Abstract
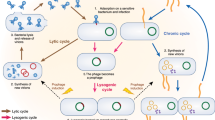
The dysregulation of intestinal microbial communities is associated with inflammatory bowel diseases (IBD). Studies aimed at understanding the contribution of the microbiota to inflammatory diseases have primarily focused on bacteria, yet the intestine harbours a viral component dominated by prokaryotic viruses known as bacteriophages (phages). Phage numbers are elevated at the intestinal mucosal surface and phages increase in abundance during IBD, suggesting that phages play an unidentified role in IBD. We used a sequence-independent approach for the selection of viral contigs and then applied quantitative metagenomics to study intestinal phages in a mouse model of colitis. We discovered that during colitis the intestinal phage population is altered and transitions from an ordered state to a stochastic dysbiosis. We identified phages specific to pathobiotic hosts associated with intestinal disease, whose abundances are altered during colitis. Additionally, phage populations in healthy and diseased mice overlapped with phages from healthy humans and humans with IBD. Our findings indicate that intestinal phage communities are altered during inflammatory disease, establishing a platform for investigating phage involvement in IBD.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Abraham, C. & Cho, J. H. Inflammatory bowel disease. N. Engl. J. Med. 361, 2066–2078 (2009).
Molodecky, N. A. et al. Increasing incidence and prevalence of the inflammatory bowel diseases with time, based on systematic review. Gastroenterology 142, 46–54 (2012).
Hooper, L. V., Midtvedt, T. & Gordon, J. I. How host–microbial interactions shape the nutrient environment of the mammalian intestine. Annu. Rev. Nutr. 22, 283–307 (2002).
Barthel, M. et al. Pretreatment of mice with streptomycin provides a Salmonella enterica serovar Typhimurium colitis model that allows analysis of both pathogen and host. Infect. Immun. 71, 2839–2858 (2003).
Ivanov, I. I. et al. Specific microbiota direct the differentiation of IL-17-producing T-helper cells in the mucosa of the small intestine. Cell Host Microbe 4, 337–349 (2008).
Mazmanian, S. K., Liu, C. H., Tzianabos, A. O. & Kasper, D. L. An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell 122, 107–118 (2005).
Reeves, A. E., Koenigsknecht, M. J., Bergin, I. L. & Young, V. B. Suppression of Clostridium difficile in the gastrointestinal tracts of germfree mice inoculated with a murine isolate from the family Lachnospiraceae. Infect. Immun. 80, 3786–3794 (2012).
Sheehan, D., Moran, C. & Shanahan, F. The microbiota in inflammatory bowel disease. J. Gastroenterol. 50, 495–507 (2015).
Belkaid, Y. & Hand, T. W. Role of the microbiota in immunity and inflammation. Cell 157, 121–141 (2014).
Minot, S. et al. The human gut virome: Inter-individual variation and dynamic response to diet. Genome Res. 21, 1616–1625 (2011).
Reyes, A. et al. Viruses in the faecal microbiota of monozygotic twins and their mothers. Nature 466, 334–338 (2010).
Dinsdale, E. A. et al. Functional metagenomic profiling of nine biomes. Nature 452, 629–632 (2008).
Oh, J. et al. Temporal stability of the human skin microbiome. Cell 165, 854–866 (2016).
Feiner, R. et al. A new perspective on lysogeny: Prophages as active regulatory switches of bacteria. Nat. Rev. Microbiol. 13, 641–650 (2015).
Winter, S. E., Lopez, C. A. & Baumler, A. J. The dynamics of gut-associated microbial communities during inflammation. EMBO Rep. 14, 319–327 (2013).
Stecher, B. The roles of inflammation, nutrient availability and the commensal microbiota in enteric pathogen infection. Microbiol. Spectr. 3, MBP-0008-2014 (2015).
Banks, D. J., Lei, B. & Musser, J. M. Prophage induction and expression of prophage-encoded virulence factors in group A Streptococcus serotype M3 strain MGAS315. Infect. Immun. 71, 7079–7086 (2003).
