Abstract
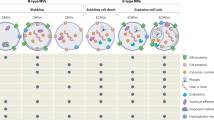
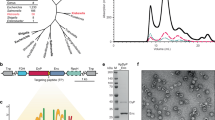
Interest in protein disulfide bond formation has recently increased because of the prominent role of disulfide bonds in bacterial virulence and survival. The first discovered pathway that introduces disulfide bonds into cell envelope proteins consists of Escherichia coli enzymes DsbA and DsbB. Since its discovery, variations on the DsbAB pathway have been found in bacteria and archaea, probably reflecting specific requirements for survival in their ecological niches. One variation found amongst Actinobacteria and Cyanobacteria is the replacement of DsbB by a homologue of human vitamin K epoxide reductase. Many Gram-positive bacteria express enzymes involved in disulfide bond formation that are similar, but non-homologous, to DsbAB. While bacterial pathways promote disulfide bond formation in the bacterial cell envelope, some archaeal extremophiles express proteins with disulfide bonds both in the cytoplasm and in the extra-cytoplasmic space, possibly to stabilize proteins in the face of extreme conditions, such as growth at high temperatures. Here, we summarize the diversity of disulfide-bond-catalysing systems across prokaryotic lineages, discuss examples for understanding the biological basis of such systems, and present perspectives on how such systems are enabling advances in biomedical engineering and drug development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bardwell, J. C. A., McGovern, K. & Beckwith, J. Identification of a protein required for disulfide bond formation in vivo. Cell 67, 581–589 (1991).
LaMantia, M. & Lennarz, W. J. The essential function of yeast protein disulfide isomerase does not reside in its isomerase activity. Cell 74, 899–908 (1993).
Frand, A. R. & Kaiser, C. A. The ERO1 gene of yeast is required for oxidation of protein dithiols in the endoplasmic reticulum. Mol. Cell 1, 161–170 (1998).
Peek, J. A. & Taylor, R. K. Characterization of a periplasmic thiol:disulfide interchange protein required for the functional maturation of secreted virulence factors of Vibrio cholerae. Proc. Natl Acad. Sci. USA 89, 6210–6214 (1992).
Kadokura, H. & Beckwith, J. Detecting folding intermediates of a protein as it passes through the bacterial translocation channel. Cell 138, 1164–1173 (2009).
Ren, B. et al. A protein disulfide oxidoreductase from the archaeon Pyrococcus furiosus contains two thioredoxin fold units. Nat. Struct. Biol. 5, 602–611 (1998).
Mallick, P., Boutz, D. R., Eisenberg, D. & Yeates, T. O. Genomic evidence that the intracellular proteins of archaeal microbes contain disulfide bonds. Proc. Natl Acad. Sci. USA 99, 9679–9684 (2002).
Kadokura, H., Tian, H., Zander, T., Bardwell, J. C. A. & Beckwith, J. Snapshots of DsbA in action: detection of proteins in the process of oxidative folding. Science 303, 534–537 (2004).
Hiniker, A. & Bardwell, J. C. A. In vivo substrate specificity of periplasmic disulfide oxidoreductases. J. Biol. Chem. 279, 12967–12973 (2004).
Yu, J. & Kroll, J. S. DsbA: a protein-folding catalyst contributing to bacterial virulence. Microbes Infect. 1, 1221–1228 (1999).
Lasica, A. M. & Jagusztyn-Krynicka, E. K. The role of Dsb proteins of Gram-negative bacteria in the process of pathogenesis. FEMS Microbiol. Rev. 31, 626–636 (2007).
Heras, B. et al. DSB proteins and bacterial pathogenicity. Nat. Rev. Microbiol. 7, 215–225 (2009).
Dutton, R. J., Boyd, D., Berkmen, M. & Beckwith, J. Bacterial species exhibit diversity in their mechanisms and capacity for protein disulfide bond formation. Proc. Natl Acad. Sci. USA 105, 11933–11938 (2008).
Sato, Y. & Inaba, K. Disulfide bond formation network in the three biological kingdoms, bacteria, fungi and mammals. FEBS J. 279, 2262–2271 (2012).
Bardwell, J. C. et al. A pathway for disulfide bond formation in vivo. Proc. Natl Acad. Sci. USA 90, 1038–1042 (1993).
Sevier, C. S. et al. The prokaryotic enzyme DsbB may share key structural features with eukaryotic disulfide bond forming oxidoreductases. Protein Sci. 14, 1630–1642 (2005).
Bevans, C. G., Krettler, C., Reinhart, C., Watzka, M. & Oldenburg, J. Phylogeny of the vitamin K 2,3-epoxide reductase (VKOR) family and evolutionary relationship to the disulfide bond formation protein B (DsbB) family. Nutrients 7, 6224–6249 (2015).
