Antimicrobial Resistance (WHO, UN, 2016).
Ghosh, T. S., Gupta, S. S., Nair, G. B. & Mande, S. S. In silico analysis of antibiotic resistance genes in the gut microflora of individuals from diverse geographies and age-groups. PLoS ONE 8, e83823 (2013).
Hu, Y. et al. Metagenome-wide analysis of antibiotic resistance genes in a large cohort of human gut microbiota. Nat. Commun. 4, 2151 (2013).
Forslund, K. et al. Country-specific antibiotic use practices impact the human gut resistome. Genome Res. 23, 1163–1169 (2013).
Sommer, M. O. A., Dantas, G. & Church, G. M. Functional characterization of the antibiotic resistance reservoir in the human microflora. Science 325, 1128–1131 (2009).
Moore, A. M. et al. Pediatric fecal microbiota harbor diverse and novel antibiotic resistance genes. PLoS ONE 8, e78822 (2013).
Gupta, S. K. et al. ARG-ANNOT, a new bioinformatic tool to discover antibiotic resistance genes in bacterial genomes. Antimicrob. Agents Chemother. 58, 212–220 (2014).
McArthur, A. G. et al. The comprehensive antibiotic resistance database. Antimicrob. Agents Chemother. 57, 3348–3357 (2013).
Zankari, E. et al. Identification of acquired antimicrobial resistance genes. J. Antimicrob. Chemother. 67, 2640–2644 (2012).
Arango-Argoty, G. et al. DeepARG: a deep learning approach for predicting antibiotic resistance genes from metagenomic data. Microbiome 6, 23 (2018).
Gibson, M. K., Forsberg, K. J. & Dantas, G. Improved annotation of antibiotic resistance determinants reveals microbial resistomes cluster by ecology. ISME J. 9, 207–216 (2015).
Wright, G. D. The antibiotic resistome: the nexus of chemical and genetic diversity. Nat. Rev. Microbiol. 5, 175–186 (2007).
Salyers, A. A., Gupta, A. & Wang, Y. Human intestinal bacteria as reservoirs for antibiotic resistance genes. Trends Microbiol. 12, 412–416 (2004).
Ghosh, S., Sadowsky, M. J., Roberts, M. C., Gralnick, J. A. & LaPara, T. M. Sphingobacterium sp. strain PM2-P1-29 harbours a functional tet(X) gene encoding for the degradation of tetracycline. J. Appl. Microbiol. 106, 1336–1342 (2009).
Stinear, T. P., Olden, D. C., Johnson, P. D., Davies, J. K. & Grayson, M. L. Enterococcal vanB resistance locus in anaerobic bacteria in human faeces. Lancet 357, 855–856 (2001).
Penders, J., Stobberingh, E. E., Savelkoul, P. H. M. & Wolffs, P. F. G. The human microbiome as a reservoir of antimicrobial resistance. Front. Microbiol. 4, 87 (2013).
Zhang, Y. & Skolnick, J. TM-Align: a protein structure alignment algorithm based on the TM-score. Nucleic Acids Res. 33, 2302–2309 (2005).
Forsberg, K. J. et al. Bacterial phylogeny structures soil resistomes across habitats. Nature 509, 612–616 (2014).
Nielsen, H. B. et al. Identification and assembly of genomes and genetic elements in complex metagenomic samples without using reference genomes. Nat. Biotechnol. 32, 822–828 (2014).
Goossens, H., Ferech, M., Van der Stichele, R. & Elseviers, M. & ESAC Project Group. Outpatient antibiotic use in Europe and association with resistance: a cross-national database study. Lancet 365, 579–587 (2005).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Berglund, F. et al. Identification of 76 novel B1 metallo-β-lactamases through large-scale screening of genomic and metagenomic data. Microbiome 5, 134 (2017).
Everard, A. et al. Cross-talk between Akkermansia muciniphila and intestinal epithelium controls diet-induced obesity. Proc. Natl Acad. Sci. USA 110, 9066–9071 (2013).
