Abstract
The endoplasmic reticulum (ER) is an architecturally diverse organelle that serves as a membrane source for the replication of multiple viruses. Flaviviruses, including yellow fever virus, West Nile virus, dengue virus and Zika virus, induce unique single-membrane ER invaginations that house the viral replication machinery1. Whether this virus-induced ER remodelling is vulnerable to antiviral pathways is unknown. Here, we show that flavivirus replication at the ER is targeted by the interferon (IFN) response. Through genome-scale CRISPR screening, we uncovered an antiviral mechanism mediated by a functional gene pairing between IFI6 (encoding IFN-α-inducible protein 6), an IFN-stimulated gene cloned over 30 years ago2, and HSPA5, which encodes the ER-resident heat shock protein 70 chaperone BiP. We reveal that IFI6 is an ER-localized integral membrane effector that is stabilized through interactions with BiP. Mechanistically, IFI6 prophylactically protects uninfected cells by preventing the formation of virus-induced ER membrane invaginations. Notably, IFI6 has little effect on other mammalian RNA viruses, including the related Flaviviridae family member hepatitis C virus, which replicates in double-membrane vesicles that protrude outwards from the ER. These findings support a model in which the IFN response is armed with a membrane-targeted effector that discriminately blocks the establishment of virus-specific ER microenvironments that are required for replication.
Similar content being viewed by others
Main
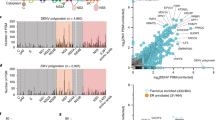
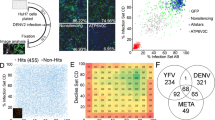
Viruses in the Flavivirus genus cause substantial human disease. Innate immune control of these viruses is mediated by interferons (IFNs)3, which induce the transcription of hundreds of IFN-stimulated genes (ISGs), some of which encode antiviral effector proteins4. To identify genes that regulate the IFN response to a flavivirus, we used a genome-scale clustered regularly interspaced short palindromic repeats (CRISPR) approach to screen for single-guide RNAs (sgRNAs) that rendered cells permissive to infection with yellow fever virus expressing Venus (YFV-Venus) despite treatment with a highly suppressive dose of IFN-α (Fig. 1a). Multiple hits from the screen reconstituted the IFN signalling pathway, as expected, or were factors involved in mRNA processing (Fig. 1b,c and Supplementary Table 1). Comparing hits to our annotated list of 448 human ISGs revealed only three overlapping genes: IFI6 (encoding IFN-α-inducible protein 6) and two IFN signalling ISGs (STAT2, encoding signal transducer and activator of transcription 2, and IRF9, encoding IFN regulatory factor 9)4 (Fig. 1d). Aside from IFI6, which was among the most significant hits (P = 2.7 × 10−13), no other classical ISGs were found with this analysis (Fig. 1d).
a, Schematic of genome-wide CRISPR screen to identify genes required for the IFN-α-induced antiviral response to YFV. MFI, mean fluorescence intensity; FSC, forward scatter. b, Manhattan dot plot of genome-wide CRISPR screen results with significance of enrichment calculated by the MAGeCK method. Genes with an FDR < 0.01 (dotted line) are coloured. The numbers in blue represent the percentage of GFP-positive cells. c, IFN signalling pathway with CRISPR hits in bold. ISRE, interferon-stimulated response element. d, Overlap of CRISPR screening hits with common ISGs. e, Western blot of Huh7.5 or Huh7.5-IFI6-KO1 cells treated with IFN. Uncropped blots are presented in Supplementary Fig. 1a. WT, wild type. f, IFN-α dose response curves in Huh7.5 or Huh7.5-IFI6-KO1 cells challenged with WNV-GFP. g, Time course of WNV production in Huh7.5-NT or Huh7.5-IFI6-KO2 cells with or without a single dose of IFN-α (1,000 U ml−1) pre-treatment (top). n = 3; ****P = 0.0001 and *P = 0.048 (two-way ANOVA on log-transformed data, Dunnet post-test). Western blots corresponding to the WNV time course (bottom). h–k, Huh7.5 cells transduced with lentivirus expressing IFI6 or an empty (Ctrl) vector were infected with viruses at 0.5–2.0 MOI as follows: time courses of YFV-Venus (h) and DENV-GFP (i), single-step growth curve of YFV (j) and infectivity of ZIKV-GFP (MR766) (k). Infectivity was monitored by FACS for GFP-based viruses and by plaque assay for non-reporter YFV. For panels h–j, n = 3; *P = 0.0173, **P = 0.0029 and ****P = 1.1 × 10−5 (two-way ANOVA, Sidak post-test). For panel k, n = 4; **P = 0.0026 (two-tailed, unpaired t-test). l, ZIKV titres in fetal human neural progenitors transduced with control or IFI6-expressing lentivirus. n = 3; *P = 0.022 (two-tailed, unpaired t-test). m, RT–PCR analysis of IFI6 mRNA induction by IFN-α (log-fold dosing from 0.1–1,000 U ml−1) in NHDF cells. n = 2. n,o, Western blot analysis of IFI6 induction by IFN-α in NHDF cells (n) or by DENV in NHDF, NHDF-NT and NHDF-IFI6-KO1 cells (o). p,q, Titres of DENV (p) or WNV (q) in NHDF, NHDF-NT and NHDF-IFI6-KO cells. In panel p, n = 3; *P = 0.032 and **P = 0.003 (one-way ANOVA on log-transformed data, Tukey post-test). In panel q, n = 1 (24 h) and n = 3 (48 h and 72 h); *P = 0.012 and **P = 0.008 (two-tailed, unpaired t-test). NS, not significant. In all graphs, the data points and error bars are explained in the Methods section (under ‘Statistics and reproducibility’).
To confirm this result, we used two strategies to deplete IFI6 via CRISPR. In the first strategy, we used a puromycin-selectable lentiCRISPRv2 vector to deliver one sgRNA targeting IFI6 (IFI6-KO1). In the second strategy, we used both puromycin-selectable and blasticidin-selectable lentiCRISPRv2 vectors to co-deliver two distinct IFI6-targeted sgRNAs (IFI6-KO2) that were also different from the sgRNA in the IFI6-KO1 strategy. We generated an IFI6-knockout line, Huh7.5-IFI6-KO1, and showed loss of IFN-induced IFI6 expression by western blot (Fig. 1e and Supplementary Fig. 1a). In infection studies, compared to control Huh7.5 cells, Huh7.5-IFI6-KO1 cells were markedly refractory to an otherwise robust IFN-mediated inhibition of West Nile virus expressing green fluorescent protein (WNV-GFP) (Fig. 1f) and YFV-Venus (Supplementary Fig. 1b). Similar results with YFV-Venus were obtained in two other cell backgrounds, A549-IFI6-KO1 and U-2 OS IFI6-KO1, and in Huh7.5-IFI6-KO2 cells (Supplementary Fig. 1c–e). To determine the effects of IFN-inducible IFI6 on a non-attenuated, non-reporter flavivirus, we assessed WNV titres in Huh7.5-NT (control cells expressing a non-targeting sgRNA) or in Huh7.5-IFI6-KO2 cells with or without IFN pre-treatment (Fig. 1g). In the absence of IFN, WNV replicated to similar titres in Huh7.5-NT and Huh7.5-IFI6-KO2 cells. IFN suppressed WNV replication by approximately 3 logs in Huh7.5-NT cells at 24 h and 48 h. At 72 h, the inhibitory effect was diminished and this correlated with decreased IFI6 expression by western blot. Notably, WNV titres were nearly completely restored at all time points in IFN-treated Huh7.5-IFI6-KO2 cells. To complement these IFI6 depletion studies, we also demonstrated that ectopically expressed IFI6 potently suppressed YFV-Venus in multiple cells types (Supplementary Fig. 1f). Time courses revealed potent and sustained IFI6-mediated inhibition of YFV-Venus and dengue virus expressing GFP (DENV-GFP) (Fig. 1h,i), as well as 2–3 log10 suppression of non-reporter YFV titres (Fig. 1j). Contemporary and prototype strains of Zika virus (ZIKV) were also inhibited by IFI6 (Fig. 1k and Supplementary Fig. 1g), although to a lesser extent than the other flaviviruses that we tested. These data establish a robust inhibitory phenotype for IFI6 against flaviviruses. In previous ectopic expression or short hairpin RNA knockdown screens, we and others also found IFI6 as inhibitory towards flaviviruses, further strengthening the reproducibility of the phenotype through multiple screening approaches4,5,6,7.