Inaba, K. et al. Dynamic nature of disulphide bond formation catalysts revealed by crystal structures of DsbB. EMBO J. 28, 779–791 (2009).
Li, W. et al. Structure of a bacterial homologue of vitamin K epoxide reductase. Nature 463, 507–512 (2010).
Bader, M., Muse, W., Ballou, D. P., Gassner, C. & Bardwell, J. C. A. Oxidative protein folding is driven by the electron transport system. Cell 98, 217–227 (1999).
Reedstrom, C. K. & Suttie, J. W. Comparative distribution, metabolism, and utilization of phylloquinone and menaquinone-9 in rat liver. P. Soc. Exp. Biol. Med. 209, 403–409 (1995).
Kadokura, H. & Beckwith, J. Mechanisms of oxidative protein folding in the bacterial cell envelope. Antioxid. Redox Sign 13, 1231–1246 (2010).
Inaba, K. Structural basis of protein disulfide bond generation in the cell. Genes Cells 15, 935–943 (2010).
Denoncin, K. & Collet, J.-F. Disulfide bond formation in the bacterial periplasm: major achievements and challenges ahead. Antioxid. Redox Sign 19, 63–71 (2012).
Hatahet, F., Boyd, D. & Beckwith, J. Disulfide bond formation in prokaryotes: history, diversity and design. BBA-Proteins Proteom. 1844, 1402–1414 (2014).
Dailey, F. E. & Berg, H. C. Mutants in disulfide bond formation that disrupt flagellar assembly in Escherichia coli. Proc. Natl Acad. Sci. USA 90, 1043–1047 (1993).
Missiakas, D., Georgopoulos, C. & Raina, S. Identification and characterization of the Escherichia coli gene dsbB, whose product is involved in the formation of disulfide bonds in vivo. Proc. Natl Acad. Sci. USA 90, 7084–7088 (1993).
Kadokura, H., Katzen, F. & Beckwith, J. Protein disulfide bond formation in prokaryotes. Annu. Rev. Biochem. 72, 111–135 (2003).
Jander, G., Martin, N. L. & Beckwith, J. Two cysteines in each periplasmic domain of the membrane protein DsbB are required for its function in protein disulfide bond formation. EMBO J. 13, 5121–5127 (1994).
Kobayashi, T. et al. Respiratory chain is required to maintain oxidized states of the DsbA–DsbB disulfide bond formation system in aerobically growing Escherichia coli cells. Proc. Natl Acad. Sci. USA 94, 11857–11862 (1997).
Bader, M. W., Xie, T., Yu, C. A. & Bardwell, J. C. A. Disulfide bonds are generated by quinone reduction. J. Biol. Chem. 275, 26082–26088 (2000).
Berkmen, M., Boyd, D. & Beckwith, J. The nonconsecutive disulfide bond of Escherichia coli phytase (AppA) renders it dependent on the protein-disulfide isomerase, DsbC. J. Biol. Chem. 280, 11387–11394 (2005).
Missiakas, D., Georgopoulos, C. & Raina, S. The Escherichia coli dsbC (xprA) gene encodes a periplasmic protein involved in disulfide bond formation. EMBO J. 13, 2013–2020 (1994).
Shevchik, V. E., Condemine, G. & Robert-Baudouy, J. Characterization of DsbC, a periplasmic protein of Erwinia chrysanthemi and Escherichia coli with disulfide isomerase activity. EMBO J. 13, 2007–2012 (1994).
Rietsch, A., Belin, D., Martin, N. & Beckwith, J. An in vivo pathway for disulfide bond isomerization in Escherichia coli. Proc. Natl Acad. Sci. USA 93, 13048–13053 (1996).
Missiakas, D., Schwager, F. & Raina, S. Identification and characterization of a new disulfide isomerase-like protein (DsbD) in Escherichia coli. EMBO J. 14, 3415–3424 (1995).
Stewart, E. J., Katzen, F. & Beckwith, J. Six conserved cysteines of the membrane protein DsbD are required for the transfer of electrons from the cytoplasm to the periplasm of Escherichia coli. EMBO J. 18, 5963–5971 (1999).
Lin, D., Rao, C. V. & Slauch, J. M. The Salmonella SPI1 type three-secretion system responds to periplasmic disulfide bond status via the flagellar apparatus and the RcsCDB system. J. Bacteriol. 190, 87–97 (2008).
Totsika, M., Heras, B., Wurpel, D. J. & Schembri, M. A. Characterization of two homologous disulfide bond systems involved in virulence factor biogenesis in uropathogenic Escherichia coli CFT073. J. Bacteriol. 191, 3901–3908 (2009).