Sokol, H. et al. Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of Crohn disease patients. Proc. Natl Acad. Sci. USA 105, 16731–16736 (2008).
Leski, T. A. et al. Multidrug-resistant tet(X)-containing hospital isolates in Sierra Leone. Int. J. Antimicrob. Agents 42, 83–86 (2013).
Plaza Oñate, F. et al. MSPminer: abundance-based reconstitution of microbial pan-genomes from shotgun metagenomic data. Bioinformatics https://doi.org/10.1093/bioinformatics/bty830 (2018).
Li, J. et al. An integrated catalog of reference genes in the human gut microbiome. Nat. Biotechnol. 32, 834–841 (2014).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473, 174–180 (2011).
de Smet, A. M. G. A. et al. Decontamination of the digestive tract and oropharynx in ICU patients. N. Engl. J. Med. 360, 20–31 (2009).
van Schaik, W. The human gut resistome. Philos. Trans. R. Soc. Lond. B Biol. Sci. 370, (2015).
Le Chatelier, E. et al. Richness of human gut microbiome correlates with metabolic markers. Nature 500, 541–546 (2013).
Pehrsson, E. C. et al. Interconnected microbiomes and resistomes in low-income human habitats. Nature 533, 212–216 (2016).
Léonard, F., Andremont, A., Leclerq, B., Labia, R. & Tancrède, C. Use of β- lactamase-producing anaerobes to prevent ceftriaxone from degrading intestinal resistance to colonization. J. Infect. Dis. 160, 274–280 (1989).
Bilinski, J. et al. Fecal microbiota transplantation in patients with blood disorders inhibits gut colonization with antibiotic-resistant bacteria: results of a prospective, single-center study. Clin. Infect. Dis. 65, 364–370 (2017).
Lupo, A., Coyne, S. & Berendonk, T. U. Origin and evolution of antibiotic resistance: the common mechanisms of emergence and spread in water bodies. Front. Microbiol. 3, 18 (2012).
Lagier, J.-C. et al. Culture of previously uncultured members of the human gut microbiota by culturomics. Nat. Microbiol. 1, 16203 (2016).
Illergård, K., Ardell, D. H. & Elofsson, A. Structure is three to ten times more conserved than sequence—a study of structural response in protein cores. Proteins 77, 499–508 (2009).
Baquero, F., Tedim, A. P. & Coque, T. M. Antibiotic resistance shaping multi-level population biology of bacteria. Front. Microbiol. 4, 15 (2013).
Martínez, J. L., Coque, T. M. & Baquero, F. What is a resistance gene? Ranking risk in resistomes. Nat. Rev. Microbiol. 13, 116–123 (2015).
Van Boeckel, T. P. et al. Global antibiotic consumption 2000 to 2010: an analysis of national pharmaceutical sales data. Lancet. Infect. Dis. 14, 742–750 (2014).
Allen, H. K., Moe, L. A., Rodbumrer, J., Gaarder, A. & Handelsman, J. Functional metagenomics reveals diverse β- lactamases in a remote Alaskan soil. ISME J. 3, 243–251 (2009).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11, 119 (2010).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012).
Powell, S. et al. eggNOGv3.0: orthologous groups covering 1133 organisms at 41 different taxonomic ranges. Nucleic Acids Res. 40, D284–D289 (2012).
Liu, Y.-Y. et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect. Dis. 16, 161–168 (2016).
Kultima, J. R. et al. MOCAT: a metagenomics assembly and gene prediction toolkit. PLoS ONE 7, e47656 (2012).
Finn, R. D., Clements, J. & Eddy, S. R. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 39, W29–W37 (2011).
Pearson, W. R. & Lipman, D. J. Improved tools for biological sequence comparison. Proc. Natl Acad. Sci. USA 85, 2444–2448 (1988).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Jones, D. T. Protein secondary structure prediction based on position-specific scoring matrices. J. Mol. Biol. 292, 195–202 (1999).