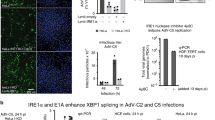
We next assessed the antiviral potential of IFI6 in more biologically relevant primary human cells. Ectopic expression of IFI6 suppressed ZIKV production in human fetal neural progenitors (Fig. 1l). IFI6 was robustly induced in both IFN-treated (Fig. 1m,n) and DENV-infected (Fig. 1o) primary normal human dermal fibroblasts (NHDFs). Despite poor replication of DENV in NHDFs, we observed enhanced DENV titres in CRIPSR-targeted NHDF-IFI6-KO1 cells relative to control cells (Fig. 1p). Similarly, when comparing WNV replication in NHDF-IFI6-KO2 cells relative to control NHDF-NT cells, we found that titres were 11-fold or 18-fold higher at 48 h or 72 h post-infection, respectively (Fig. 1q). Together, these data indicate that IFI6 is necessary for a robust anti-flavivirus IFN response, is sufficient to prophylactically suppress infection and can inhibit flavivirus replication in primary human cells.
IFI6 encodes a small, 130 amino acid protein in the IFI-6-16 family, which has four conserved paralogues in humans (IFI6, IFI27, IFI27L1 and IFI27L2) and three in mice (Ifi27, Ifi27l2a Ifi27l2b)8 (Supplementary Fig. 2a). IFI6 is only present in higher-order mammals. In humans, only IFI6 and IFI27 are IFN inducible2,9 (Supplementary Fig. 2b). However, IFI27 did not inhibit YFV infection (Supplementary Fig. 2c), which suggests distinct functions for these related proteins.
As IFI6 overexpression suppressed infection similar to IFN-α (compare Fig. 1f and Fig. 1h), we tested whether IFI6 modulates cellular antiviral gene expression. In contrast to the antiviral transcription factor IRF1, ectopically expressed IFI6 did not induce the classical ISGs IFIT1, IFITM3, OAS2 and RSAD2, or significantly alter the global cellular transcriptome (Supplementary Fig. 3a,b). Furthermore, Huh7.5-IFI6-KO1 cells treated with IFN-α were not impaired in their ability to induce other ISGs (Supplementary Fig. 3c). These data indicate that IFI6 does not mediate antiviral signalling and is probably a direct effector.
Another screening hit, HSPA5, encodes BiP (also known as GRP78), a heat shock protein 70 chaperone in the endoplasmic reticulum (ER) that assists in protein folding and the surveillance of misfolded proteins10. We hypothesized that BiP may modulate the antiviral effects of IFI6. We first assessed IFI6 and BiP mRNA and protein levels in IFN-treated cells (Fig. 2a). As expected, IFI6 mRNA and protein were highly induced over a time course of IFN treatment. HSPA5 mRNA levels were largely unaffected by IFN, whereas BiP protein levels increased slightly at each time point after IFN treatment. To validate the CRISPR screen, we depleted BiP via CRISPR and found that YFV infection was de-repressed by more than fivefold in the presence of IFN-α (Fig. 2b). In the absence of IFN-α, we detected a modest 1.5-fold loss of infection in BiP-depleted cells, suggesting a minor role for this chaperone during YFV infection. We then knocked out BiP in cells ectopically expressing IFI6 and observed a similar (approximately fourfold) de-repression of YFV infection as with IFN-α treatment (Fig. 2c). The antiviral activity of IFN-α could be restored in BiP-depleted cells by complementing cells with sgRNA-resistant BiP (Fig. 2d). Notably, the depletion of BiP resulted in near-complete loss of ectopic IFI6 expression (Fig. 2e). These data indicate that, in the absence of BiP, IFI6 is targeted for degradation. This was confirmed by restoring IFI6 expression in BiP-depleted cells with the proteasome inhibitor MG132 (Fig. 2f). To test whether BiP chaperone activity was required for its regulation of IFI6, we reconstituted BiP-depleted cells with CRISPR-resistant wild-type BiP or three ATPase-defective mutants11. The mutants could not rescue IFI6 levels (Fig. 2g), suggesting that BiP chaperone activity is required for IFI6 protein expression. To assess potential interactions between BiP and IFI6, we immunoprecipitated ectopically expressed FLAG-tagged IFI6 and found that BiP was indeed pulled down (Fig. 2h). The interaction was abrogated by ATP, which binds to the chaperone and induces a conformational change that releases substrates12. We next tested whether endogenous, IFN-induced IFI6 interacts with BiP. As our IFI6 antibody has a high background (Supplementary Fig. 1a), we used CRISPR-mediated ‘gene tagging’ to introduce a 3×FLAG epitope at the carboxy terminus of endogenous IFI6 in U-2 OS cells13. PCR with reverse transcription (RT–PCR) and western blotting confirmed IFN-α-mediated induction of endogenous FLAG-tagged IFI6 (Fig. 2i). In a proximity ligation assay, we detected a robust interaction signal for BiP and IFI6 in IFN-treated gene-tagged cells (Fig. 2j and Supplementary Fig. 4a). Thus, BiP probably functions as a regulator of the IFN response to flaviviruses by facilitating proper folding and/or localization of IFI6 at the ER membrane.

The functional link between IFI6 and BiP, a lumenal ER chaperone, was unexpected as several reports indicate that IFI6 is a mitochondrial protein14,15,16. Thus, we sought to resolve IFI6 subcellular localization. The amino terminus of IFI6 has a predicted signal peptide (Supplementary Fig. 4b), suggesting targeting to the secretory pathway (ER, Golgi, vesicles, and so on). Indeed, the expression of IFI6 bearing a C-terminal 3× FLAG epitope (IFI6-3×F) revealed a reticular staining pattern that overlapped with the ER translocon member Sec61β-GFP, but was distinct from mitochondria-localized GFP (Fig. 3a). As overexpression can mislocalize proteins, we assessed the localization of endogenous IFI6 in our gene-tagged cells (Fig. 2i). Immunofluorescence of IFN-treated, gene-tagged cells demonstrated a reticular pattern for endogenous IFI6 that overlapped with Sec61β-GFP (Fig. 3b) and was similar to ectopically expressed IFI6-3×F (Fig. 3a). Immunogold electron microscopy in COS-7 cells revealed that ectopically expressed IFI6 bearing a C-terminal haemagglutinin (HA) tag (IFI6-HA) localized extensively to cytosolic membranes that resembled ER structures, but not within nuclei or mitochondria (Fig. 3c). We confirmed by immunofluorescence that IFI6-HA staining in COS-7 cells was reticular and distinct from a mitochondrial GFP signal (Fig. 3d). Finally, we fused the putative N-terminal signal peptide of IFI6 to GFP and found that the signal from this fusion protein, (N32)IFI6-GFP, overlapped with the common ER epitope KDEL (Fig. 3e). Taken together, these results suggest that, contrary to previous reports, IFI6 is localized to the ER, the site of flavivirus replication.
a, Immunofluorescence microscopy of Huh7.5 cells stably expressing IFI6-3×F and transfected with plasmids expressing Sec61β-GFP (top) or mitochondrial-targeted GFP (Mito-GFP) (bottom). Cells were stained with anti-FLAG antibody (red) or anti-GFP antibody (green). Scale bars, 10 μm. b, Gene-tagged U-2 OS cells (Fig. 2i) expressing Sec61β-GFP were treated with IFN-α (1,000 U ml−1) and analysed by immunofluorescence microscopy. Scale bars, 10 μm. c, Immunogold electron microscopy of COS-7 cells stably expressing IFI6-HA. Cy, cytoplasm; Mi, mitochondria; Nu, nucleus. d, Immunofluorescence microscopy of COS-7 cells expressing IFI6-HA (red) and mitochondrial-targeted GFP (green). Scale bars, 10 μm. e, Immunofluorescence microscopy of COS-7 cells transiently expressing (N32)IFI6-GFP. Cells were stained with anti-KDEL antibody (red) or anti-GFP antibody (green). SP, signal peptide. Scale bars, 10 μm. f, Membrane flotation assay. Huh7.5 cells expressing IFI6-3×F were lysed in the presence of 1 M NaCl, 0.1 M Na2CO3 (pH 11.5) or 0.5% Triton X-100, and membranes were fractionated on a Histodenz gradient by ultracentrifugation. Collected fractions were analysed by western blot with antibodies against calnexin or FLAG. g, Secretion assay in 293T cells. Cells transiently expressing IFI6-3×F or CCL2-1×F were incubated in serum-free media with or without 1 μg ml−1 Brefeldin A (BFA) for 5 h. Supernatants (Sup) and cell lysates were analysed by western blot with anti-FLAG or anti-actin antibodies.