Kwon, A.-R. & Choi, E.-C. Role of disulfide bond of arylsulfate sulfotransferase in the catalytic activity. Arch. Pharm. Res. 28, 561–565 (2005).
Grimshaw, J. P. A. et al. DsbL and DsbI form a specific dithiol oxidase system for periplasmic arylsulfate sulfotransferase in uropathogenic Escherichia coli. J. Mol. Biol. 380, 667–680 (2008).
Lin, D., Kim, B. & Slauch, J. M. DsbL and DsbI contribute to periplasmic disulfide bond formation in Salmonella enterica serovar Typhimurium. Microbiology 155, 4014–4024 (2009).
Miki, T., Okada, N. & Danbara, H. Two periplasmic bisulfide oxidoreductases, DsbA and SrgA, target outer membrane protein SpiA, a component of the Salmonella pathogenicity island 2 type III secretion system. J. Biol. Chem. 279, 34631–34642 (2004).
Ha, U., Wang, Y. & Jin, S. DsbA of Pseudomonas aeruginosa is essential for multiple virulence factors. Infect. Immun. 71, 1590–1595 (2003).
Kim, S. H., Park, S. Y., Heo, Y. J. & Cho, Y. H. Drosophila melanogaster-based screening for multihost virulence factors of Pseudomonas aeruginosa PA14 and identification of a virulence-attenuating factor, HudA. Infect. Immun. 76, 4152–4162 (2008).
Lasica, A. M., Wyszynska, A., Szymanek, K., Majewski, P. & Jagusztyn-Krynicka, E. K. Campylobacter protein oxidation influences epithelial cell invasion or intracellular survival as well as intestinal tract colonization in chickens. J. Appl. Genet. 51, 383–393 (2010).
Grabowska, A. D. et al. Functional and bioinformatics analysis of two Campylobacter jejuni homologs of the thiol-disulfide oxidoreductase, DsbA. PLoS ONE 9, e106247 (2014).
Kpadeh, Z. Z., Day, S. R., Mills, B. W. & Hoffman, P. S. Legionella pneumophila utilizes a single-player disulfide-bond oxidoreductase system to manage disulfide bond formation and isomerization. Mol. Microbiol. 95, 1054–1069 (2015).
Heras, B. et al. Structural and functional characterization of three DsbA paralogues from Salmonella enterica serovar typhimurium. J. Biol. Chem. 285, 18423–18432 (2010).
Arts, I. S. et al. Dissecting the machinery that introduces disulfide bonds in Pseudomonas aeruginosa. mBio 4, e00912-13 (2013).
Tinsley, C. R., Voulhoux, R., Beretti, J. L., Tommassen, J. & Nassif, X. Three homologues, including two membrane-bound proteins, of the disulfide oxidoreductase DsbA in Neisseria meningitidis: effects on bacterial growth and biogenesis of functional type IV pili. J. Biol. Chem. 279, 27078–27087 (2004).
Sinha, S., Langford, P. R. & Kroll, J. S. Functional diversity of three different DsbA proteins from Neisseria meningitidis. Microbiology 150, 2993–3000 (2004).
Sinha, S., Ambur, O. H., Langford, P. R., Tønjum, T. & Kroll, J. S. Reduced DNA binding and uptake in the absence of DsbA1 and DsbA2 of Neisseria meningitidis due to inefficient folding of the outer-membrane secretin PilQ. Microbiology 154, 217–225 (2008).
Kpadeh, Z. Z. et al. Disulfide bond oxidoreductase DsbA2 of Legionella pneumophila exhibits protein disulfide isomerase activity. J. Bacteriol. 195, 1825–1833 (2013).
Ren, G., Champion, M. M. & Huntley, J. F. Identification of disulfide bond isomerase substrates reveals bacterial virulence factors. Mol. Microbiol. 94, 926–944 (2014).
Qin, A. et al. FipB, an essential virulence factor of Francisella tularensis subsp. tularensis, has dual roles in disulfide bond formation. J. Bacteriol. 196, 3571–3581 (2014).
Arredondo, S. A., Chen, T. F., Riggs, A. F., Gilbert, H. F. & Georgious, G. Role of dimerization in the catalytic properties of the Escherichia coli disulfide isomerase DsbC. J. Biol. Chem. 284, 23972–23979 (2009).
Jameson-Lee, M., Garduño, R. A. & Hoffman, P. S. DsbA2 (27kDa Com1-like protein) of Legionella pneumophila. catalyses extracytoplasmic disulphide-bond formation in proteins including the Dot/Icm type IV secretion system. Mol. Microbiol 80, 835–852 (2011).
