Abstract
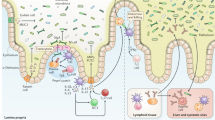
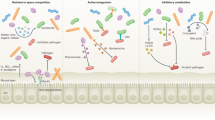
The human body is colonized by a diverse community of microorganisms collectively referred to as the microbiota. Here, we describe how the human microbiota influences susceptibility to infectious diseases using examples from the respiratory, gastrointestinal and female reproductive tract. We will discuss how interactions between the host, the indigenous microbiota and non-native microorganisms, including bacteria, viruses and fungi, can alter the outcome of infections. This Review Article will highlight the complex mechanisms by which the microbiota mediates colonization resistance, both directly and indirectly, against infectious agents. Strategies for the therapeutic modulation of the microbiota to prevent or treat infectious diseases will be discussed, and we will review potential therapies that directly target the microbiota, including prebiotics, probiotics, synbiotics and faecal microbiota transplantation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Aagaard, K. et al. The placenta harbors a unique microbiome. Sci. Transl. Med. 6, 237ra265 (2014).
Wampach, L. et al. Colonization and succession within the human gut microbiome by archaea, bacteria, and microeukaryotes during the first year of life. Front. Microbiol. 8, 738 (2017).
Nkamga, V. D., Henrissat, B. & Drancourt, M. Archaea: essential inhabitants of the human digestive microbiota. Hum. Microbiome J. 3, 1–8 (2017).
Parfrey, L. W., Walters, W. A. & Knight, R. Microbial eukaryotes in the human microbiome: ecology, evolution, and future directions. Front. Microbiol. 2, 153 (2011).
Reynolds, L. A. & Finlay, B. B. Early life factors that affect allergy development. Nat. Rev. Immunol. 17, 518–528 (2017).
Jandhyala, S. M. et al. Role of the normal gut microbiota. World J. Gastroenterol. 21, 8787–8803 (2015).
Su, C. et al. Helminth-induced alterations of the gut microbiota exacerbate bacterial colitis. Mucosal Immunol. 11, 144–157 (2018).
Lozupone, C. A., Stombaugh, J. I., Gordon, J. I., Jansson, J. K. & Knight, R. Diversity, stability and resilience of the human gut microbiota. Nature 489, 220–230 (2012).
Lawley, T. D. & Walker, A. W. Intestinal colonization resistance. Immunology 138, 1–11 (2013).
Koch, R. Die Ätiologie der milzbrand-krankheit, begründet auf die entwicklungsgeschichte des Bacillus anthracis. Cohns Beitr. Biol. Pflanzen 2, 277–310 (1876).
Falkow, S. Molecular Koch’s postulates applied to bacterial pathogenicity—a personal recollection 15 years later. Nat. Rev. Microbiol. 2, 67–72 (2004).
Young, V. B. The role of the microbiome in human health and disease: an introduction for clinicians. Br. Med. J. 356, j831 (2017).
Buffie, C. G. & Pamer, E. G. Microbiota-mediated colonization resistance against intestinal pathogens. Nat. Rev. Immunol. 13, 790–801 (2013).
Round, J. L. & Mazmanian, S. K. The gut microbiota shapes intestinal immune responses during health and disease. Nat. Rev. Immunol. 9, 313–323 (2009).
Walter, J., Maldonado-Gómez, M. X. & Martínez, I. To engraft or not to engraft: an ecological framework for gut microbiome modulation with live microbes. Curr. Opin. Biotechnol. 49, 129–139 (2018).
Costello, E. K., Stagaman, K., Dethlefsen, L., Bohannan, B. J. M. & Relman, D. A. The application of ecological theory toward an understanding of the human microbiome. Science 336, 1255–1262 (2012).
Shea, K. & Chesson, P. Community ecology theory as a framework for biological invasions. Trends Ecol. Evol. 17, 170–176 (2002).
Maldonado-Gomez, M. X. et al. Stable engraftment of Bifidobacterium longum AH1206 in the human gut depends on individualized features of the resident microbiome. Cell Host Microbe 20, 515–526 (2016).
Theriot, C. M. et al. Antibiotic-induced shifts in the mouse gut microbiome and metabolome increase susceptibility to Clostridium difficile infection. Nat. Commun. 5, 3114 (2014).