IFI6 is highly hydrophobic and has putative transmembrane helices (Supplementary Fig. 4c,d), suggesting that it may be membrane associated. To test this, we monitored IFI6-3×F migration in a membrane flotation assay17,18. IFI6-3×F migrated to similar fractions as the ER integral membrane protein calnexin (Fig. 3f). Solubilization of membranes with Triton X-100 shifted the entire pool of IFI6 and calnexin to higher densities. By contrast, disrupting electrostatic interactions with a high salt concentration did not affect the migration pattern. Treatment with sodium carbonate (pH 11.5), which disrupts peripheral membrane protein association19, caused a small proportion of IFI6 to shift to higher densities, although most of the signal was retained at a lower density. Similar results were obtained with a different gradient composition (Supplementary Fig. 4e). These data indicate that IFI6 is primarily an integral membrane protein, although we cannot rule out that some of the protein may associate to membranes through peripheral interactions. Supporting this conclusion, we did not detect secretion of IFI6 from cells, whereas the secretion of CC-motif ligand 2 (CCL2), a control, was robust and readily blocked by Brefeldin A treatment (Fig. 3g).
To gain insight into the mechanism of action of IFI6, we assessed which viral replication step was targeted. We first used a reporter DENV expressing Phytonus pyralis firefly luciferase (Fluc)7. This virus recapitulates the full replication cycle, with Fluc expressed at low levels during post-entry primary viral RNA translation and at higher levels during genome amplification. IFI6 suppressed DENV-Fluc at late, but not early, time points, suggesting that viral entry and primary RNA translation were probably intact, whereas viral genome replication was inhibited (Fig. 4a). Similar results were obtained with a YFV subgenomic replicon expressing Renilla reniformis luciferase (Rluc) (Supplementary Fig. 5a), which uncouples viral RNA translation and replication from entry and virion morphogenesis20.
a, DENV-Fluc infectivity in control or IFI6-expressing Huh7.5 cells, monitored by luciferase relative light units (RLU). n = 3; ****P = 8.5 × 10−5 (24 h), ****P = 1 × 10−13 (48 h) and ****P = 3 × 10−13 (72 h) (two-way ANOVA on log-transformed data, Sidak post-test). b, Transmission electron micrographs of control or IFI6-expressing cells infected with YFV. Scale bars, 0.5 μM. c, YFV-infected control or IFI6-expressing cells were stained with antibody against dsRNA (green) or with Hoechst stain (blue). Scale bars, 20 μM. d,e, Time-of-addition experiment. Huh7.5 cells were transduced with control or IFI6-expressing lentivirus at various time points prior to, with or after YFV infection (d). Two days later, infectivity was quantified by FACS (e, top). n = 3; *P = 0.0115, ****P = 3.8 × 10−7 (−8 h) and ****P = 7.5 × 10−9 (−24 h) (two-way ANOVA, Sidak post-test). Protein levels for viral NS1 and IFI6 were determined by western blot (e, bottom). f, Laser scanning confocal microscopy of Huh7.5 cells expressing IFI6-3×F and infected with DENV. Cells were stained with anti-FLAG antibody (red) or anti-NS4B antibody (green). Scale bars, 10 μM. For cell 2, the white line in the ‘zoom’ panel represents the plane for the z-stack shown in the three-dimensional (3D) side view. g, Pearson correlation coefficients of IFI6 and NS4B localization in 12 cells. n = 3. h, HCV-Gluc infectivity in control, IFI6-expressing or IRF1-expressing Huh7.5 cells monitored by luciferase RLU. n = 3; ****P = 0.0001 (two-way ANOVA on log-transformed data, Dunnett post-test). i, HCV-Ypet infectivity in control or IFI6-expressing cells, monitored by FACS. n = 3 (two-tailed, unpaired t-test). j, HCV-Ypet infectivity in Huh7.5-NT or Huh7.5-IFI6-KO2 cells with or without a single dose of IFN-α (100 U ml−1) pre-treatment. n = 3 (two-way ANOVA, Sidak post-test). k, Co-infection of YFV-Venus and HCV-Gluc in Huh7.5 cells. HCV replication was monitored by Gluc RLU in cell supernatants; YFV infectivity was monitored by FACS. n = 3; ****P = 2 × 10−11 (two-tailed, unpaired t-test). l, Human coronavirus infectivity in control or IFI6-expressing Huh7.5 cells, monitored by FACS. n = 3 (two-tailed, unpaired t-test). m, A model of IFI6 antiviral function. IFN-inducible IFI6 localizes to the ER membrane, is stabilized by BiP and prevents the formation of flavivirus-induced invaginations. IFI6 has no effect on viruses that induce double-membrane vesicles during replication (for example, HCV). NS, not significant. In all graphs, the data points and error bars are explained in the Methods section (under ‘Statistics and reproducibility’).
After viral RNA translation, the flavivirus polyprotein is cleaved by both the viral protease NS3 and the host signal peptidases to initiate replication. We tested whether IFI6 affected cleavage of DENV proteins or a known host target, stimulator of interferon genes protein (STING)21. IFI6 had no effect on the ability of DENV NS2B–NS3 to cleave itself or STING (Supplementary Fig. 6a). Similarly, ectopically expressed DENV C-prM and 2K-NS4B, which are both processed in a signal peptidase complex subunit 1 (SPCS1)-dependent manner22, were properly cleaved in the presence and absence of IFI6 (Supplementary Fig. 6b,c). Together, these data indicate that IFI6 does not interfere with polyprotein processing events mediated by the viral protease or SPCS1.
After polyprotein processing, flavivirus infection is characterized by dramatic rearrangement of ER membranes, including the formation of single-membrane invaginations that house the replication machinery1. Using transmission electron microscopy, we found that IFI6 expression markedly suppressed the formation of these ‘replication organelles’ (Fig. 4b). This correlated with near-complete suppression of viral double-stranded RNA (dsRNA) replication intermediates (Fig. 4c). We attempted to overcome IFI6-mediated inhibition by rescuing a serially passaged YFV mutant. However, after passaging YFV 20 times in IFI6-expressing cells, we were unable to rescue an escape variant (Supplementary Fig. 7), indicating that the IFI6-mediated replication block is not easily overcome by mutation. Taken together, these replication cycle studies suggest that IFI6 specifically targets an early step in the process of genome amplification, after viral entry, translation and polyprotein processing.
Two models may explain how IFI6 inhibits flavivirus replication: in model 1, IFI6 inhibits the viral polymerase or viral RNA amplification, thereby preventing the production of viral proteins that induce replication organelle formation; in model 2, IFI6 has no effect on replication per se, but rather interferes with the ability of the viral proteins to invaginate the ER membrane, which is required for robust replication. To distinguish these two possibilities, we performed a time-of-addition experiment in which IFI6 was expressed either before or after infection. We found that expressing IFI6 at the same time as or after viral infection rendered IFI6 incapable of inhibiting replication, as shown by infectivity assays and viral NS1 protein expression (Fig. 4d,e). Thus, despite being expressed in virally infected cells, IFI6 cannot inhibit viral replication once it is already established. This ‘point-of-no-return’ phenotype indicates that IFI6 is primarily a prophylactic antiviral effector and suggests that it may target virus-induced membrane alterations at the ER (model 2) rather than viral genome amplification (model 1). In support of this conclusion, when we infected IFI6-expressing cells at a high enough multiplicity of infection (MOI) of DENV to partially overcome the antiviral effect, we observed that IFI6 colocalized with ER-resident DENV NS4B (Fig. 4f,g, and Supplementary Fig. 8). We also found that, in some cells, IFI6 relocalized to NS4B-rich large cytoplasmic punctae, which are characteristic of DENV infection17 (Fig. 4f). As NS4B helps to form the membrane scaffold that is required for replication organelle biogenesis23,24,25, we propose that IFI6, if expressed before infection, suppresses flavivirus replication by thwarting virus-induced ER perturbations.