Sali, A. & Blundell, T. L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Wallner, B. & Elofsson, A. Can correct protein models be identified?. Protein Sci. 12, 1073–1086 (2003).
Wiederstein, M. & Sippl, M. J. ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res. 35, W407–W410 (2007).
Ortiz, A. R., Strauss, C. E. M. & Olmea, O. MAMMOTH (matching molecular models obtained from theory): an automated method for model comparison. Protein Sci. Publ. Protein Soc. 11, 2606–2621 (2002).
Tibshirani, R. Regression shrinkage and selection via the lasso. J R. Stat. Soc B 58, 267–288 (1996).
Fan, R. E., Chang, K. W., Hsieh, C. J., Wang, X. R. & Lin, C. J. A library for large linear classification. JMLR 9, 1871–1874 .
Siguier, P., Perochon, J., Lestrade, L., Mahillon, J. & Chandler, M. ISfinder: the reference centre for bacterial insertion sequences. Nucleic Acids Res. 34, D32–D36 (2006).
Guglielmini, J., Quintais, L., Garcillán-Barcia, M. P., de la Cruz, F. & Rocha, E. P. C. The repertoire of ICE in prokaryotes underscores the unity, diversity, and ubiquity of conjugation. PLoS Genet. 7, e1002222 (2011).
Cury, J., Jové, T., Touchon, M., Néron, B. & Rocha, E. P. Identification and analysis of integrons and cassette arrays in bacterial genomes. Nucleic Acids Res. 44, 4539–4550 (2016).
Holmes, I., Harris, K. & Quince, C. Dirichlet multinomial mixtures: generative models for microbial metagenomics. PLoS ONE 7, e30126 (2012).
Ding, T. & Schloss, P. D. Dynamics and associations of microbial community types across the human body. Nature 509, 357–360 (2014).
Dray, S. & Legendre, P. Testing the species traits-environment relationships: the fourth-corner problem revisited. Ecology 89, 3400–3412 (2008).
Godon, J. J., Zumstein, E., Dabert, P., Habouzit, F. & Moletta, R. Molecular microbial diversity of an anaerobic digestor as determined by small-subunit rDNA sequence analysis. Appl. Environ. Microbiol. 63, 2802–2813 (1997).
Suau, A. et al. Direct analysis of genes encoding 16S rRNA from complex communities reveals many novel molecular species within the human gut. Appl. Environ. Microbiol. 65, 4799–4807 (1999).
Pons, N. et al. METEOR—A Platform for Quantitative Metagenomic Profiling of Complex Ecosystems (JOBIM Montpellier, 2010); http://www.jobim2010.fr/sites/default/files/presentations/27Pons.pdf
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Quereda, J. J. et al. Bacteriocin from epidemic Listeria strains alters the host intestinal microbiota to favor infection. Proc. Natl Acad. Sci. USA 113, 5706–5711 (2016).
Robert, P. & Escoufier, Y. A unifying tool for linear multivariate statistical methods: the RV-coefficient. Appl. Stat. 25, 257 (1976).
Acknowledgements
The authors are grateful to the GENOTOUL (Toulouse, France), GENOUEST (Rennes, France), ABIMS (Roscoff, France), MIGALE (Jouy-en-Josas) and TGCC-GENCI (Institut Curie) calculation clusters. The authors also thank B. Perichon (Institut Pasteur, Paris, France) for providing ARD sequences from Acinetobacter baumannii, P. Siguier (CNRS, Toulouse, France) for helping the search of insertion sequences with ISfinder, J. Guglielmini (Institut Pasteur, Paris, France) for his assistance in finding conjugative elements, S. Volant (Institut Pasteur, Paris, France) for the design of the statistical model in SHAMAN, T. Jové (University of Limoges, France) for his assistance in finding integrons, M. Petitjean (IAME Research Center, Paris, France) for her assistance in bioinformatic analyses, and F. Plaza-Oñate and M. Almeida for their help with MSPs. The project was funded in part by the European Union Seventh Framework Programme (FP7-HEALTH-2011-single-stage) under grant agreement number 282004, EvoTAR. IRYCIS authors acknowledge the European Development Regional Fund ‘A way to achieve Europe’ for co-founding the Spanish R&D National Plan 2012–2019 Work (PI15-0512), CIBER (CIBERESP; CB06/02/0053) and the Government of Madrid (InGeMICS- B2017/BMD-3691). V.F.L. was further funded by a Research Award Grant 2016 of the European Society for Clinical Microbiology and Infectious Diseases.