Duerkop, B. A., Clements, C. V., Rollins, D., Rodrigues, J. L. & Hooper, L. V. A composite bacteriophage alters colonization by an intestinal commensal bacterium. Proc. Natl Acad. Sci. USA 109, 17621–17626 (2012).
Diard, M. et al. Inflammation boosts bacteriophage transfer between Salmonella spp. Science 355, 1211–1215 (2017).
Wang, W. et al. Metagenomic analysis of microbiome in colon tissue from subjects with inflammatory bowel diseases reveals interplay of viruses and bacteria. Inflamm. Bowel Dis. 21, 1419–1427 (2015).
Norman, J. M. et al. Disease-specific alterations in the enteric virome in inflammatory bowel disease. Cell 160, 447–460 (2015).
Lepage, P. et al. Dysbiosis in inflammatory bowel disease: A role for bacteriophages? Gut 57, 424–425 (2008).
Barr, J. J. et al. Bacteriophage adhering to mucus provide a non-host-derived immunity. Proc. Natl Acad. Sci. USA 110, 10771–10776 (2013).
Ostanin, D. V. et al. T cell transfer model of chronic colitis: Concepts, considerations, and tricks of the trade. Am. J. Physiol. Gastrointest. Liver Physiol. 296, G135–146 (2009).
Powrie, F., Correa-Oliveira, R., Mauze, S. & Coffman, R. L. Regulatory interactions between CD45RBhigh and CD45RBlow CD4+ T cells are important for the balance between protective and pathogenic cell-mediated immunity. J. Exp. Med. 179, 589–600 (1994).
Kang, D. W. et al. Microbiota Transfer Therapy alters gut ecosystem and improves gastrointestinal and autism symptoms: An open-label study. Microbiome 5, 10 (2017).
Berry, D. et al. Phylotype-level 16S rRNA analysis reveals new bacterial indicators of health state in acute murine colitis. ISME J. 6, 2091–2106 (2012).
Hughes, E. R. et al. Microbial respiration and formate oxidation as metabolic signatures of inflammation-associated dysbiosis. Cell Host Microbe 21, 208–219 (2017).
Lupp, C. et al. Host-mediated inflammation disrupts the intestinal microbiota and promotes the overgrowth of Enterobacteriaceae. Cell Host Microbe 2, 204 (2007).
Roux, S., Emerson, J. B., Eloe-Fadrosh, E. A. & Sullivan, M. B. Benchmarking viromics: An in silico evaluation of metagenome-enabled estimates of viral community composition and diversity. PeerJ 5, e3817 (2017).
Paez-Espino, D., Pavlopoulos, G. A., Ivanova, N. N. & Kyrpides, N. C. Nontargeted virus sequence discovery pipeline and virus clustering for metagenomic data. Nat. Protoc. 12, 1673–1682 (2017).
Roux, S., Enault, F., Hurwitz, B. L. & Sullivan, M. B. VirSorter: Mining viral signal from microbial genomic data. PeerJ 3, e985 (2015).
Paez-Espino, D. et al. IMG/VR: A database of cultured and uncultured DNA viruses and retroviruses. Nucleic Acids Res. 45, D457–D465 (2017).
Bailly-Bechet, M., Vergassola, M. & Rocha, E. Causes for the intriguing presence of tRNAs in phages. Genome Res. 17, 1486–1495 (2007).
Kristensen, D. M. et al. Orthologous gene clusters and taxon signature genes for viruses of prokaryotes. J. Bacteriol. 195, 941–950 (2013).
Paez-Espino, D. et al. Uncovering Earth’s virome. Nature 536, 425–430 (2016).
Uchiyama, J. et al. In silico analysis of AHJD-like viruses, Staphylococcus aureus phages S24-1 and S13’, and study of phage S24-1 adsorption. Microbiologyopen 3, 257–270 (2014).
Prangishvili, D., Forterre, P. & Garrett, R. A. Viruses of the Archaea: A unifying view. Nat. Rev. Microbiol. 4, 837–848 (2006).
