Abstract
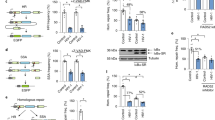
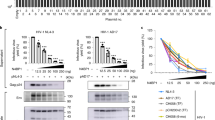
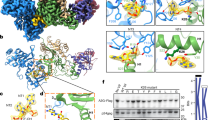
To evade host immune defences, human immunodeficiency viruses 1 and 2 (HIV-1 and HIV-2) have evolved auxiliary proteins that target cell restriction factors. Viral protein X (Vpx) from the HIV-2/SIVsmm lineage enhances viral infection by antagonizing SAMHD1 (refs 1,2), but this antagonism is not sufficient to explain all Vpx phenotypes. Here, through a proteomic screen, we identified another Vpx target—HUSH (TASOR, MPP8 and periphilin)—a complex involved in position-effect variegation3. HUSH downregulation by Vpx is observed in primary cells and HIV-2-infected cells. Vpx binds HUSH and induces its proteasomal degradation through the recruitment of the DCAF1 ubiquitin ligase adaptor, independently from SAMHD1 antagonism. As a consequence, Vpx is able to reactivate HIV latent proviruses, unlike Vpx mutants, which are unable to induce HUSH degradation. Although antagonism of human HUSH is not conserved among all lentiviral lineages including HIV-1, it is a feature of viral protein R (Vpr) from simian immunodeficiency viruses (SIVs) of African green monkeys and from the divergent SIV of l’Hoest's monkey, arguing in favour of an ancient lentiviral species-specific vpx/vpr gene function. Altogether, our results suggest the HUSH complex as a restriction factor, active in primary CD4+ T cells and counteracted by Vpx, therefore providing a molecular link between intrinsic immunity and epigenetic control.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hrecka, K. et al. Vpx relieves inhibition of HIV-1 infection of macrophages mediated by the SAMHD1 protein. Nature 474, 658–661 (2011).
Laguette, N. et al. SAMHD1 is the dendritic- and myeloid-cell-specific HIV-1 restriction factor counteracted by Vpx. Nature 474, 654–657 (2011).
Tchasovnikarova, I. A. et al. Gene silencing. Epigenetic silencing by the HUSH complex mediates position-effect variegation in human cells. Science 348, 1481–1485 (2015).
Matheron, S. et al. Factors associated with clinical progression in HIV-2 infected-patients: the French ANRS cohort. AIDS 17, 2593–2601 (2003).
Saleh, S., Vranckx, L., Gijsbers, R., Christ, F. & Debyser, Z. Insight into HIV-2 latency may disclose strategies for a cure for HIV-1 infection. J. Virus Erad. 3, 7–14 (2017).
Yu, X. F., Yu, Q. C., Essex, M. & Lee, T. H. The vpx gene of simian immunodeficiency virus facilitates efficient viral replication in fresh lymphocytes and macrophage. J. Virol. 65, 5088–5091 (1991).
Guyader, M., Emerman, M., Montagnier, L. & Peden, K. VPX mutants of HIV-2 are infectious in established cell lines but display a severe defect in peripheral blood lymphocytes. EMBO J. 8, 1169–1175 (1989).
Schaller, T., Bauby, H., Hue, S., Malim, M. H. & Goujon, C. New insights into an X-traordinary viral protein. Front. Microbiol. 5, 126 (2014).
Baldauf, H. M. et al. Vpx overcomes a SAMHD1-independent block to HIV reverse transcription that is specific to resting CD4 T cells. Proc. Natl Acad. Sci. USA 114, 2729–2734 (2017).
Fujita, M., Nomaguchi, M., Adachi, A. & Otsuka, M. SAMHD1-dependent and -independent functions of HIV-2/SIV Vpx protein. Front. Microbiol. 3, 297 (2012).
Lahouassa, H. et al. HIV-1 Vpr degrades the HLTF DNA translocase in T cells and macrophages. Proc. Natl Acad. Sci. USA 113, 5311–5316 (2016).
Liu, L. et al. A whole genome screen for HIV restriction factors. Retrovirology 8, 94 (2011).