Our model also predicts that IFI6 would not target RNA viruses that do not replicate on ER membranes. Indeed, ectopic expression of IFI6 did not inhibit other diverse RNA viruses, including Sindbis virus (SINV; Togaviridae family), coxsackie B virus (CVB; Picornaviridae family) or measles virus (Paramyxoviridae family) (Supplementary Fig. 5b). The loss of IFI6 did not affect IFN-mediated inhibition of SINV or CVB infection (Supplementary Fig. 5c). We next tested the inhibitory potential of IFI6 against hepatitis C virus (HCV), which, like DENV or YFV, is a Flaviviridae family member (genus Hepacivirus). Using multiple HCV constructs in both gain-of-function and loss-of-function assays, we found that IFI6 lacked inhibitory activity against HCV (Fig. 4h–j and Supplementary Fig. 5d). This was unexpected as HCV also replicates at the ER. However, rather than induce single-membrane invaginations like the flaviviruses, HCV infection results in the formation of double-membrane vesicles that protrude out from the ER1. To determine whether HCV subverts the antiviral effects of IFI6, which was proposed recently26, we co-infected cells with both YFV and HCV reporter viruses, predicting that IFI6 may have reduced activity against YFV if HCV counteracts its function. However, IFI6 still inhibited YFV, but not HCV (Fig. 4k), suggesting that HCV does not antagonize IFI6 in a manner that interferes with its anti-flavivirus activity (Supplementary Fig. 5e). Other groups have reported that ectopically expressed IFI6 inhibits HCV27,28, but the magnitude of inhibition in those studies was typically on the order of twofold, rather than the multiple log-fold inhibition that we observed with flaviviruses. Moreover, supporting our data, three independent loss-of-function screens to identify genes that mediate the IFN response to HCV failed to identify IFI6 as a hit29,30,31. Beyond HCV, we also found that IFI6 does not inhibit a coronavirus (Fig. 4l), which, like HCV, forms double-membrane vesicles at the ER32. Together, these data support a model in which IFI6 inhibits ER membrane perturbations specific to flaviviruses, but not to other viruses that generate different ER structures (Fig. 4m).
Our study reveals IFI6 to be an ER integral membrane protein and key IFN-inducible antiviral effector. The unique viral specificity of IFI6 suggests a mechanism that may be tailored for flavivirus-induced single-membrane invaginations at the ER, but not the ER-derived double-membrane vesicles formed during the replication of HCV and coronavirus. If the host has evolved multiple mechanisms to inhibit membrane-derived viral replication compartments, then future studies may reveal IFI6-like molecules that target other viruses replicating at the ER or distinct membranous organelles. Our findings further raise the intriguing possibility that this antiviral strategy may be leveraged for therapeutic intervention of flavivirus infection.
Methods
Viruses and cells
Huh7.5, U-2 OS, A549, 293T (from C. Rice, The Rockefeller University), 293T (from N. Conrad, UT Southwestern Medical Center) and COS-7 (from N. Alto, UT Southwestern Medical Center) cells and all derivatives were maintained in ‘complete’ DMEM (Gibco) supplemented with 10% FBS (Gibco) and 1× non-essential amino acids (NEAA; Gibco). Human STAT1−/− fibroblasts (from J.-L. Casanova, The Rockefeller University) were maintained in RPMI supplemented with 10% FBS (Gibco) and 1× NEAA (Gibco). BHK21-J (from C. Rice) were grown in MEM (Gibco) supplemented with 10% FBS and 1× NEAA. Stable cells expressing antibiotic resistance genes were grown in complete media supplemented with puromycin (Sigma) at 4 μg ml−1 or blasticidin (Gibco) at 15 μg ml−1. NHDFs were purchased from Lonza and cultured in fibroblast basal media (American Type Culture Collection (ATCC) or Lonza) as recommended by the supplier. Human fetal neural progenitors were cultured as previously described33. Cell lines were routinely tested for mycoplasma using a PCR-based assay (Vendor GeM Mycoplasma Detection Kit, MP0025-1KT, Sigma). When applicable, cell lines were authenticated with short tandem repeat (STR) analysis using the ATCC Cell Line Authentication service.
The generation and propagation of the following viruses or replicons have been previously described4,5: YFV strain 17D expressing Venus GFP (YFV-Venus), HCV genotype 2a intragenotypic chimera expressing Ypet GFP (HCV-Ypet), CVB-GFP, Sindbis virus AR86 expressing GFP (SINV-GFP), WNV-GFP, measles virus Edmonston lineage expressing GFP, HCV replicon expressing Gaussia luciferase (Bi-Gluc-JFH-SG) and ZIKV strain PRVABC59 (ref. 33). Infectious HCV-Gluc was generated from the infectious clone Jc1FLAG(p7-nsGluc2A) as previously described34. An infectious clone of non-reporter WNV (strain TX02) was kindly provided by I. Frolov (University of Alabama Birmingham) and the virus was propagated as described above for WNV-GFP. DENV-Fluc with a L52F mutation in the gene encoding NS4B was derived from pDENV2-IC30P-A and propagated as previously described7. A ZIKV MR766-GFP infectious clone (kindly provided by M. Evans, Icahn School of Medicine at Mount Sinai) was used to generate the virus as described35. The infectious clone pACNR-FLYF-17Dx (kindly provided by C. Rice) was used to generate non-reporter YFV-17D. Briefly, the plasmid was linearized with XhoI and the purified DNA was used as a template for transcription with the mMessage mMachine SP6 Transcription kit (Thermo Fisher). RNA was purified with the RNeasy Mini Kit (Qiagen) and electroporated into BHK21-J cells. Virus-containing supernatants were collected, clarified by centrifugation and stored at −80 °C. Human coronavirus OC43 (ATCC strain VR-1558) was propagated in HCT-8 cells as specified by the ATCC. Viral titres were determined by antibody staining (MAB9012, Millipore) and flow cytometry36.
Plasmids and molecular cloning
To generate a C-terminal 3×FLAG-tagged IFI6 variant, pENTR221.IFI6 from the ISG library previously described4 was digested with PstI and XhoI (NEB). Full-length IFI6 containing a glycine-serine linker was PCR amplified from pENTR221.IFI6. A 3×FLAG epitope was PCR amplified from pcDNA4/TO/GFP-3×FLAG (kindly provided by I. D’Orso). The three fragments were combined with the Gibson Assembly Cloning Kit (New England Biolabs) according to the manufacturer’s instructions, to give the final pENTR221.IFI6-3×FLAG construct. A similar HA-tagged construct was generated by replacing the 3×FLAG epitope with annealed oligos encoding the HA epitope, to generate pENTR221.IFI6-HA.
To generate a GFP expressing the putative IFI6 signal peptide at its N terminus, full-length enhanced GFP (eGFP) was PCR amplified with primers containing SacI or XhoI sites and directionally cloned into a SacI/XhoI-digested pENTR221.IFI6 vector. The resulting plasmid was named pENTR221.IFI6(N32)-eGFP.
To generate CCL2 with a 1×FLAG tag on the C terminus, pENTR221.CCL2 from the ISG library previously described4 was modified as follows. A DNA fragment containing a 1×FLAG coding sequence flanked by CCL2 homology arms was synthesized and inserted into pENTR221.CCL2 with the Gibson Assembly Cloning Kit (New England Biolabs) according to the manufacturer’s instructions.
To generate DENV-2K-NS4B-HA, the 2K-NS4B region from pDENV2-IC30P-A was PCR amplified and cloned into a lentiviral expression plasmid, pTRIP.XKB-GFP (gift from C. Rice), replacing the GFP sequence. A similar construct expressing DENV C-prM was generated by PCR.
The Gateway-compatible lentiviral SCRPSY-DEST plasmid co-expressing TagRFP and a puromycin resistance cassette has been previously described7. A derivative of this plasmid in which TagRFP was replaced with a nuclear-localized TagBFP (Evrogen) was generated and named SCRPSY-DEST-nlsBFP. A second derivative of SCRPSY-DEST, named SCRBBL-DEST, was generated by removing the TagRFP-2A-PuroR cassette and subcloning a PCR-amplified blasticidin resistance gene in its place. The previously described pTRIP.CMV.IVSB.ires.TagRFP-DEST vector4 was modified to remove the IRES-TagRFP cassette, generating a non-reporter Gateway-compatible lentiviral vector named pTRIP.CMV.IVSB-DEST. Lentiviral expression constructs were generated by combining ENTR and DEST vectors in the recombination reaction using LR Clonase II (Invitrogen) according to the manufacturer’s instructions. All pENTR constructs were propagated in DH5-α cells, whereas lentiviral vectors were grown in DH5-α or MDS42RecA cells (Scarab Genomics).
To complement cells that had genomic BiP edited by CRISPR, an overexpression construct of BiP containing six silent mutations in the region targeted by BiP CRISPR guide 3 was generated. A synthetic gene fragment of BiP containing silent mutations was cloned into pENTR221-BiP (kindly provided by N. Alto) after digestion with AflII and PmlI restriction enzymes, using Gibson Assembly Cloning Kit (New England Biolabs) according to the manufacturer’s instructions. Constructs expressing catalytic mutant versions of guide 3 CRISPR-resistant BiP (T37G, E201G and T229G) were generated using site-directed mutagenesis and verified by sequencing. The oligo sequences used in this study are listed in Supplementary Table 2.