Jernberg, C., Lofmark, S., Edlund, C. & Jansson, J. K. Long-term ecological impacts of antibiotic administration on the human intestinal microbiota. ISME J. 1, 56–66 (2007).
Buffie, C. G. et al. Profound alterations of intestinal microbiota following a single dose of clindamycin results in sustained susceptibility to Clostridium difficile-induced colitis. Infect. Immun. 80, 62–73 (2012).
Jenior, M. L., Leslie, J. L., Young, V. B. & Schloss, P. D. Clostridium difficile colonizes alternative nutrient niches during infection across distinct murine gut microbiomes. mSystems 2, e00063-17 (2017).
Fleming-Davies, A. et al. In Women in Mathematical Biology (eds Layton, A. & Miller, L.) 137–161 (Springer, New York, 2017).
Theriot, C. M. & Young, V. B. Interactions between the gastrointestinal microbiome and Clostridium difficile. Annu. Rev. Microbiol. 69, 445–461 (2015).
Theriot, C. M., Bowman, A. A. & Young, V. B. Antibiotic-induced alterations of the gut microbiota alter secondary bile acid production and allow for Clostridium difficile spore germination and outgrowth in the large intestine. mSphere 1, e00045-15 (2016).
Thanissery, R., Winston, J. A. & Theriot, C. M. Inhibition of spore germination, growth, and toxin activity of clinically relevant C. difficile strains by gut microbiota derived secondary bile acids. Anaerobe 45, 86–100 (2017).
Kato, K., Lillehoj, E. P., Lu, W. & Kim, K. C. MUC1: the first respiratory mucin with an anti-inflammatory function. J. Clin. Med. 6, 110 (2017).
Andersch-Bjorkman, Y., Thomsson, K. A., Holmen Larsson, J. M., Ekerhovd, E. & Hansson, G. C. Large scale identification of proteins, mucins, and their O-glycosylation in the endocervical mucus during the menstrual cycle. Mol. Cell. Proteomics 6, 708–716 (2007).
Kim, K. C. Role of epithelial mucins during airway infection. Pulm. Pharmacol. Ther. 25, 415–419 (2012).
Lesuffleur, T., Zweibaum, A. & Real, F. X. Mucins in normal and neoplastic human gastrointestinal tissues. Crit. Rev. Oncol. Hematol. 17, 153–180 (1994).
Specian, R. D. & Neutra, M. R. Regulation of intestinal goblet cell secretion. I. Role of parasympathetic stimulation. Am. J. Physiol. 242, G370–G379 (1982).
Van der Sluis, M. et al. Muc2-deficient mice spontaneously develop colitis, indicating that MUC2 is critical for colonic protection. Gastroenterology 131, 117–129 (2006).
Schroeder, B. O. et al. Bifidobacteria or fiber protects against diet-induced microbiota-mediated colonic mucus feterioration. Cell Host Microbe 23, 27–40 (2018).
Hill, D. R. et al. Bacterial colonization stimulates a complex physiological response in the immature human intestinal epithelium. eLife 6, e29132 (2017).
Engevik, M. A. et al. Human Clostridium difficile infection: altered mucus production and composition. Am. J. Physiol. Gastrointest. Liver Physiol. 308, G510–G524 (2015).
Desai, M. S. et al. A dietary fiber-deprived gut microbiota degrades the colonic mucus barrier and enhances pathogen susceptibility. Cell 167, 1339–1353.e1321 (2016).
Ménard, S., Cerf-Bensussan, N. & Heyman, M. Multiple facets of intestinal permeability and epithelial handling of dietary antigens. Mucosal Immunol. 3, 247–259 (2010).
König, J. et al. Human intestinal barrier function in health and disease. Clin. Transl. Gastroenterol. 7, e196 (2016).
Zeissig, S. et al. Changes in expression and distribution of claudin 2, 5 and 8 lead to discontinuous tight junctions and barrier dysfunction in active Crohn’s disease. Gut 56, 61–72 (2007).
Fasano, A. et al. Vibrio cholerae produces a second enterotoxin, which affects intestinal tight junctions. Proc. Natl Acad. Sci. USA 88, 5242–5246 (1991).