Manrique, P. et al. Healthy human gut phageome. Proc. Natl Acad. Sci. USA 113, 10400–10405 (2016).
Govind, R., Fralick, J. A. & Rolfe, R. D. Genomic organization and molecular characterization of Clostridium difficile bacteriophage PhiCD119. J. Bacteriol. 188, 2568–2577 (2006).
Mesyanzhinov, V. V. et al. The genome of bacteriophage phiKZ of Pseudomonas aeruginosa. J. Mol. Biol. 317, 1–19 (2002).
Esposito, D. et al. The complete nucleotide sequence of bacteriophage HP1 DNA. Nucleic Acids Res. 24, 2360–2368 (1996).
Ferrenberg, S. et al. Changes in assembly processes in soil bacterial communities following a wildfire disturbance. ISME J. 7, 1102–1111 (2013).
Lee, S. H., Sorensen, J. W., Grady, K. L., Tobin, T. C. & Shade, A. Divergent extremes but convergent recovery of bacterial and archaeal soil communities to an ongoing subterranean coal mine fire. ISME J. 11, 1447–1459 (2017).
Winter, S. E. et al. Host-derived nitrate boosts growth of E. coli in the inflamed gut. Science 339, 708–711 (2013).
Turnbaugh, P. J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006).
Jiang, W. et al. Dysbiosis gut microbiota associated with inflammation and impaired mucosal immune function in intestine of humans with non-alcoholic fatty liver disease. Sci. Rep. 5, 8096 (2015).
Duplessis, M. & Moineau, S. Identification of a genetic determinant responsible for host specificity in Streptococcus thermophilus bacteriophages. Mol. Microbiol. 41, 325–336 (2001).
Holmfeldt, K., Middelboe, M., Nybroe, O. & Riemann, L. Large variabilities in host strain susceptibility and phage host range govern interactions between lytic marine phages and their Flavobacterium hosts. Appl. Environ. Microbiol. 73, 6730–6739 (2007).
Duerkop, B. A., Huo, W., Bhardwaj, P., Palmer, K. L. & Hooper, L. V. Molecular basis for lytic bacteriophage resistance in Enterococci. mBio 7, e01304–01316 (2016).
Mombaerts, P. et al. RAG-1-deficient mice have no mature B and T lymphocytes. Cell 68, 869–877 (1992).
Cash, H. L., Whitham, C. V., Behrendt, C. L. & Hooper, L. V. Symbiotic bacteria direct expression of an intestinal bactericidal lectin. Science 313, 1126–1130 (2006).
Kleiner, M., Hooper, L. V. & Duerkop, B. A. Evaluation of methods to purify virus-like particles for metagenomic sequencing of intestinal viromes. BMC Genom. 16, 7 (2015).
Roux, S., Krupovic, M., Debroas, D., Forterre, P. & Enault, F. Assessment of viral community functional potential from viral metagenomes may be hampered by contamination with cellular sequences. Open Biol. 3, 130160 (2013).
Enault, F. et al. Phages rarely encode antibiotic resistance genes: A cautionary tale for virome analyses. ISME J. 11, 237–247 (2017).
BBMap: short read aligner and other bioinformatic tools v36.99 (Bushnell, B., 2018); https://sourceforge.net/projects/bbmap/
phyloFlash v.2.0 (Gruber-Vodicka, H., Pruesse, E.A. & Seah, B., 2018); https://github.com/HRGV/phyloFlash
Quast, C. et al. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 41, D590–596 (2013).
Peng, Y., Leung, H. C., Yiu, S. M. & Chin, F. Y. IDBA-UD: A de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinformatics 28, 1420–1428 (2012).
Li, W. & Godzik, A. Cd-hit: A fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659 (2006).
Finn, R. D., Clements, J. & Eddy, S. R. HMMER web server: Interactive sequence similarity searching. Nucleic Acids Res. 39, W29–37 (2011).