Bocian-Ostrzycka, K. M., Grzeszczuk, M. J., Dziewit, L. & Jagusztyn-Krynicka, E. K. Diversity of the Epsilonproteobacteria Dsb (disulfide bond) systems. Front. Microbiol. 6, 570 (2015).
Raczko, A. M. et al. Characterization of new DsbB-like thiol-oxidoreductases of Campylobacter jejuni and Helicobacter pylori and classification of the DsbB family based on phylogenomic, structural and functional criteria. Microbiology 151, 219–231 (2005).
Yoon, J. Y. et al. Structural and functional characterization of Helicobacter pylori. DsbG. FEBS Lett. 585, 3862–3867 (2011).
Roszczenko, P., Radomska, K. A., Wywial, E., Collet, J. F. & Jagusztyn-Krynicka, E. K. A novel insight into the oxidoreductase activity of Helicobacter pylori HP0231 protein. PLoS ONE 7, e46563 (2012).
Lester, J. et al. Characterization of Helicobacter pylori HP0231 (DsbK): role in disulfide bond formation, redox homeostasis and production of Helicobacter cystein-rich protein HcpE. Mol. Microbiol. 96, 110–133 (2015).
Bocian-Ostrzycka, K. M. et al. Engineering of Helicobacter pylori dimeric oxidoreductase Dsbk (HP0231). Front. Microbiol. 7, 1158 (2016).
Landeta, C. et al. Compounds targeting disulfide bond forming enzyme DsbB of Gram-negative bacteria. Nat. Chem. Biol. 11, 292–298 (2015).
Meehan, B. M., Landeta, C., Boyd, D. & Beckwith, J. The essential cell division protein FtsN contains a critical disulfide bond in a non-essential domain. Mol. Microbiol. 103, 413–422 (2016).
Meehan, B. M., Landeta, C., Boyd, D. & Beckwith, J. The disulfide bond formation pathway is essential for anaerobic growth of Escherichia coli. J. Bacteriol. 199, e00120-17 (2017).
Hizukuri, Y., Yakushi, T., Kawagishi, I. & Homma, M. Role of the intramolecular disulfide bond in FlgI, the flagellar P-ring component of Escherichia coli. J. Bacteriol. 188, 4190–4197 (2006).
Dai, K., Xu, Y., Lutkenhaus, J. & Lutkenhaus, J. O. E. Topological characterization of the essential Escherichia coli cell division protein FtsN. J. Bacteriol. 178, 1328–1334 (1996).
Bojkovic, J. et al. Characterization of an Acinetobacter baumannii lptD deletion strain: permeability defects and response to inhibition of lipopolysaccharide and fatty acid biosynthesis. J. Bacteriol. 198, 731–741 (2016).
Peterson, K. M. & Mekalanos, J. J. Characterization of the Vibrio cholerae ToxR regulon: identification of novel genes involved in intestinal colonization. Infect. Immun. 56, 2822–2829 (1988).
Yu, J., Webb, H. & Hirst, T. R. A homologue of the Escherichia coli DsbA protein involved in disulphide bond formation is required for enterotoxin biogenesis in Vibrio cholerae. Mol. Microbiol. 6, 1949–1958 (1992).
Pogliano, J., Lynch, A. S., Belin, D., Lin, E. C. & Beckwith, J. Regulation of Escherichia coli cell envelope proteins involved in protein folding and degradation by the Cpx two-component system. Genes Dev. 11, 1169–1182 (1997).
Grabowska, A. D. et al. Campylobacter jejuni dsb gene expression is regulated by iron in a Fur-dependent manner and by a translational coupling mechanism. BMC Microbiol. 11, 166 (2011).
Bayan, N., Houssin, C., Chami, M. & Leblon, G. Mycomembrane and S-layer: two important structures of Corynebacterium glutamicum cell envelope with promising biotechnology applications. J. Biotechol. 104, 55–67 (2003).
Matias, V. R. F. & Beveridge, T. J. Native cell wall organization shown by cryo-electron microscopy confirms the existence of a periplasmic space in Staphylococcus aureus. J. Bacteriol. 188, 1011–1021 (2006).
Matias, V. R. F. & Beveridge, T. J. Cryo-electron microscopy reveals native polymeric cell wall structure in Bacillus subtilis 168 and the existence of a periplasmic space. Mol. Microbiol. 56, 240–251 (2005).
Burkovski, A. & Burkovski, A. Cell envelope of Corynebacteria: structure and influence on pathogenicity. ISRN Microbiol. 2013, 935736 (2013).
Singh, A. K., Bhattacharyya-Pakrasi, M. & Pakrasi, H. B. Identification of an atypical membrane protein involved in the formation of protein disulfide bonds in oxygenic photosynthetic organisms. J. Biol. Chem. 283, 15762–15770 (2008).