Nusrat, A. et al. Clostridium difficile toxins disrupt epithelial barrier function by altering membrane microdomain localization of tight junction proteins. Infect. Immun. 69, 1329–1336 (2001).
Leslie, J. L. et al. Persistence and toxin production by Clostridium difficile within human intestinal organoids result in disruption of epithelial paracellular barrier function. Infect. Immun. 83, 138–145 (2015).
Hering, N. A. et al. Yersinia enterocolitica induces epithelial barrier dysfunction through regional tight junction changes in colonic HT-29/B6 cell monolayers. Lab. Invest. 91, 310–324 (2010).
Short, K. R. et al. Influenza virus damages the alveolar barrier by disrupting epithelial cell tight junctions. Eur. Respir. J. 47, 954–966 (2016).
De La Serre, C. B. et al. Propensity to high-fat diet-induced obesity in rats is associated with changes in the gut microbiota and gut inflammation. Am. J. Physiol. Gastrointest. Liver Physiol. 299, G440–G448 (2010).
Cani, P. D. et al. Selective increases of bifidobacteria in gut microflora improve high-fat-diet-induced diabetes in mice through a mechanism associated with endotoxaemia. Diabetologia 50, 2374–2383 (2007).
Ahmad, R., Rah, B., Bastola, D., Dhawan, P. & Singh, A. B. Obesity-induces organ and tissue specific tight junction restructuring and barrier deregulation by claudin switching. Sci. Rep. 7, 5125 (2017).
Zhang, L. J. & Gallo, R. L. Antimicrobial peptides. Curr. Biol. 26, R14–R19 (2016).
Jiang, Z. et al. Effects of net charge and the number of positively charged residues on the biological activity of amphipathic α-helical cationic antimicrobial peptides. Biopolymers 90, 369–383 (2008).
Wang, Z. & Wang, G. APD: the antimicrobial peptide database. Nucleic Acids Res. 32, D590–D592 (2004).
Muytjens, C. M. J., Yu, Y. & Diamandis, E. P. Discovery of antimicrobial peptides in cervical-vaginal fluid from healthy nonpregnant women via an integrated proteome and peptidome analysis. Proteomics 17, 1600461 (2017).
Hiemstra, P. S., Amatngalim, G. D., van der Does, A. M. & Taube, C. Antimicrobial peptides and innate lung defenses: role in infectious and noninfectious lung diseases and therapeutic applications. Chest 149, 545–551 (2016).
Natividad, J. M. et al. Differential induction of antimicrobial REGIII by the intestinal microbiota and Bifidobacterium breve NCC2950. Appl. Environ. Microbiol. 79, 7745–7754 (2013).
Wong, J. M., de Souza, R., Kendall, C. W., Emam, A. & Jenkins, D. J. Colonic health: fermentation and short chain fatty acids. J. Clin. Gastroenterol. 40, 235–243 (2006).
Zhao, Y. et al. GPR43 mediates microbiota metabolite SCFA regulation of antimicrobial peptide expression in intestinal epithelial cells via activation of mTOR and STAT3. Mucosal Immunol. 11, 752–762 (2018).
McHan, F. & Shotts, E. B. Effect of short-chain fatty acids on the growth of Salmonella typhimurium in an in vitro system. Avian Dis. 37, 396–398 (1993).
Horswill, A. R., Dudding, A. R. & Escalante-Semerena, J. C. Studies of propionate toxicity in Salmonella enterica identify 2-methylcitrate as a potent inhibitor of cell growth. J. Biol. Chem. 276, 19094–19101 (2001).
Jacobson, A. et al. A gut commensal-produced metabolite mediates colonization resistance to Salmonella infection. Cell Host Microbe 24, 296–307 (2018).
Ricke, S. C. Perspectives on the use of organic acids and short chain fatty acids as antimicrobials. Poult. Sci. 82, 632–639 (2003).
Hung, C. C. et al. The intestinal fatty acid propionate inhibits Salmonella invasion through the post-translational control of HilD. Mol. Microbiol. 87, 1045–1060 (2013).
Gantois, I. et al. Butyrate specifically down-regulates Salmonella pathogenicity island 1 gene expression. Appl. Environ. Microbiol. 72, 946–949 (2006).