Camacho, C. et al. BLAST+: Architecture and applications. BMC Bioinformatics 10, 421 (2009).
Strous, M., Kraft, B., Bisdorf, R. & Tegetmeyer, H. E. The binning of metagenomic contigs for microbial physiology of mixed cultures. Front. Microbiol. 3, 410 (2012).
Rho, M., Wu, Y. W., Tang, H., Doak, T. G. & Ye, Y. Diverse CRISPRs evolving in human microbiomes. PLoS Genet. 8, e1002441 (2012).
Laslett, D. & Canback, B. ARAGORN, a program to detect tRNA genes and tmRNA genes in nucleotide sequences. Nucleic Acids Res. 32, 11–16 (2004).
Chen, I. A. et al. IMG/M: Integrated genome and metagenome comparative data analysis system. Nucleic Acids Res. 45, D507–D516 (2017).
Edwards, R. A., McNair, K., Faust, K., Raes, J. & Dutilh, B. E. Computational approaches to predict bacteriophage–host relationships. FEMS Microbiol. Rev. 40, 258–272 (2016).
Acknowledgements
We would like to thank C. Boyd, T. Leal and K. Ruhn for assistance with animals and X. Dong and F. Santoriello for bioinformatics assistance. This work was supported by NIH R01DK070855 (L.V.H.), the Howard Hughes Medical Institute (L.V.H.), NIH K01DK102436 (B.A.D.), start-up funds from the University of Colorado School of Medicine (B.A.D.), the Government of Canada’s Banting Postdoctoral Fellowship (M.K.) and the NC State Chancellor’s Faculty Excellence Program Cluster on Microbiomes and Complex Microbial Communities (M.K.). This work was partly conducted by the US Department of Energy Joint Genome Institute, a DOE Office of Science User Facility, under contract number DE-AC02-05CH11231.
Author information
Authors and Affiliations
Contributions
B.A.D., M.K. and L.V.H. designed the study. B.A.D., M.K., D.P.E. and B.H. performed experiments. B.A.D., M.K., D.P.E. and B.B. performed bioinformatic analyses. B.A.D., M.K., D.P.E., W.Z., S.E.W., N.C.K. and L.V.H. analysed data. B.A.D., M.K. and L.V.H. wrote the paper with input from all of the authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–10
Supplementary Table 1
VLP contigs containing virus-like genes determined using a VPF database
Supplementary Table 2
VLP contigs determined to be phages using VirSorter
Supplementary Table 3
VLP contigs grouped into genetically related viral clusters
Supplementary Table 4
VLP reads mapped to phage genomes from NCBI
Supplementary Table 5
VLP read mapping abundances against the IMG/VR database
Supplementary Table 6
VLP reads mapped to the curated VLP contig database
Supplementary Table 7
VLP reads mapped to curated VLP contig database at day 42
Supplementary Table 8
Contigs with high read recruitment in T-cell-treated animals
Supplementary Table 9
Phage taxonomy or host assignment per contig
Supplementary Table 10
P value
Rights and permissions
About this article
Cite this article
Duerkop, B.A., Kleiner, M., Paez-Espino, D. et al. Murine colitis reveals a disease-associated bacteriophage community. Nat Microbiol 3, 1023–1031 (2018). https://doi.org/10.1038/s41564-018-0210-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-018-0210-y
This article is cited by
-
MetaBinner: a high-performance and stand-alone ensemble binning method to recover individual genomes from complex microbial communities
Genome Biology (2023)
-
Respiratory eukaryotic virome expansion and bacteriophage deficiency characterize childhood asthma
Scientific Reports (2023)
-
Characterization of Phietavirus Henu 2 in the virome of individuals with acute gastroenteritis
Virus Genes (2023)
-
Transplantation of bacteriophages from ulcerative colitis patients shifts the gut bacteriome and exacerbates the severity of DSS colitis
Microbiome (2022)
-
Longitudinal gut virome analysis identifies specific viral signatures that precede necrotizing enterocolitis onset in preterm infants
Nature Microbiology (2022)