Author information
Authors and Affiliations
Contributions
E.R., A.G. and J.T. performed the analysis. E.R., A.G., J.T., W.v.S., A.d.B. and S.P.K. wrote the manuscript. A.S.A. and N.M. handled the data management. T.C., S.H.A., I.C. and J.L.M. performed the gene synthesis experiments. J.L.M., T.M.C., V.F.L., F.B., A.d.B., J.D., S.P.K., F.H. and S.D.E. discussed the protocol and results. L.M., T.G., V.d.L., N.A., B.F., I.W., A.A., W.v.S., M.R., X.Z. and R.J.L. recruited the patients and collected the samples. H.B., V.L., A.L. and F.L. handled the wet lab experiments. N.P., P.L. and J.M.B. managed the informatics and the calculation clusters. K.W. and N.P. designed the website (http://mgps.eu/Mustard/).
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–17, Supplementary Notes, Supplementary References.
Supplementary Table 1
The 6,095 pdARDs that were found in the 3.9 million protein catalogue. PCM score missing values means that the candidate could not be modelled with the negative template, so that the PCM score was considered to be over 50%.
Supplementary Table 2
Description of the 16 candidates sharing at least 40% amino acid identity with a reference ARD but being not predicted as an ARD by PCM. The TM score represents the degree of correct alignment of the structure generated by PCM and a reference structure (the highest score being 1).
Supplementary Table 3
Description of the 49 pdARDs found in plasmids and/or phages from GenBank.
Supplementary Table 4
Description of the 82 pdARDs shared by ≥2 species. Insertion sequences, conjugative elements and integrons were searched the same way as described in the Methods section.
Supplementary Table 5
Details on the 74 MGS that were found to be differentially abundant between subjects with no recent antibiotic exposure (n = 31) to antibiotics and subjects under chronic exposure to antibiotics (n = 30) using the Wald unpaired test. Padj, adjusted P-value (Benjamini–Hochberg correction).
Supplementary Table 6
Details on the 133 MGS that were found to be differentially abundant between subjects (n = 10) before and after SDD using the Wald paired test. Padj, adjusted P-value (Benjamini–Hochberg correction).
Supplementary Table 7
Predictions of ARDs in the functional metagenomics dataset from soils9 by the PCM method.
Rights and permissions
About this article
Cite this article
Ruppé, E., Ghozlane, A., Tap, J. et al. Prediction of the intestinal resistome by a three-dimensional structure-based method. Nat Microbiol 4, 112–123 (2019). https://doi.org/10.1038/s41564-018-0292-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-018-0292-6
This article is cited by
-
Sequence-structure-function characterization of the emerging tetracycline destructase family of antibiotic resistance enzymes
Communications Biology (2024)
-
The effect of long-term application of nitrogen-rich fertilizers on soil resistome: A study of conventional and organic cropping systems
Soil Ecology Letters (2024)
-
Perturbation and resilience of the gut microbiome up to 3 months after β-lactams exposure in healthy volunteers suggest an important role of microbial β-lactamases
Microbiome (2024)
-
Landscape of global urban environmental resistome and its association with local socioeconomic and medical status
Science China Life Sciences (2024)
-
Latent antibiotic resistance genes are abundant, diverse, and mobile in human, animal, and environmental microbiomes
Microbiome (2023)