Hryckowian, A. J. et al. Microbiota-accessible carbohydrates suppress Clostridium difficile infection in a murine model. Nat. Microbiol. 3, 662–669 (2018).
Benveniste, J., Lespinats, G. & Salomon, J. Serum and secretory IgA in axenic and holoxenic mice. J. Immunol. 107, 1656–1662 (1971).
Kawamoto, S. et al. The inhibitory receptor PD-1 regulates IgA selection and bacterial composition in the gut. Science 336, 485–489 (2012).
Kubinak, J. L. et al. MyD88 signaling in T cells directs IgA-mediated control of the microbiota to promote health. Cell Host Microbe 17, 153–163 (2015).
Lecuyer, E. et al. Segmented filamentous bacterium uses secondary and tertiary lymphoid tissues to induce gut IgA and specific T helper 17 cell responses. Immunity 40, 608–620 (2014).
Flannigan, K. L. et al. IL-17A-mediated neutrophil recruitment limits expansion of segmented filamentous bacteria. Mucosal Immunol. 10, 673–684 (2016).
Robak, O. H. et al. Antibiotic treatment-induced secondary IgA deficiency enhances susceptibility to Pseudomonas aeruginosa pneumonia. J. Clin. Invest. 128, 3535–3545 (2018).
Blander, J. M., Longman, R. S., Iliev, I. D., Sonnenberg, G. F. & Artis, D. Regulation of inflammation by microbiota interactions with the host. Nat. Immunol. 18, 851–860 (2017).
Sonnenburg, J. L. & Backhed, F. Diet–microbiota interactions as moderators of human metabolism. Nature 535, 56–64 (2016).
Verdu, E. F., Galipeau, H. J. & Jabri, B. Novel players in coeliac disease pathogenesis: role of the gut microbiota. Nat. Rev. Gastroenterol. Hepatol. 12, 497–506 (2015).
Bercik, P., Verdu, E. F. & Collins, S. M. Is irritable bowel syndrome a low-grade inflammatory bowel disease? Gastroenterol. Clin. North Am. 34, 235–245 (2005).
Martin, R. et al. Faecalibacterium prausnitzii prevents physiological damages in a chronic low-grade inflammation murine model. BMC Microbiol. 15, 67 (2015).
Zacho, J., Benfield, T., Tybjaerg-Hansen, A. & Nordestgaard, B. G. Increased baseline C-reactive protein concentrations are associated with increased risk of infections: results from 2 large Danish population cohorts. Clin. Chem. 62, 335–342 (2016).
Ravel, J. et al. Vaginal microbiome of reproductive-age women. Proc. Natl Acad. Sci. USA 108, 4680–4687 (2011).
Wessels, J. M. et al. Association of high-risk sexual behaviour with diversity of the vaginal microbiota and abundance of Lactobacillus. PLoS ONE 12, e0187612 (2017).
Borgdorff, H. et al. Lactobacillus-dominated cervicovaginal microbiota associated with reduced HIV/STI prevalence and genital HIV viral load in African women. ISME J. 8, 1781–1793 (2014).
Van de Wijgert, J. H. H. M. et al. The vaginal microbiota: what have we learned after a decade of molecular characterization? PLoS ONE 9, e105998 (2014).
MacIntyre, D. A. et al. The vaginal microbiome during pregnancy and the postpartum period in a European population. Sci. Rep. 5, 8988 (2015).
Borgdorff, H. et al. The association between ethnicity and vaginal microbiota composition in Amsterdam, the Netherlands. PLoS ONE 12, e0181135 (2017).
Fettweis, J. M. et al. Differences in vaginal microbiome in African American women versus women of European ancestry. Microbiology 160, 2272–2282 (2014).
Łaniewski, P. et al. Linking cervicovaginal immune signatures, HPV and microbiota composition in cervical carcinogenesis in non-Hispanic and Hispanic women. Sci. Rep. 8, 7593 (2018).
Redondo-Lopez, V., Cook, R. L. & Sobel, J. D. Emerging role of lactobacilli in the control and maintenance of the vaginal bacterial microflora. Rev. Infect. Dis. 12, 856–872 (1990).
Hillier, S. L., Krohn, M. A., Rabe, L. K., Klebanoff, S. J. & Eschenbach, D. A. The normal vaginal flora, H2O2-producing lactobacilli, and bacterial vaginosis in pregnant women. Clin. Infect. Dis. 16, S273–S281 (1993).