Goodstadt, L. & Ponting, C. P. Vitamin K epoxide reductase: homology, active site and catalytic mechanism. Trends Biochem. Sci. 29, 289–292 (2004).
Wang, X., Dutton, R. J., Beckwith, J. & Boyd, D. Membrane topology and mutational analysis of Mycobacterium tuberculosis VKOR, a protein involved in disulfide bond formation and a homologue of human vitamin K epoxide reductase. Antioxid. Redox Sign 14, 1413–1420 (2011).
Premkumar, L. et al. Rv2969c, essential for optimal growth in Mycobacterium tuberculosis, is a DsbA-like enzyme that interacts with VKOR-derived peptides and has atypical features of DsbA-like disulfide oxidases. Acta Crystallogr. Sect. D 69, 1981–1994 (2013).
Kurosu, M. & Begari, E. Vitamin K2 in electron transport system: are enzymes involved in vitamin K2 biosynthesis promising drug targets? Molecules 15, 1531–1553 (2010).
Dutton, R. J. et al. Inhibition of bacterial disulfide bond formation by the anticoagulant warfarin. Proc. Natl Acad. Sci. USA 107, 297–301 (2010).
Daniels, R. et al. Disulfide bond formation and cysteine exclusion in gram-positive bacteria. J. Biol. Chem. 285, 3300–3309 (2010).
Patarroyo, M. A. et al. Functional characterization of Mycobacterium tuberculosis Rv2969c membrane protein. Biochem. Bioph. Res. Co. 372, 935–940 (2008).
Chim, N., Harmston, C. A., Guzman, D. J. & Goulding, C. W. Structural and biochemical characterization of the essential DsbA-like disulfide bond forming protein from Mycobacterium tuberculosis. BMC Struct. Biol. 13, 23 (2013).
Goulding, C. W. et al. Gram-positive DsbE proteins function differently from gram-negative DsbE homologs: a structure to function analysis of DsbE from mycobacterium tuberculosis. J. Biol. Chem. 279, 3516–3524 (2004).
Chim, N. et al. An extracellular disulfide bond forming protein (DsbF) from Mycobacterium tuberculosis: structural, biochemical, and gene expression analysis. J. Mol. Biol. 396, 1211–1226 (2010).
Sassetti, C. M. & Rubin, E. J. Genetic requirements for mycobacterial survival during infection. Proc. Natl Acad. Sci. USA 100, 12989–12994 (2003).
Reardon-Robinson, M. E. et al. A disulfide bond-forming machine is linked to the sortase-mediated pilus assembly pathway in the Gram-positive bacterium Actinomyces oris. J. Biol. Chem. 290, 21393–21405 (2015).
Reardon-Robinson, M. E. & Ton-That, H. Disulfide-bond-forming pathways in Gram-positive bacteria. J. Bacteriol. 198, 746–754 (2016).
Reardon-Robinson, M. E. et al. A thiol-disulfide oxidoreductase of the Gram-positive pathogen Corynebacterium diphtheriae is essential for viability, pilus assembly, toxin production and virulence. Mol. Microbiol. 98, 1037–1050 (2015).
Ishihara, T. et al. Cloning and characterization of the gene for a protein thiol-disulfide oxidoreductase in Bacillus brevis. J. Bacteriol. 177, 745–749 (1995).
Bolhuis, A. Functional analysis of paralogous thiol-disulfide oxidoreductases in Bacillus subtilis. J. Biol. Chem. 274, 24531–24538 (1999).
Crow, A. et al. Crystal structure and biophysical properties of Bacillus subtilis BdbD. An oxidizing thiol:disulfide oxidoreductase containing a novel metal site. J. Biol. Chem. 284, 23719–23733 (2009).
Meima, R. et al. The bdbDC operon of Bacillus subtilis encodes thiol-disulfide oxidoreductases required for competence development. J. Biol. Chem. 277, 6994–7001 (2002).
Draskovic, I. & Dubnau, D. Biogenesis of a putative channel protein, ComEC, required for DNA uptake: membrane topology, oligomerization and formation of disulphide bonds. Mol. Microbiol. 55, 881–896 (2005).
Dorenbos, R. et al. Thiol-disulfide oxidoreductases are essential for the production of the lantibiotic sublancin 168. J. Biol. Chem. 277, 16682–16688 (2002).
Kouwen, T. R. H. M. et al. Thiol-disulphide oxidoreductase modules in the low-GC Gram-positive bacteria. Mol. Microbiol. 64, 984–999 (2007).
Dumoulin, A., Grauschopf, U., Bischoff, M., Thöny-Meyer, L. & Berger-Bächi, B. Staphylococcus aureus DsbA is a membrane-bound lipoprotein with thiol-disulfide oxidoreductase activity. Arch. Microbiol. 184, 117–128 (2005).