Liu, N.et al. Selective silencing of euchromatic L1s revealed by genome-wide screens for L1 regulators. Nature 553, 228–232 2018).
Le Rouzic, E. et al. HIV1 Vpr arrests the cell cycle by recruiting DCAF1/VprBP, a receptor of the Cul4-DDB1 ubiquitin ligase. Cell Cycle 6, 182–188 (2007).
Jordan, A., Bisgrove, D. & Verdin, E. HIV reproducibly establishes a latent infection after acute infection of T cells in vitro. EMBO J. 22, 1868–1877 (2003).
Romani, B. et al. HIV-1 Vpr reactivates latent HIV-1 provirus by inducing depletion of class I HDACs on chromatin. Sci. Rep. 6, 31924 (2016).
Lim, E. S. et al. The ability of primate lentiviruses to degrade the monocyte restriction factor SAMHD1 preceded the birth of the viral accessory protein Vpx. Cell Host Microbe. 11, 194–204 (2012).
Daugherty, M. D. & Malik, H. S. Rules of engagement: molecular insights from host–virus arms races. Annu. Rev. Genet. 46, 677–700 (2012).
Duggal, N. K. & Emerman, M. Evolutionary conflicts between viruses and restriction factors shape immunity. Nat. Rev. Immunol. 12, 687–695 (2012).
Chen, H. C., Martinez, J. P., Zorita, E., Meyerhans, A. & Filion, G. J. Position effects influence HIV latency reversal. Nat. Struct. Mol. Biol. 24, 47–54 (2017).
Gramberg, T., Sunseri, N. & Landau, N. R. Evidence for an activation domain at the amino terminus of simian immunodeficiency virus Vpx. J. Virol. 84, 1387–1396 (2010).
Goujon, C. et al. With a little help from a friend: increasing HIV transduction of monocyte-derived dendritic cells with virion-like particles of SIV(MAC). Gene Ther. 13, 991–994 (2006).
Ueno, F. et al. Vpx and Vpr proteins of HIV-2 up-regulate the viral infectivity by a distinct mechanism in lymphocytic cells. Microbes Infect. 5, 387–395 (2003).
Bergamaschi, A. et al. The human immunodeficiency virus type 2 Vpx protein usurps the CUL4A-DDB1 DCAF1 ubiquitin ligase to overcome a postentry block in macrophage infection. J. Virol. 83, 4854–4860 (2009).
Manel, N. et al. A cryptic sensor for HIV-1 activates antiviral innate immunity in dendritic cells. Nature 467, 214–217 (2010).
Gerard, A. et al. The integrase cofactor LEDGF/p75 associates with Iws1 and Spt6 for postintegration silencing of HIV-1 gene expression in latently infected cells. Cell Host Microbe 17, 107–117 (2015).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Huh, J. W. et al. Molecular evolution of the periphilin gene in relation to human endogenous retrovirus m element. J. Mol. Evol. 62, 730–737 (2006).
Loytynoja, A. & Goldman, N. Phylogeny-aware gap placement prevents errors in sequence alignment and evolutionary analysis. Science 320, 1632–1635 (2008).
Kosakovsky Pond, S. L., Posada, D., Gravenor, M. B., Woelk, C. H. & Frost, S. D. Automated phylogenetic detection of recombination using a genetic algorithm. Mol. Biol. Evol. 23, 1891–1901 (2006).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010).
Pond, S. L., Frost, S. D. & Muse, S. V. HyPhy: hypothesis testing using phylogenies. Bioinformatics 21, 676–679 (2005).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Gueguen, L. et al. Bio++: efficient extensible libraries and tools for computational molecular evolution. Mol. Biol. Evol. 30, 1745–1750 (2013).
Murrell, B. et al. Gene-wide identification of episodic selection. Mol. Biol. Evol. 32, 1365–1371 (2015).
Guegen, L. & Duret, L. Unbiased estimate of synonymous and non-synonymous substitution rates with non-stationary base composition. Mol. Biol. Evol. 35, 734–742 (2017).
Murrell, B. et al. FUBAR: a fast, unconstrained bayesian approximation for inferring selection. Mol. Biol. Evol. 30, 1196–1205 (2013).