Sewankambo, N. et al. HIV-1 infection associated with abnormal vaginal flora morphology and bacterial vaginosis. Lancet 350, 546–550 (1997).
Gupta, K. et al. Inverse association of H2O2-producing lactobacilli and vaginal Escherichia coli colonization in women with recurrent urinary tract infections. J. Infect. Dis. 178, 446–450 (1998).
Wiesenfeld, H. C., Hillier, S. L., Krohn, M. A., Landers, D. V. & Sweet, R. L. Bacterial vaginosis is a strong predictor of Neisseria gonorrhoeae and Chlamydia trachomatis infection. Clin. Infect. Dis. 36, 663–668 (2003).
Lai, S. K. et al. Human immunodeficiency virus type 1 is trapped by acidic but not by neutralized human cervicovaginal mucus. J. Virol. 83, 11196–11200 (2009).
Di Paola, M. et al. Characterization of cervico-vaginal microbiota in women developing persistent high-risk human papillomavirus infection. Sci. Rep. 7, 10200 (2017).
Anahtar, M. N. et al. Cervicovaginal bacteria are a major modulator of host inflammatory responses in the female genital tract. Immunity 42, 965–976 (2015).
Keller, M. J. et al. Longitudinal assessment of systemic and genital tract inflammatory markers and endogenous genital tract E. coli inhibitory activity in HIV-infected and uninfected women. Am. J. Reprod. Immunol. 75, 631–642 (2016).
Doerflinger, S. Y., Throop, A. L. & Herbst-Kralovetz, M. M. Bacteria in the vaginal microbiome alter the innate immune response and barrier properties of the human vaginal epithelia in a species-specific manner. J. Infect. Dis. 209, 1989–1999 (2014).
Lennard, K. et al. Microbial composition predicts genital tract inflammation and persistent bacterial vaginosis in South African adolescent females. Infect. Immun. 86, e00410–17 (2018).
Masson, L. et al. Defining genital tract cytokine signatures of sexually transmitted infections and bacterial vaginosis in women at high risk of HIV infection: a cross-sectional study. Sex. Transm. Infect. 90, 580–587 (2014).
Martin, H. L. et al. Vaginal lactobacilli, microbial flora, and risk of human immunodeficiency virus type 1 and sexually transmitted disease acquisition. J. Infect. Dis. 180, 1863–1868 (1999).
McClelland, R. S. et al. Evaluation of the association between the concentrations of key vaginal bacteria and the increased risk of HIV acquisition in African women from five cohorts: a nested case-control study. Lancet Infect. Dis. 18, 554–564 (2018).
Furci, L., Sironi, F., Tolazzi, M., Vassena, L. & Lusso, P. α-defensins block the early steps of HIV-1 infection: interference with the binding of gp120 to CD4. Blood 109, 2928–2935 (2007).
Pace, B. T., Lackner, A. A., Porter, E. & Pahar, B. The role of defensins in HIV pathogenesis. Mediators Inflamm. 2017, 5186904 (2017).
Hearps, A. C. et al. Vaginal lactic acid elicits an anti-inflammatory response from human cervicovaginal epithelial cells and inhibits production of pro-inflammatory mediators associated with HIV acquisition. Mucosal Immunol. 10, 1480–1490 (2017).
Pecora, D. V. A comparison of transtracheal aspiration with other methods of determining the bacterial flora of the lower respiratory tract. N. Eng. J. Med. 269, 664–666 (1963).
Dickson, R. P. et al. Analysis of culture-dependent versus culture-independent techniques for identification of bacteria in clinically obtained bronchoalveolar lavage fluid. J. Clin. Microbiol. 52, 3605–3613 (2014).
Sibley, C. D. et al. Culture enriched molecular profiling of the cystic fibrosis airway microbiome. PLoS ONE 6, e22702 (2011).
Collins, A. M. et al. Bronchoalveolar lavage (BAL) for research; obtaining adequate sample yield. J. Vis. Exp. 85, e4345 (2014).
Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Segal, L. N. et al. Enrichment of lung microbiome with supraglottic taxa is associated with increased pulmonary inflammation. Microbiome 1, 19 (2013).