Heras, B. et al. Staphylococcus aureus DsbA does not have a destabilizing disulfide: a new paradigm for bacterial oxidative folding. J. Biol. Chem. 283, 4261–4271 (2008).
van der Kooi-Pol, M. M. et al. Requirement of signal peptidase ComC and thiol-disulfide oxidoreductase DsbA for optimal cell surface display of pseudopilin ComGC in Staphylococcus aureus. Appl. Environ. Microb. 78, 7124–7127 (2012).
Lacy, D. B., Tepp, W., Cohen, A. C., DasGupta, B. R. & Stevens, R. C. Crystal structure of botulinum neurotoxin type A and implications for toxicity. Nat. Struct. Biol. 5, 898–902 (1998).
Marvaud, J. C. et al. botr/A is a positive regulator of botulinum neurotoxin and associated non-toxin protein genes in Clostridium botulinum A. Mol. Microbiol. 29, 1009–1018 (1998).
Baker, M. D., Gendlina, I., Collins, C. M. & Acharya, K. R. Crystal structure of a dimeric form of streptococcal pyrogenic exotoxin A (SpeA1). Protein Sci. 13, 2285–2290 (2004).
Davey, L., Ng, C. K. W., Halperin, S. A. & Lee, S. F. Functional analysis of paralogous thiol-disulfide oxidoreductases in Streptococcus gordonii. J. Biol. Chem. 288, 16416–16429 (2013).
Davey, L., Cohen, A., Leblanc, J., Halperin, S. A. & Lee, S. F. The disulfide oxidoreductase SdbA is active in Streptococcus gordonii using a single C-terminal cysteine of the CXXC motif. Mol. Microbiol. 99, 236–253 (2016).
Chng, S. S. et al. Overexpression of the rhodanese PspE, a single cysteine-containing protein, restores disulphide bond formation to an Escherichia coli strain lacking DsbA. Mol. Microbiol. 85, 996–1006 (2012).
Derman, A., Prinz, W. A., Belin, D. & Beckwith, J. Mutations that allow disulfide bond formation in the cytoplasm of Escherichia coli. Science 262, 1744–1747 (1993).
Bessette, P. H., Aslund, F., Beckwith, J. & Georgiou, G. Efficient folding of proteins with multiple disulfide bonds in the Escherichia coli cytoplasm. Proc. Natl Acad. Sci. USA 96, 13703–13708 (1999).
Hatahet, F. & Ruddock, L. W. Topological plasticity of enzymes involved in disulfide bond formation allows catalysis in either the periplasm or the cytoplasm. J. Mol. Biol. 425, 3268–3276 (2013).
Ladenstein, R. & Ren, B. Protein disulfides and protein disulfide oxidoreductases in hyperthermophiles. FEBS J. 273, 4170–4185 (2006).
Jorda, J. & Yeates, T. O. Widespread disulfide bonding in proteins from thermophilic archaea. Archaea 2011, 409156 (2011).
Beeby, M. et al. The genomics of disulfide bonding and protein stabilization in thermophiles. PLoS Biol. 3, 1549–1558 (2005).
Ladenstein, R. & Ren, B. Reconsideration of an early dogma, saying ‘there is no evidence for disulfide bonds in proteins from archaea’. Extremophiles 12, 29–38 (2008).
Hibender, S., Landeta, C., Berkmen, M., Beckwith, J. & Boyd, D. Aeropyrum pernix membrane topology of protein VKOR promotes protein disulfide bond formation in two subcellular compartments. Microbiology 12, 1864–1879 (2017).
Pedone, E., Ren, B., Ladenstein, R., Rossi, M. & Bartolucci, S. Fuctional properties of the protein disulfide oxidoreductase from the Archaeon: Pryococcus furioses. Eur. J. Biochem. 271, 3437–3448 (2004).
Pedone, E., Limauro, D., D’Alterio, R., Rossi, M. & Bartolucci, S. Characterization of a multifunctional protein disulfide oxidoreductase from Sulfolobus solfataricus. FEBS J. 273, 5407–5420 (2006).
D’Ambrosio, K. et al. A novel member of the protein disulfide oxidoreductase family from Aeropyrum pernix K1: structure, function and electrostatics. J. Mol. Biol. 362, 743–752 (2006).
Alvarez, A. F., Rodriguez, C. & Georgellis, D. Ubiquinone and menaquinone electron carriers represent the yin and yang in the redox regulation of the ArcB sensor kinase. J. Bacteriol. 195, 3054–3061 (2013).
Maskos, K., Huber-Wunderlich, M. & Glockshuber, R. DsbA and DsbC-catalyzed oxidative folding of proteins with complex disulfide bridge patterns in vitro and in vivo. J. Mol. Biol. 325, 495–513 (2003).