Bell, S. M. & Bedford, T. Modern-day SIV viral diversity generated by extensive recombination and cross-species transmission. PLoS Pathog. 13, e1006466 (2017).
Gifford, R. J. et al. A transitional endogenous lentivirus from the genome of a basal primate and implications for lentivirus evolution. Proc. Natl Acad. Sci. USA 105, 20362–20367 (2008).
Acknowledgements
The authors thank C. Pique and her group and M. Mangeney for discussions, S. Emiliani and A. Abdouni for their advice and protocols for conducting the ChIP experiment, L. Guéguen and L. Picard for the Bio++ analyses and their input on evolutionary analyses, S. Jacquet for her comments on the manuscript, and A. Cimarelli, head of the ‘Host–Pathogen Interaction during Lentiviral Infection’ team at CIRI Lyon. The authors also thank M. Mangeney, G. Lê-Bury and F. Niedergang for their frequent gifts of primary cells, E. Verdin for J-Lat clones, A. Adachi and N. Manel for HIV-2 constructs, N. Landau for SIV packaging constructs and A. Cimarelli for HIV-1- and SIV-based transfer vectors, M.-L. Blondot for her contribution to the SILAC screen and E. Le Rouzic for raising some Vpx mutants. The authors acknowledge the Cytometry and Immunobiology Facility of the Cochin Institute and the 3P5 proteomic facility of Paris Descartes University. The authors also thank all the contributors of publically available genome sequences. This work was supported by grants from the Agence Nationale de la Recherche sur le SIDA et les hépatites virales (ANRS), SIDACTION, Fondation de France and Fondation pour la Recherche Médicale (FRM, grant no. DEQ20140329528 to F.M.-G.). G.C. acknowedges a fellowship from the French government, S.M.-M. received a fellowship from the Fondation pour la Recherche Médicale (FRM, grant no. DEQ20140329528 to F.M.-G.) and H.L. from SIDACTION. L.E. is supported by the CNRS and by grants from the amfAR (Mathilde Krim Phase II Fellowship no. 109140-58-RKHF), the Fondation pour la Recherche Médicale (FRM ‘Projet Innovant’ no. ING20160435028), the FINOVI (‘Recently Settled Scientist’ grant), the ANRS (no. ECTZ19143) and the ANR LABEX ECOFECT (ANR-11-LABX-0048 of Université de Lyon, within the programme ‘Investissements d’Avenir’ (ANR-11-IDEX-0007) operated by the French National Research Agency).
Author information
Authors and Affiliations
Contributions
G.C., S.M.-M. and F.M.-G. conceived the study. G.C., S.M.-M., R.M., M.M.M., M.M. and F.M.-G. designed experiments and interpreted data. G.C., S.M.-M., R.M., M.M.M. and M.M. performed experiments. H.L. contributed to the SILAC screen design and performance. M.L. contributed to the proteomic analysis of SILAC samples. B.C.R. contributed to the design and performance of experiments in primary cells. L.E. designed and performed the phylogenetic and evolutionary analyses, and contributed to the design of the evolutionary-guided functional experiments. G.C. and F.M.-G. wrote the paper. All authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–6, Supplementary Tables 2–4, blots for Supplementary Figures 1–4.
Supplementary Table 1
TASOR (FAM208A) is the second best downregulated protein in presence of Vpx.
Rights and permissions
About this article
Cite this article
Chougui, G., Munir-Matloob, S., Matkovic, R. et al. HIV-2/SIV viral protein X counteracts HUSH repressor complex. Nat Microbiol 3, 891–897 (2018). https://doi.org/10.1038/s41564-018-0179-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-018-0179-6
This article is cited by
-
The HIV-1 gag p6: a promising target for therapeutic intervention
Retrovirology (2024)
-
Keep quiet: the HUSH complex in transcriptional silencing and disease
Nature Structural & Molecular Biology (2024)
-
HIV-2/SIV Vpx antagonises NF-κB activation by targeting p65
Retrovirology (2022)
-
The KT Jeang Retrovirology prize 2022: Florence Margottin-Goguet
Retrovirology (2022)
-
HUSH-mediated HIV silencing is independent of TASOR phosphorylation on threonine 819
Retrovirology (2022)