Dickson, R. P. et al. Spatial variation in the healthy human lung microbiome and the adapted island model of lung biogeography. Ann. Am. Thorac. Soc. 12, 821–830 (2015).
Hilty, M. et al. Disordered microbial communities in asthmatic airways. PLoS ONE 5, e8578 (2010).
Charlson, E. S. et al. Assessing bacterial populations in the lung by replicate analysis of samples from the upper and lower respiratory tracts. PLoS ONE 7, e42786 (2012).
Bassis, C. M. et al. Analysis of the upper respiratory tract microbiotas as the source of the lung and gastric microbiotas in healthy individuals. mBio 6, e00037 (2015).
Dickson, R. P. et al. Bacterial topography of the healthy human lower respiratory tract. mBio https://doi.org/10.1128/mBio.02287-16 (2017).
Dickson, R. P. & Huffnagle, G. B. The lung microbiome: new principles for respiratory bacteriology in health and disease. PLoS Pathog. 11, e1004923 (2015).
Dickson, R. P., Erb-Downward, J. R. & Huffnagle, G. B. Homeostasis and its disruption in the lung microbiome. Am. J. Physiol. Lung Cell. Mol. Physiol. 309, L1047–L1055 (2015).
Shenoy, M. K. et al. Immune response and mortality risk relate to distinct lung microbiomes in patients with HIV and pneumonia. Am. J. Respir. Crit. Care Med. 195, 104–114 (2016).
Segal, L. N. et al. Enrichment of the lung microbiome with oral taxa is associated with lung inflammation of a Th17 phenotype. Nat. Microbiol. 1, 16031 (2016).
Murdock, B. J. et al. Interleukin-17 drives pulmonary eosinophilia following repeated exposure to Aspergillus fumigatus conidia. Infect. Immun. 80, 1424–1436 (2012).
Thomas, D. W. et al. Probiotics and prebiotics in pediatrics. Pediatrics 126, 1217–1231 (2010).
Hill, C. et al. Expert consensus document. The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat. Rev. Gastroenterol. Hepatol. 11, 506–514 (2014).
Luoto, R. et al. Prebiotic and probiotic supplementation prevents rhinovirus infections in preterm infants: a randomized, placebo-controlled trial. J. Allergy Clin. Immunol. 133, 405–413 (2014).
Lohner, S., Kullenberg, D., Antes, G., Decsi, T. & Meerpohl, J. J. Prebiotics in healthy infants and children for prevention of acute infectious diseases: a systematic review and meta-analysis. Nutr. Rev. 72, 523–531 (2014).
Rycroft, C. E., Jones, M. R., Gibson, G. R. & Rastall, R. A. A comparative in vitro evaluation of the fermentation properties of prebiotic oligosaccharides. J. Appl. Microbiol. 91, 878–887 (2001).
Marx, S. P., Winkler, S. & Hartmeier, W. Metabolization of β-(2,6)-linked fructose-oligosaccharides by different bifidobacteria. FEMS Microbiol. Lett. 182, 163–169 (2000).
Guigoz, Y., Rochat, F., Perruisseau-Carrier, G., Rochat, I. & Schiffrin, E. J. Effects of oligosaccharide on the faecal flora and non-specific immune system in elderly people. Nutr. Res. 22, 13–25 (2002).
Koga, Y. et al. Age-associated effect of kestose on Faecalibacterium prausnitzii and symptoms in the atopic dermatitis infants. Pediatr. Res. 80, 844–851 (2016).
Pokusaeva, K., Fitzgerald, G. F. & van Sinderen, D. Carbohydrate metabolism in Bifidobacteria. Genes Nutr. 6, 285–306 (2011).
Pan, X.-D., Chen, F.-Q., Wu, T.-X., Tang, H.-G. & Zhao, Z.-Y. Prebiotic oligosaccharides change the concentrations of short-chain fatty acids and the microbial population of mouse bowel. J. Zhejiang. Univ. Sci. B 10, 258–263 (2009).
Correa-Oliveira, R., Fachi, J. L., Vieira, A., Sato, F. T. & Vinolo, M. A. Regulation of immune cell function by short-chain fatty acids. Clin. Transl. Immunol. 5, e73 (2016).