Lentiviral transduction, virus infections and replicon studies
Lentiviral production and transductions were performed as previously described4. Viral infections for GFP-expressing reporter viruses and for ZIKV were carried out as previously described4,5,33. For the non-reporter YFV-17D growth curve, cells were infected with a MOI of 2 in DMEM supplemented with 1% FBS for 1 h. Media were aspirated and cells were washed with serum-free DMEM four times and replaced with 500 μl complete DMEM. Virus yields in supernatants were quantified by plaque assay. Studies using the HCV subgenomic replicon (Bi-Gluc-JFH[SG]) were carried out as previously described4. The YFV-17D subgenomic replicon YFRP-Rluc (kindly provided by R. Kuhn) was propagated and used to generate viral RNA as previously described20. Assays to detect YFV-Rluc replicon activity were conducted similar to HCV replicon studies, with the detection of intracellular Rluc using the Renilla Luciferase Assay System (Promega). For IFN-mediated inhibition studies, cells were treated with the indicated dose of human IFN-α2a (11100-1, PBL Assay Science) for 4 h or 16 h prior to infection. Infections proceeded for approximately one round of viral replication and cells were collected for analysis by flow cytometry. For studies in human fetal neural progenitors, approximately 100,000 cells per well were plated onto 24-well plates. Two or three days later, cells were transduced with SCRPSY-Empty or SCRPSY-IFI6 lentivirus for 2 days in 1 ml human fetal neural progenitor proliferation media. Two days post-transduction, cells were infected with approximately 0.5 MOI ZIKV (PRVABC59) for 1–2 h. Cells were washed three times with media and supernatants were collected at 24, 48 and 72 h. Supernatants were titred by limiting dilution on STAT1−/− fibroblasts, using 4G2 (D1-4G2-4-15, ATCC) staining as the readout36.
CRISPR–Cas9 cloning, gene targeting and viral infection studies
Oligos encoding sgRNAs for generating knockout cells using CRISPR–Cas9 were cloned into the lentiCRISPRv2 plasmid (a gift from F. Zhang, Addgene plasmid 52961) as previously described37,38. The oligo sequences for sgRNAs targeting IFI6, BiP or non-targeting controls are listed in Supplementary Table 2. LentiCRISPRv2 clones containing guide sequences were sequenced, purified and used for lentiviral production as described above. For generating heterogeneous knockout cell populations, Huh7.5, A549 or U-2 OS cells were infected with the lentiCRISPRv2-derived lentivirus for 48 h, then reseeded into complete DMEM containing 1–4 µg ml−1 puromycin for 3 days to select for transduced cells.
Single-cell clones of Huh7.5 cells targeted for IFI6 knockout via CRISPR could not be propagated. To overcome this limitation, CRISPR-targeted Huh7.5 cells (using sgRNA ‘BR2’) were diluted with parental Huh7.5 cells at a ratio of 1/2,000. Mixed cells were plated at 100 cells per well in 96-well plates. Once confluent, cells were passaged to a 48-well format in the presence of 4 μg ml−1 puromycin to kill off non-targeted cells. Surviving populations derived in this manner were propagated and expanded for 6 weeks before cryopreserving stock cultures. Using this strategy, an IFI6-targeted knockout cell line was generated and named ‘IFI6-KO1’. In another strategy, two distinct guides targeting IFI6 (‘g1g2’) were co-expressed, one by lentiCRISPRv2 (blasticidin selectable) and the other by lentiCRISPRv2 (puromycin selectable). Double drug selection led to bulk populations of cells, with each cell receiving two independent CRISPR guides targeting IFI6. These were named ‘IFI6-KO2’ cells.
The following methods were used for cells targeted via the IFI6-KO1 strategy. The day before IFN treatment and infection, 70,000–150,000 cells were plated onto 24-well plates. Cells were pre-treated with serial dilutions of IFN-α 4 h before infection. The IFN-containing media were removed and cells were incubated with 0.5–1.0 MOI virus (CVB-GFP, SINV-GFP, YFV-Venus or WNV-GFP) for 1–2 h before being brought to volume with complete DMEM. After approximately one viral life cycle, cells were harvested and analysed by flow cytometry.
The following methods were used for cells targeted via the IFI6-KO2 strategy. For YFV-Venus infections: 100,000 cells were plated in 24-well plates. Cells were treated with 1,000 U ml−1 IFN-α overnight at time of plating. The next day, cells were infected with 7 MOI of YFV-Venus for 2 h at 37 °C. Cells were harvested 24 h later for flow cytometry analysis. For HCV infection: 80,000 cells were plated in 24-well plates. Cells were treated with 100 U ml−1 IFN-α overnight at time of plating. The next day, cells were infected with approximately 1 MOI of HCV (BiYPetJC1Flag2) for 1 h at 37 °C. Cells were harvested for flow cytometry analysis 48 h later. For WNV infections: 100,000 cells were plated in 24-well plates. Cells were treated with 1,000 U ml−1 IFN-α overnight at time of plating. The next day, cells were infected with 0.01 MOI of WNV for 2 h at 37 °C. Cells were washed four times with DMEM/3% FBS media. Supernatant was collected at 24, 48 and 72 h and titred by plaque assay on BHK cells.
Genome-wide CRISPR screen
The genome-wide CRISPR screen using the Brunello CRISPR knockout pooled library is described in detail in Supplementary Methods. Briefly, Huh7.5 cells were transduced with the Brunello CRISPR library. Puromycin-selected cell populations were treated with IFN-α overnight, followed by YFV-Venus infection for 24 h. Venus-positive cells that were no longer sensitive to IFN-α-mediated inhibition were collected by fluorescence-activated cell sorting (FACS). Genomic DNA was isolated from these cells, and sgRNA sequences were amplified by PCR and deep sequenced. Data were analysed using the MAGeCK method39.
NHDF experiments
For lentiviral transductions, NHDFs were plated at approximately 100,000 cells per well onto 6-well plates. The next day, cells were transduced via spinoculation with lentiCRISPRv2 (expressing non-targeting sgRNAs or sgRNAs targeting IFI6) in 2 ml fibroblast basal media at 37 °C for at 800g. Two days post-transduction, cells were pooled and placed under selection with 4 μg ml−1 puromycin and 15 μg ml−1 blasticidin. Three days after selection, cells were plated for experiments or passaged once before plating for experiments. For DENV infections, 100,000 cells per well were plated onto 6-well plates. The next day, cells were infected with DENV at an MOI of 2 for 2 h in 1 ml fibroblast basal media. The supernatant was aspirated and cells were washed four times with 500 μl media and 1.5 ml media was added back. Forty-eight hours post-infection, supernatants were collected and titred by limiting dilution on STAT1−/− fibroblasts as described above. Cell lysates were collected for western blot to detect IFI6 expression. For WNV infections, 40,000–80,000 cells per well were plated onto 6-well plates. The next day, cells were infected with WNV at an MOI of 0.01 for 1 h in 1 ml fibroblast basal media. The supernatant was aspirated and cells were washed four times with 500 µl media and a final volume of 1 ml was placed on the cells. Supernatant was collected at 24, 48 and 72 h for titring by plaque assay on BHK cells. To detect IFI6 induction by IFN in NHDFs, 100,000 cells per well were plated onto 6-well plates (for RNA isolation) or 200,000 cells were plated onto 6-cm dishes (for protein isolation). Cells were treated with IFN at indicated doses for 4 h or 24 h. Cells collected for RNA were processed using the Qiagen RNeasy protocol and IFI6 mRNA was detected by quantitative RT–PCR as described above. For protein detection, cells were collected, resuspended in lysis buffer and mixed with 1× SDS loading buffer lacking β-mercaptoethanol (BME). Samples were sonicated but not boiled, and loaded onto Tris-tricine low-molecular-weight gels and analysed for IFI6 expression as described above.
Endogenous IFI6 gene tagging in U-2 OS cells
Oligos encoding an sgRNA near the C-terminal coding region of IFI6 were cloned into LentiCRISPRV2 as described above. An IFI6-based donor vector containing homology arms flanking the protospacer adjacent motif site by 800 bp in both directions and a 3×FLAG sequence was synthesized in a pUC57 backbone (Genewiz). The protospacer adjacent motif site codons were altered to avoid re-targeting of the site once DNA repair had occurred. For transfections, 200,000 U-2 OS cells were seeded onto 6-well plates in complete DMEM containing 0.1 μM SCR-7, a DNA ligase IV inhibitor (Tocris). The donor vector and the LentiCRISPRv2 vector were transfected with X-tremeGENE 9 (Roche) into U-2 OS cells at a ratio of 2/1 with a final DNA amount of 2 μg per well. Two days post-transfection, cells were re-plated in complete DMEM with 1 μg ml−1 puromycin and 0.1 μM SCR-7 for 24 h, then replaced with complete DMEM with 0.1 μM SCR-7 and no puromycin. Cells were maintained for at least 1 week before use in experiments.