Lobstein, J. et al. SHuffle, a novel Escherichia coli protein expression strain capable of correctly folding disulfide bonded proteins in its cytoplasm. Microb. Cell Fact. 11, 753 (2012).
Robinson, M. et al. Efficient expression of full-length antibodies in the cytoplasm of engineered bacteria. Nat. Commun. 6, 8072 (2015).
Ritz, D., Lim, J., Reynolds, C. M., Poole, L. B. & Beckwith, J. Conversion of a peroxiredoxin into a disulfide reductase by a triplet repeat expansion. Science 294, 158–161 (2001).
Davey, L., Halperin, S. A. & Lee, S. F. Thiol-disulfide exchange in Gram-positive Firmicutes. Trends Microbiol. 24, 902–915 (2016).
Fleury, Y. et al. Covalent structure, synthesis, and structure-function studies of Mesentericin Y 10537, a defensive peptide from Gram-positive Bacteria Leuconostoc mesenteroides. J. Biol. Chem. 271, 14421–14429 (1996).
Kawai, Y. et al. Primary amino acid and DNA sequences of gassericin T, a lactacin F-family bacteriocin produced by Lactobacillus gasseri SBT2055. Biosci. Biotech. Bioch. 64, 2201–2208 (2000).
O’Shea, E. F. et al. Bactofencin A, a new type of cationic bacteriocin with unusual immunity. mBio 4, e00498-13 (2013).
Oppegård, C., Fimland, G., Anonsen, J. H. & Nissen-Meyer, J. The pediocin PA-1 accessory protein ensures correct disulfide bond formation in the antimicrobial peptide pediocin PA-1. Biochemistry 54, 2967–2974 (2015).
Früh, V. et al. Application of fragment-based drug discovery to membrane proteins: identification of ligands of the integral membrane enzyme DsbB. Chem. Biol. 17, 881–891 (2010).
Duprez, W. et al. Virtual screening of peptide and peptidomimetic fragments targeted to inhibit bacterial dithiol oxidase DsbA. PLoS ONE 10, e0133805 (2015).
Adams, L. A. et al. Application of fragment-based screening to the design of inhibitors of Escherichia coli DsbA. Angew. Chem. Int. Ed. 54, 2179–2184 (2015).
Halili, M. A. et al. Small molecule inhibitors of disulfide bond formation by the bacterial DsbA–DsbB dual enzyme system. ACS Chem. Biol. 10, 957–964 (2015).
Mohanty, B. et al. Fragment library screening identifies hits that bind to the non-catalytic surface of Pseudomonas aeruginosa DsbA1. PLoS ONE 12, e0173436 (2017).
Smith, R. P., Paxman, J. J., Scanlon, M. J. & Heras, B. Targeting bacterial Dsb proteins for the development of anti-virulence agents. Molecules 21, 811 (2016).
Duprez, W. et al. Peptide inhibitors of the Escherichia coli DsbA oxidative machinery essential for bacterial virulence. J. Med. Chem. 58, 577–587 (2015).
Landeta, C. et al. Inhibition of virulence-promoting disulfide bond formation enzyme DsbB is blocked by mutating residues in two distinct regions. J. Biol. Chem. 292, 6529–6541 (2017).
Kang, H. J., Coulibaly, F., Clow, F., Proft, T. & Baker, E. N. Stabilizing isopeptide bonds revealed in Gram-positive bacterial pilus structure. Science 318, 1625–1628 (2007).
Kang, H. J. & Baker, E. N. Structure and assembly of Gram-positive bacterial pili: unique covalent polymers. Curr. Opin. Struc. Biol. 22, 200–207 (2012).
Wikoff, W. R. et al. Topologically linked protein rings in the bacteriophage HK97 capsid. Science 289, 2129–2133 (2000).
Vincent-Sealy, L., Thomas, J. D., Commander, P. & Salmond, G. P. C. Secreted proteins. Microbiology 145, 1945–1958 (1999).
Kloek, aP., Brooks, D. M. & Kunkel, B. N. A dsbA mutant of Pseudomonas syringae exhibits reduced virulence and partial impairment of type III secretion. Mol. Plant Pathol. 1, 139–150 (2000).
Gonzalez, M. D., Lichtensteiger, C. A. & Vimr, E. R. Adaptation of signature-tagged mutagenesis to Escherichia coli K1 and the infant-rat model of invasive disease. FEMS Microbiol. Lett. 198, 125–128 (2001).
Herbert, Ma et al. Signature tagged mutagenesis of Haemophilus influenzae identifies genes required for in vivo survival. Microb. Pathog. 33, 211–223 (2002).