Maslowski, K. M. et al. Regulation of inflammatory responses by gut microbiota and chemoattractant receptor GPR43. Nature 461, 1282–1286 (2009).
Venkataraman, A. et al. Variable responses of human microbiomes to dietary supplementation with resistant starch. Microbiome 4, 33 (2016).
Kunz, C., Rudloff, S., Baier, W., Klein, N. & Strobel, S. Oligosaccharides in human milk: structural, functional, and metabolic aspects. Annu. Rev. Nutr. 20, 699–722 (2000).
Shoaf, K., Mulvey, G. L., Armstrong, G. D. & Hutkins, R. W. Prebiotic galactooligosaccharides reduce adherence of enteropathogenic Escherichia coli to tissue culture cells. Infect. Immun. 74, 6920–6928 (2006).
Zhang, K. et al. The non-toxigenic Clostridium difficile CD37 protects mice against infection with a BI/NAP1/027 type of C. difficile strain. Anaerobe 36, 49–52 (2015).
Gerding, D. N. et al. Administration of spores of nontoxigenic Clostridium difficile strain M3 for prevention of recurrent C. difficile infection: a randomized clinical trial. JAMA 313, 1719–1727 (2015).
Cotter, P. D., Hill, C. & Ross, R. P. Bacteriocins: developing innate immunity for food. Nat. Rev. Microbiol. 3, 777–788 (2005).
Servin, A. L. Antagonistic activities of lactobacilli and bifidobacteria against microbial pathogens. FEMS Microbiol. Rev. 28, 405–440 (2004).
Mukai, T., Kaneko, S., Matsumoto, M. & Ohori, H. Binding of Bifidobacterium bifidum and Lactobacillus reuteri to the carbohydrate moieties of intestinal glycolipids recognized by peanut agglutinin. Int. J. Food Microbiol. 90, 357–362 (2004).
Liu, L. et al. Global, regional, and national causes of child mortality in 2000–13, with projections to inform post-2015 priorities: an updated systematic analysis. Lancet 385, 430–440 (2015).
Thaver, D. & Zaidi, A. K. Burden of neonatal infections in developing countries: a review of evidence from community-based studies. Pediatr. Infect. Dis. J. 28, S3–S9 (2009).
Panigrahi, P. et al. A randomized synbiotic trial to prevent sepsis among infants in rural India. Nature 548, 407–412 (2017).
Kassam, Z., Lee, C. H., Yuan, Y. & Hunt, R. H. Fecal microbiota transplantation for Clostridium difficile infection: systematic review and meta-analysis. Am. J. Gastroenterol. 108, 500–508 (2013).
Van Nood, E. et al. Duodenal infusion of donor feces for recurrent Clostridium difficile. N. Engl. J. Med. 368, 407–415 (2013).
Manges, A. R., Steiner, T. S. & Wright, A. J. Fecal microbiota transplantation for the intestinal decolonization of extensively antimicrobial-resistant opportunistic pathogens: a review. Infect. Dis. (Lond) 48, 587–592 (2016).
Khoruts, A. & Sadowsky, M. J. Understanding the mechanisms of faecal microbiota transplantation. Nat. Rev. Gastroenterol. Hepatol. 13, 508–516 (2016).
Weingarden, A. et al. Dynamic changes in short- and long-term bacterial composition following fecal microbiota transplantation for recurrent Clostridium difficile infection. Microbiome 3, 10 (2015).
Hamilton, M. J., Weingarden, A. R., Unno, T., Khoruts, A. & Sadowsky, M. J. High-throughput DNA sequence analysis reveals stable engraftment of gut microbiota following transplantation of previously frozen fecal bacteria. Gut Microbes 4, 125–135 (2013).
Seekatz, A. M. et al. Recovery of the gut microbiome following fecal microbiota transplantation. mBio 5, e00893-14 (2014).
Smillie, C. S. et al. Strain tracking reveals the determinants of bacterial engraftment in the human gut following fecal microbiota transplantation. Cell Host Microbe 23, 229–240.e5 (2018).
Staley, C., Kelly, C. R., Brandt, L. J., Khoruts, A. & Sadowsky, M. J. Complete microbiota engraftment is not essential for recovery from recurrent Clostridium difficile infection following fecal microbiota transplantation. mBio 7, e01965-16 (2016).