Immunofluorescence and confocal microscopy
Cells were fixed in 4% paraformaldehyde in PBS for 15 min, permeabilized with 0.2% Triton-X 100 in PBS for 5 min and blocked with 10% BSA, 5% goat serum and 50 mM glycine in PBS for 30 min. Primary antibody incubation for 2 h at room temperature was followed by secondary incubation with an Alexa Fluor-conjugated secondary antibody (AF-488 for green channels and AF-555 for red channels, Life Technologies). Cells were mounted with ProLong Diamond with DAPI (Life Technologies) and imaged with a Zeiss Axiovert 200 microscope or Zeiss Observer Z.1 microscope, unless otherwise indicated.
dsRNA
Approximately 20,000 Huh7.5 cells stably expressing IFI6 or an empty vector were plated into 8-well chamber slides. Cells were infected with 1.5 MOI YFV-17D for 1 h. Cells were incubated for 48 h and, following the protocol described above, were stained with J2 dsRNA antibody (1:200, Scicons) and Hoechst stain (1:12,000, Thermo Fisher) and mounted with ProLong Gold (Life Technologies).
Sec61β-mEmerald and pTRIP.Mito-GFP
mEmerald-Sec61-C-18 was a gift from M. Davidson (Addgene plasmid 54249). COS-7 or Huh7.5 stably expressing IFI6 3×FLAG were plated at 5,000 cells per well into 8-well chamber slides. The next day, cells were transfected with 75 ng per well (Huh7.5) or 20 ng per well (COS-7) of mEmerald-Sec61β or 75 ng per well of pTRIP.Mito.eGFP (kindly provided by C. Rice). Cells were stained as described above with anti-GFP antibody (1:4,000, 6556, Abcam) and FLAG-M2 (1:1,000, 3165, Sigma).
U-2 OS HDR cells
U-2 OS cells were plated at 5,000 cells per well onto 8-well chamber slides. The next day, cells were transfected with 75 ng per well of mEmerald-Sec61β and 6 h post-transfection treated with 1,000 U ml−1 IFN-α. Cells were stained with antibodies against FLAG and GFP as described above.
N32-GFP localization
COS-7 cells were plated at 5,000 cells per well onto 8-well chamber slides. The next day, cells were transfected with 75 ng per well of SCRPSY.IFI6(N32)GFP-nlsBFP. Twenty-four hours post-transfection, cells were stained with antibodies against FLAG and the ER retention KDEL sequence (1:250, ADI-SPA-827, Enzo).
DENV NS4B localization
IFI6-3×F-expressing Huh7.5 cells were plated at 5,000 cells per well into 8-well chamber slides. Cells were infected with DENV at a MOI of 2 for 2 h. Cells were incubated for 48 h and stained with antibodies targeting DENV NS4B (1:200, Thermo Fisher) and FLAG (1:1,000, Sigma). Images were acquired using a Zeiss 880 laser scanning confocal microscope with Airyscan for super-resolution capability. Images were deconvolved using AutoQuant X3. Deconvolved images were analysed for colocalization using Imaris 7.7.2, where a background subtraction was performed and a threshold for pixel intensity was automatically determined by the software. The Pearson values shown indicate the overlap in the automatically determined region of interest.
Electron microscopy and immunogold labelling
Approximately 2 × 106 Huh7.5 cells stably transduced with SCRPSY.empty or SCRPSY.IFI6 lentivirus were seeded into 10-cm2 tissue culture dishes. The next day, cells were mock infected or infected with YFV-17D at 1 MOI. Cells were processed 24 h post-infection for electron microscopy as previously described33. For immunogold labelling, COS-7 cells stably expressing SCRPSY.IFI6-HA were fixed for 30 min at room temperature with 4% paraformaldehyde and 0.1% glutaraldehyde in 0.1 M sodium phosphate buffer (pH 7.4). Cells were permeabilized with 0.25% saponin in phosphate buffer for 30 min and blocked with 5% goat serum in 0.01% saponin in phosphate buffer for 1 h. Cells were then incubated with anti-HA antibody (1:1,000 dilution, 901501, BioLegend) overnight at 4 °C. The next day, cells were washed four times with phosphate buffer, followed by incubation with 1.4-nm gold-conjugated fragment antigen-binding goat antibodies to mouse IgG (1:100, 7202, Nanoprobes) for 2 h at room temperature. After washing five times with phosphate buffer, cells were further fixed with 1% glutaraldehyde and washed three times with phosphate buffer. After rinsing with water, the immunogold-labelled samples were gold enhanced for 2.5 min using the gold enhancement kit (Nanoprobes) and washed again with water and 0.1 M sodium cacodylate buffer. Cells were then post-fixed in 1% osmium tetroxide and 0.8% K3[Fe(CN6)] in 0.1 M sodium cacodylate buffer for 1 h at room temperature and en bloc stained with 2% aqueous uranyl acetate. Samples were then dehydrated with increasing concentrations of ethanol, infiltrated with Embed-812 resin and polymerized in a 60 °C oven overnight. Blocks were sectioned with a diamond knife (Diatome) on a Leica Ultracut UCT (7) ultramicrotome (Leica Microsystems) and collected onto copper grids. Images were acquired on a Tecnai G2 spirit transmission electron microscope (FEI) equipped with a LaB6 source using a voltage of 120 kV.
RNA and protein detection in cell cultures
Quantitative RT–PCR for ISGs
For gene expression assays, total RNA was isolated from cells stably expressing IFI6 or, alternatively, after treatment with 0 or 100 U ml−1 IFN-α. RNA was isolated using a RNeasy Mini Kit (Qiagen). Total RNA (50 ng) was analysed by quantitative RT–PCR using the QuantiFast SYBR Green RT–PCR kit (Qiagen). Commercially available QuantiTect primers specific for IFI27, IFI27L1, IFI27L2, IRF1, IFITM3, RSAD2, IFIT1, HSPA5 and the housekeeping control gene RPS11 (Qiagen) were used according to the manufacturer’s instructions. Laboratory-generated IFI6 and OAS2 primer sets were also used (Supplementary Table 2). Reactions were run on an ABI7500 Fast Real Time PCR System and gene expression was calculated using the ΔΔCT method.
RNA sequencing
Gene expression analysis by RNA sequencing was performed as previously described40. The RNA sequencing data have been deposited to the NCBI Gene Expression Omnibus with the accession number GSE105771.
Western blot
For protein expression assays, cells were lysed in RIPA buffer (25 mM Tris (pH 7.5), 150 mM NaCl, 0.1% SDS, 0.5% sodium deoxycholate, 1% Triton X-100 and 1× Complete Protease Inhibitor Cocktail (Roche)) or NP-40 lysis buffer (50 mM Tris (pH 7.5), 150 mM NaCl, 1 mM EDTA, 1% NP-40 and 1× Complete Protease Inhibitor Cocktail) to obtain a post-nuclear lysate. The protein concentration of cell lysates was determined by Bradford Assay (Pierce). Alternatively, cell pellets were directly boiled in 2× SDS–PAGE sample buffer (100 mM Tris (pH 6.8), 20% glycerol, 4% SDS, 2% BME and 0.1% Bromophenol blue) or 2× Tricine sample buffer (200 mM Tris-HCl (pH 6.8), 40% glycerol, 2% SDS and 0.04% Coomassie blue). Lysates were separated on 12% polyacrylamide gels using the Laemmli method. For endogenous IFI6 detection, 10% Tris-Tricine gels were used. Proteins were blotted to PVDF (polyvinylidene difluoride) membranes (Bio-Rad) and processed for western blotting. Blots were blocked overnight in 3% or 5% milk in 1– TBST (50 mM Tris-Cl (pH 7.5), 150 mM NaCl and 0.05% Tween-20), followed by incubation with primary and secondary antibodies for 1 h and 30 min, respectively. Proteins were visualized by incubating blots with enhanced chemiluminescent substrate (ECL, Pierce) and exposing blots to autoradiography film (Denville Scientific). The antibodies used in the study include: anti-FLAG M2 (F3165, Sigma), anti-FLAG polyclonal (F7425, Sigma), anti-β-actin (ab6276, Abcam), anti-BiP (PA5-34941, Thermo Fisher), anti-calnexin (ADI-SPA-860-D, Enzo Life Sciences), anti-KDEL (ADI-SPA-827D, Enzo Life Science), anti-RFP (AB233, Evrogen), anti-DENV NS4A (GTX132069, GeneTex), anti-NS1 (gift from M. Diamond), rabbit IgG control (ab27478, Abcam), mouse IgG control (ab81032, Abcam), goat anti-rabbit horseradish peroxidase and goat anti-mouse horseradish peroxidase (Pierce). For the detection of endogenous IFI6, a rabbit polyclonal antibody was custom generated by ProSci, Inc. The antigen consisted of a peptide corresponding to the C terminus of IFI6 (LMGYATHKYLDSEED) and containing an N-terminal cysteine to assist in conjugation to the carrier protein. Antibody was further isolated by immunoaffinity purification of the serum.