Rosadini, C. V., Wong, S. M. S. & Akerley, B. J. The periplasmic disulfide oxidoreductase DsbA contributes to Haemophilus influenzae pathogenesis. Infect. Immun. 76, 1498–1508 (2008).
Sabarth, N., Hurwitz, R., Meyer, T. F. & Bumann, D. Multiparameter selection of Helicobacter pylori antigens identifies two novel antigens with high protective efficacy. Infect. Immun. 70, 6499–6503 (2002).
Zhong, Y. et al. Helicobacter pylori HP0231 influences bacterial virulence and is essential for gastric colonization. PLoS ONE 11, e0154643 (2016).
Burall, L. S. et al. Proteus mirabilis genes that contribute to pathogenesis of urinary tract infection: identification of 25 signature-tagged mutants attenuated at least 100-fold. Infect. Immun. 72, 2922–2938 (2004).
Godlewska, R. et al. Helicobacter pylori protein oxidation influences the colonization process. Int. J. Med. Microbiol. 296, 321–324 (2006).
Jiang, B.-L. et al. DsbB is required for the pathogenesis process of Xanthomonas campestris pv. campestris. Mol. Plant Microbe In. 21, 1036–1045 (2008).
Qin, A., Scott, D. W. & Mann, B. J. Francisella tularensis subsp. tularensis Schu S4 disulfide bond formation protein B, but not an RND-type efflux pump, is required for virulence. Infect. Immun. 76, 3086–3092 (2008).
Qin, A., Scott, D. W., Thompson, J. A. & Mann, B. J. Identification of an essential Francisella tularensis subsp. tularensis virulence factor. Infect. Immun. 77, 152–161 (2009).
Straskova, A. et al. Proteome analysis of an attenuated Francisella tularensis dsbA mutant: identification of potential DsbA substrate proteins. J. Proteome Res. 8, 5336–5346 (2009).
Schmidt, M. et al. Francisella tularensis subsp. holarctica DsbA homologue: a thioredoxin-like protein with chaperone function. Microbiol. 159, 2364–2374 (2013).
Vilches, S., Jimenez, N., Merino, S. & Tomas, J. M. The Aeromonas DsbA mutation decreased their virulence by triggering type III secretion system but not flagella production. Microb. Pathog. 52, 130–139 (2012).
Ireland, P. M. et al. Disarming Burkholderia pseudomallei: structural and functional characterization of a disulfide oxidoreductase (DsbA) required for virulence in vivo. Antioxid. Redox Signal. 20, 606–617 (2014).
Ciccarelli, F. D. et al. Toward automatic reconstruction of a highly resolved tree of life. Science 311, 1283–1287 (2006).
Debarbieux, L. & Beckwith, J. O. N. On the functional interchangeability, oxidant versus reductant, of members of the thioredoxin superfamily. J. Bacteriol. 182, 723–727 (2000).
McMahon, R. M., Premkumar, L. & Martin, J. L. Four structural subclasses of the antivirulence drug target disulfide oxidoreductase DsbA provide a platform for design of subclass-specific inhibitors. BBA-Proteins Proteom. 1844, 1391–1401 (2014).
Acknowledgements
This work was supported by US National Institute of General Medical Sciences grants GMO41883 (to J.B. and D.B.) and by an industry research agreement with F. Hoffmann-La Roche Ltd. and F. Hoffmann-La Roche Inc. (to J.B. and D.B.). J.B. is an American Cancer Society Professor.
Author information
Authors and Affiliations
Contributions
C.L. and J.B. wrote the manuscript. D.B. performed the bioinformatic analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Table 1 and Supplementary References.
Rights and permissions
About this article
Cite this article
Landeta, C., Boyd, D. & Beckwith, J. Disulfide bond formation in prokaryotes. Nat Microbiol 3, 270–280 (2018). https://doi.org/10.1038/s41564-017-0106-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-017-0106-2
This article is cited by
-
Regulation of P450 TleB catalytic flow for the synthesis of sulfur-containing indole alkaloids by substrate structure-directed strategy and protein engineering
Science China Chemistry (2023)
-
The transcriptomic landscape of Magnetospirillum gryphiswaldense during magnetosome biomineralization
BMC Genomics (2022)
-
A high-throughput cell-based assay pipeline for the preclinical development of bacterial DsbA inhibitors as antivirulence therapeutics
Scientific Reports (2021)
-
Lifestyle-specific S-nitrosylation of protein cysteine thiols regulates Escherichia coli biofilm formation and resistance to oxidative stress
npj Biofilms and Microbiomes (2021)
-
Revisiting long-chain fatty acid metabolism in Escherichia coli: integration with stress responses
Current Genetics (2021)