Ott, S. J. et al. Efficacy of sterile fecal filtrate transfer for treating patients with Clostridium difficile infection. Gastroenterology 152, 799–811.e7 (2017).
Zuo, T. et al. Bacteriophage transfer during faecal microbiota transplantation in Clostridium difficile infection is associated with treatment outcome. Gut 67, 634–643 (2018).
Morton, E. R. et al. Variation in rural African gut microbiota is strongly correlated with colonization by Entamoeba and subsistence. PLoS Genet. 11, e1005658 (2015).
Gilchrist, C. A. et al. Role of the gut microbiota of children in diarrhea due to the protozoan parasite Entamoeba histolytica. J. Infect. Dis. 213, 1579–1585 (2016).
Burgess, S. L., Gilchrist, C. A., Lynn, T. C. & Petri, W. A. Jr Parasitic protozoa and interactions with the host intestinal microbiota. Infect. Immun. 85, e00101–17 (2017).
Sui, Y. et al. Influence of gut microbiome on mucosal immune activation and SHIV viral transmission in naive macaques. Mucosal Immunol. 11, 1219–1229 (2018).
Dillon, S. M. et al. An altered intestinal mucosal microbiome in HIV-1 infection is associated with mucosal and systemic immune activation and endotoxemia. Mucosal Immunol. 7, 983–994 (2014).
Ichinohe, T. et al. Microbiota regulates immune defense against respiratory tract influenza A virus infection. Proc. Natl Acad. Sci. USA 108, 5354–5359 (2011).
Rudner, X. L., Happel, K. I., Young, E. A. & Shellito, J. E. Interleukin-23 (IL-23)-IL-17 cytokine axis in murine Pneumocystis carinii infection. Infect. Immun. 75, 3055–3061 (2007).
Lukacs, N. W. et al. Respiratory virus-induced TLR7 activation controls IL-17-associated increased mucus via IL-23 regulation. J. Immunol. 185, 2231–2239 (2010).
Jespers, V. et al. A longitudinal analysis of the vaginal microbiota and vaginal immune mediators in women from sub-Saharan Africa. Sci. Rep. 7, 11974 (2017).
Gosmann, C. et al. Lactobacillus-deficient cervicovaginal bacterial communities are associated with increased HIV acquisition in young South African women. Immunity 46, 29–37 (2017).
Christensen-Quick, A. et al. Human Th17 cells lack HIV-inhibitory RNases and are highly permissive to productive HIV infection. J. Virol. 90, 7833–7847 (2016).
Prescott, S. L. History of medicine: origin of the term microbiome and why it matters. Hum. Microbiome J. 4, 24–25 (2017).
Acknowledgements
The authors thank D. R. Hill, K. Rao and C. M. Bassis for helpful feedback on earlier versions of this manuscript, and D. R. Hill for many helpful conversations regarding this manuscript. This work was supported by a grant awarded to V.B.Y. from the National Institute of Allergy and Infectious Diseases at the National Institutes of Health (U01-AI124255).
Author information
Authors and Affiliations
Contributions
All authors contributed to the conceptualization, writing and preparation of this manuscript, as well as the creation of the figures.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Libertucci, J., Young, V.B. The role of the microbiota in infectious diseases. Nat Microbiol 4, 35–45 (2019). https://doi.org/10.1038/s41564-018-0278-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-018-0278-4
This article is cited by
-
Integrative study of pulmonary microbiome, transcriptome and clinical outcomes in Mycoplasma pneumoniae pneumonia
Respiratory Research (2024)
-
Two intestinal microbiota-derived metabolites, deoxycholic acid and butyrate, synergize to enhance host defense peptide synthesis and alleviate necrotic enteritis
Journal of Animal Science and Biotechnology (2024)
-
Interactions between Balantidium ctenopharyngodoni and microbiota reveal its low pathogenicity in the hindgut of grass carp
BMC Microbiology (2024)
-
Assessment of the nasal microbiota in dogs with fungal rhinitis before and after cure and in dogs with chronic idiopathic rhinitis
BMC Microbiology (2023)
-
The interplay between human papillomavirus and vaginal microbiota in cervical cancer development
Virology Journal (2023)