Membrane flotation assay
Huh7.5 cells stably expressing IFI6-3×FLAG were collected from confluent 15-cm2 tissue culture dishes and resuspended in 2 ml cold 0.25 M sucrose. Cells were lysed in a 2-ml tight-fitting Dounce homogenizer with 200 strokes on ice to yield approximately 90% lysis. The lysate was centrifuged at 2,500g for 10 min at 4 °C to pellet debris. The supernatant was transferred to a new tube and pelleted at 20,000g for 10 min at 4 °C to pellet membranes. Pellets were resuspended in 2 ml PBS, PBS with 1 M NaCl, PBS with 0.1 M Na2CO3 (pH 11.5) or PBS with 0.5% Triton X-100 and incubated on ice for 30 min. The samples were mixed with 2 ml cold 60% Histodenz (Sigma) and transferred to an ultracentrifuge tube (Beckman Coulter Ultra-Clear, 14 × 95 mm). To avoid mixing, the sample was overlaid with gentle pipetting of 4 ml cold 20% Histodenz in PBS/sucrose and finally with 4 ml cold 10% Histodenz in PBS/sucrose. An additional 10% Histodenz in PBS/sucrose was used to bring the tube volume to ~1 mm from the rim of the tube. Alternatively, in some experiments, samples were loaded onto an iodixanol (Sigma) gradient. Samples were centrifuged at 35,000 r.p.m. (~209,000g) in a SW40Ti rotor for 16 h. Samples were collected in 1-ml fractions from the top of the tube and mixed with 1 ml 2× SDS–PAGE sample loading buffer. Aliquots (20 μl) were analysed by western blot with anti-FLAG, anti-calnexin or anti-RFP antibodies as described above.
Immunoprecipitation assay
Approximately 1.5 × 106 Huh7.5 cells stably expressing IFI6-3×FLAG were collected for each condition and pelleted. Cells were resuspended and lysed gently at 4 °C with nutation for 15 min in 250 μl of a buffer containing 20 mM Tris-HCl (pH 7.5), 1.5 mM MgCl2, 150 mM NaCl, 1% NP-40, 5% glycerol and 1 protease inhibitor pellet (Roche) per 10 ml of solution. When used, ATP was included at a final concentration of 2 mM (A6559, Sigma). Cells were then pelleted at 16,000g for 8 min at 4 °C. An affinity gel containing 4% agarose beads with FLAG-M2 antibody covalently bound (F2426, Sigma) was equilibrated in washing buffer (20 mM Tris-HCl (pH 7.5), 250 mM NaCl, 0.2% NP-40, 1.5 mM MgCl2 and 5% glycerol). Cleared lysate (5–10%) was collected as inputs and stored at −80 °C. The remaining lysate was added to the beads and incubated overnight at 4 °C on a rotator. The next morning, the supernatant was collected and beads were washed four times with washing buffer. A 3×FLAG peptide (4799, Sigma) was used to elute the bound products off of the column at 4 °C with constant vortexing for 1 h. Input samples and eluted products were run on a SDS–PAGE gel and transferred to a PVDF membrane, which was probed with antibodies against BiP or FLAG (rabbit polyclonal).
Secretion assay
293T cells plated at 400,000 cells per well in 6-well plates were transfected with pTRIP.CMV.IVSB-CCL2-1×FLAG or pTRIP.CMV.IVSB-IFI6-3×FLAG plasmids using X-tremeGENE 9 (Roche). The next day, media were removed and 1 ml Optimem (Gibco) with or without 1 μg ml−1 Brefeldin A was added to cells. After 5 h, supernatants were removed and proteins were precipitated with trichloroacetic acid. Cells were harvested with Accumax and post-nuclear cell lysates were analysed by western blot with anti-FLAG or anti-actin antibodies as described above.
Protein cleavage assays
293T, 293T control sgRNA or 293T SPCS1-KO cells (the latter two kindly provided by M. Diamond) were plated at 400,000 cells per well in 6-well plates. As indicated, cells were transfected with pTRIP.XKB-2K-NS4B-HA, pTRIP.XKB-C-prM-HA, pTRIP.CMV.IVSB-IFI6-3×FLAG, pTRIP.CMV.IVSB-GFP-1×FLAG, pQCXIP-DENV-NS2B-NS3 (wild type or S135A; gifts from M. Gack) and MRX-HA-STING (gift from N. Yan) plasmids using X-tremeGENE 9 (Roche). The next day, cells were harvested with Accumax and post-nuclear cell lysates were analysed by western blot with anti-FLAG, anti-HA or anti-actin antibodies as described above.
Statistics and reproducibility
Statistical analyses were performed with GraphPad Prism software. In all figures, the data points and bar graphs represent the mean of independent biological replicates. In all graphs, the error bars represent the standard deviation and are only shown for experiments with n = 3 or greater. In bar graphs, the individual data points are overlaid. The genome-wide CRISPR screen was performed once at 900× library coverage (Fig. 1b). Western blots are representative of multiple biological replicates showing similar results, as follows: IFN-induced IFI6 expression in Huh7.5 cells, n = 3 (Fig. 1e); IFI6 and actin expression in WNV-infected Huh7.5 cells, n = 2 (Fig. 1g); IFN-induced IFI6 expression in NHDF, n = 4 (Fig. 1n), DENV-induced IFI6 expression in NHDF, n = 6 (Fig. 1o); IFI6 and BiP protein levels after IFN treatment in Huh7.5 cells, n = 3 (Fig. 2a and Supplementary Fig. 1a); the effect of BiP depletion on IFI6 levels, n = 3 (Fig. 2e); the rescue of IFI6 expression by proteasome inhibition, n = 3 (Fig. 2f); the effect of ATPase-defective BiP mutants on IFI6 expression, n = 3 (Fig. 2g); BiP immunoprecipitation, n = 3 (Fig. 2h); endogenous IFI6-3×FLAG expression in gene-tagged U2OS cells, n = 2 (Fig. 2i); the membrane flotation assay, n = 2 (Fig. 3f and Supplementary Fig. 4e); the secretion assay, n = 2 (Fig. 3g); the IFI6 time-of-addition experiment, n = 3 (Fig. 4e); and polyprotein processing, n = 2 (Supplementary Fig. 6a,b) and n = 3 (Supplementary Fig. 6c). Uncropped blots are presented in Supplementary Fig. 9. Microscopy-based data are representative of multiple biological replicates showing similar results, as follows: the proximity ligation assay, n = 3 (Fig. 2j and Supplementary Fig. 4a); immunofluorescence microscopy, n = 3 (Figs. 3a,d and 4f and Supplementary Fig. 8) or n = 2 (Figs. 3b and 4c); and electron microscopy, n = 3 (Fig. 4b). One of the three electron microscopy experiments was blinded before analysis. Microscopy data in Fig. 3c,e represent multiple fields of view from one biological replicate.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data sets generated and/or analysed during the current study are available from the corresponding author upon request.
References
Neufeldt, C. J., Cortese, M., Acosta, E. G. & Bartenschlager, R. Rewiring cellular networks by members of the Flaviviridae family. Nat. Rev. Microbiol. 16, 125–142 (2018).
Kelly, J. M. et al. Characterization of a human gene inducible by alpha- and beta-interferons and its expression in mouse cells. EMBO J. 5, 1601–1606 (1986).
Shresta, S. et al. Interferon-dependent immunity is essential for resistance to primary dengue virus infection in mice, whereas T- and B-cell-dependent immunity are less critical. J. Virol. 78, 2701–2710 (2004).
Schoggins, J. W. et al. A diverse range of gene products are effectors of the type I interferon antiviral response. Nature 472, 481–485 (2011).
Schoggins, J. W. et al. Pan-viral specificity of IFN-induced genes reveals new roles for cGAS in innate immunity. Nature 505, 691–695 (2014).
Li, J. et al. A short hairpin RNA screen of interferon-stimulated genes identifies a novel negative regulator of the cellular antiviral response. mBio 4, e00385-13 (2013).
Schoggins, J. W. et al. Dengue reporter viruses reveal viral dynamics in interferon receptor-deficient mice and sensitivity to interferon effectors in vitro. Proc. Natl Acad. Sci. USA 109, 14610–14615 (2012).
Parker, N. & Porter, A. C. Identification of a novel gene family that includes the interferon-inducible human genes 6-16 and ISG12. BMC Genomics 5, 8 (2004).
Gjermandsen, I. M., Justesen, J. & Martensen, P. M. The interferon-induced gene ISG12 is regulated by various cytokines as the gene 6-16 in human cell lines. Cytokine 12, 233–238 (2000).
Wang, J., Lee, J., Liem, D. & Ping, P. HSPA5 gene encoding Hsp70 chaperone BiP in the endoplasmic reticulum. Gene 618, 14–23 (2017).
Gaut, J. R. & Hendershot, L. M. Mutations within the nucleotide binding site of immunoglobulin-binding protein inhibit ATPase activity and interfere with release of immunoglobulin heavy chain. J. Biol. Chem. 268, 7248–7255 (1993).
Munro, S. & Pelham, H. R. An Hsp70-like protein in the ER: identity with the 78 kd glucose-regulated protein and immunoglobulin heavy chain binding protein. Cell 46, 291–300 (1986).
Lackner, D. H. et al. A generic strategy for CRISPR–Cas9-mediated gene tagging. Nat. Commun. 6, 10237 (2015).
Cheriyath, V. et al. G1P3, an IFN-induced survival factor, antagonizes TRAIL-induced apoptosis in human myeloma cells. J. Clin. Invest. 117, 3107–3117 (2007).
Cheriyath, V. et al. G1P3, an interferon- and estrogen-induced survival protein contributes to hyperplasia, tamoxifen resistance and poor outcomes in breast cancer. Oncogene 31, 2222–2236 (2012).
Tahara, E. et al. G1P3, an interferon inducible gene 6-16, is expressed in gastric cancers and inhibits mitochondrial-mediated apoptosis in gastric cancer cell line TMK-1 cell. Cancer Immunol. Immunother. 54, 729–740 (2005).
Miller, S., Sparacio, S. & Bartenschlager, R. Subcellular localization and membrane topology of the dengue virus type 2 non-structural protein 4B. J. Biol. Chem. 281, 8854–8863 (2006).
Vogt, D. A. & Ott, M. Membrane flotation assay. Bio. Protoc. 5, e1435 (2015).
Fujiki, Y., Hubbard, A. L., Fowler, S. & Lazarow, P. B. Isolation of intracellular membranes by means of sodium carbonate treatment: application to endoplasmic reticulum. J. Cell Biol. 93, 97–102 (1982).
Jones, C. T., Patkar, C. G. & Kuhn, R. J. Construction and applications of yellow fever virus replicons. Virology 331, 247–259 (2005).
Aguirre, S. et al. DENV inhibits type I IFN production in infected cells by cleaving human STING. PLoS Pathog. 8, e1002934 (2012).
Zhang, R. et al. A CRISPR screen defines a signal peptide processing pathway required by flaviviruses. Nature 535, 164–168 (2016).
Gillespie, L. K., Hoenen, A., Morgan, G. & Mackenzie, J. M. The endoplasmic reticulum provides the membrane platform for biogenesis of the flavivirus replication complex. J. Virol. 84, 10438–10447 (2010).
Roosendaal, J., Westaway, E. G., Khromykh, A. & Mackenzie, J. M. Regulated cleavages at the West Nile virus NS4A-2K-NS4B junctions play a major role in rearranging cytoplasmic membranes and Golgi trafficking of the NS4A protein. J. Virol. 80, 4623–4632 (2006).
Miller, S., Kastner, S., Krijnse-Locker, J., Buhler, S. & Bartenschlager, R. The non-structural protein 4A of dengue virus is an integral membrane protein inducing membrane alterations in a 2K-regulated manner. J. Biol. Chem. 282, 8873–8882 (2007).
Qi, H. et al. Systematic identification of anti-interferon function on hepatitis C virus genome reveals p7 as an immune evasion protein. Proc. Natl Acad. Sci. USA 114, 2018–2023 (2017).
Itsui, Y. et al. Expressional screening of interferon-stimulated genes for antiviral activity against hepatitis C virus replication. J. Viral Hepat. 13, 690–700 (2006).
Meyer, K. et al. Interferon-α inducible protein 6 impairs EGFR activation by CD81 and inhibits hepatitis C virus infection. Sci. Rep. 5, 9012 (2015).
Metz, P. et al. Identification of type I and type II interferon-induced effectors controlling hepatitis C virus replication. Hepatology 56, 2082–2093 (2012).
Fusco, D. N. et al. A genetic screen identifies interferon-α effector genes required to suppress hepatitis C virus replication. Gastroenterology 144, 1438–1449 (2013).
Zhao, H. et al. A functional genomic screen reveals novel host genes that mediate interferon-alpha’s effects against hepatitis C virus. J. Hepatol. 56, 326–333 (2012).
Knoops, K. et al. SARS-coronavirus replication is supported by a reticulovesicular network of modified endoplasmic reticulum. PLoS Biol. 6, e226 (2008).
Hanners, N. W. et al. Western Zika virus in human fetal neural progenitors persists long term with partial cytopathic and limited immunogenic effects. Cell Rep. 15, 2315–2322 (2016).
Marukian, S. et al. Cell culture-produced hepatitis C virus does not infect peripheral blood mononuclear cells. Hepatology 48, 1843–1850 (2008).
Schwarz, M. C. et al. Rescue of the 1947 Zika virus prototype strain with a Cytomegalovirus promoter-driven cDNA clone. mSphere 1, e00246-16 (2016).
Grigorov, B., Rabilloud, J., Lawrence, P. & Gerlier, D. Rapid titration of measles and other viruses: optimization with determination of replication cycle length. PLoS ONE 6, e24135 (2011).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Shalem, O. et al. Genome-scale CRISPR–Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Perelman, S. S. et al. Cell-based screen identifies human interferon-stimulated regulators of Listeria monocytogenes infection. PLoS Pathog. 12, e1006102 (2016).
Acknowledgements
We thank N. Alto and J. Pfeiffer for critical manuscript feedback, M. Diamond and M. Henne for helpful discussions and C. Rice for reagents. We acknowledge the technical support of the UT Southwestern Live Cell Imaging Core and Electron Microscopy Core. This work was supported in part by NIH grants AI117922 and DK095031 (J.W.S.), the UT Southwestern Endowed Scholars program (J.W.S.), the Rita Allen Foundation (J.W.S.), the Clayton Foundation for Research (J.W.S.) and the National Science Foundation Graduate Research Fellowship Program grant 2016217834 (I.N.B.). C.X. was partially supported by NIH grant UL1TR001105. Additional support for R.B.R. and K.B.M. was obtained from NIH T32 Training grants AI007520 and AI005284, respectively. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH or the other funders.
Author information
Authors and Affiliations
Contributions
R.B.R., M.B.O. and J.W.S. designed the project. R.B.R., M.B.O., J.L.E., M.B.M., I.N.B., K.B.M., P.C.D.-R. and J.W.S. performed the experimental work. C.D. and G.K. provided the human fetal neural progenitor cultures. All authors contributed to data analysis and interpretation. A.K. and C.X. processed the CRISPR data sets. R.B.R., M.B.O. and J.W.S. drafted the manuscript text. All authors contributed to the editing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–9, Supplementary Tables 1 and 2, Supplementary Methods and Supplementary References.
Supplementary Table 1
MAGeCK analysis output of CRISPR screen.
Rights and permissions
About this article
Cite this article
Richardson, R.B., Ohlson, M.B., Eitson, J.L. et al. A CRISPR screen identifies IFI6 as an ER-resident interferon effector that blocks flavivirus replication. Nat Microbiol 3, 1214–1223 (2018). https://doi.org/10.1038/s41564-018-0244-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-018-0244-1
This article is cited by
-
CRISPR–Cas system to discover host-virus interactions in Flaviviridae
Virology Journal (2023)
-
LY6E is a pan-coronavirus restriction factor in the respiratory tract
Nature Microbiology (2023)
-
Zika virus targets human trophoblast stem cells and prevents syncytialization in placental trophoblast organoids
Nature Communications (2023)
-
Molecular factors shaping whether HCMV infection is productive or latent
Nature Microbiology (2023)
-
Downregulation of Microcephaly-Causing Genes as a Mechanism for ZIKV Teratogenesis: A Meta-analysis of RNA-Seq Studies
Journal of Molecular Neuroscience (2023)