Abstract
Inflammatory bowel disease (IBD) is a group of chronic diseases of the digestive tract that affects millions of people worldwide. Genetic, environmental and microbial factors have been implicated in the onset and exacerbation of IBD. However, the mechanisms associating gut microbial dysbioses and aberrant immune responses remain largely unknown. The integrative Human Microbiome Project seeks to close these gaps by examining the dynamics of microbiome functionality in disease by profiling the gut microbiomes of >100 individuals sampled over a 1-year period. Here, we present the first results based on 78 paired faecal metagenomes and metatranscriptomes, and 222 additional metagenomes from 59 patients with Crohn’s disease, 34 with ulcerative colitis and 24 non-IBD control patients. We demonstrate several cases in which measures of microbial gene expression in the inflamed gut can be informative relative to metagenomic profiles of functional potential. First, although many microbial organisms exhibited concordant DNA and RNA abundances, we also detected species-specific biases in transcriptional activity, revealing predominant transcription of pathways by individual microorganisms per host (for example, by Faecalibacterium prausnitzii). Thus, a loss of these organisms in disease may have more far-reaching consequences than suggested by their genomic abundances. Furthermore, we identified organisms that were metagenomically abundant but inactive or dormant in the gut with little or no expression (for example, Dialister invisus). Last, certain disease-specific microbial characteristics were more pronounced or only detectable at the transcript level, such as pathways that were predominantly expressed by different organisms in patients with IBD (for example, Bacteroides vulgatus and Alistipes putredinis). This provides potential insights into gut microbial pathway transcription that can vary over time, inducing phenotypical changes that are complementary to those linked to metagenomic abundances. The study’s results highlight the strength of analysing both the activity and the presence of gut microorganisms to provide insight into the role of the microbiome in IBD.
Similar content being viewed by others
Main
Inflammatory bowel disease (IBD) is a group of chronic inflammatory disorders that affects all or part of the digestive tract, and incidence rates are increasing worldwide1. IBD is a lifelong disease with no effective long-term treatment options, and an estimated 25–30% of all patients present with symptoms before 20 years of age2,3. Several human genetic mutations are implicated in an increased susceptibility to IBD; however, not everyone who carries these mutations develops IBD, indicating that additional exposures are also involved. Recognizing that the gut microbiome is one such factor altered in patients with Crohn’s disease (CD) and ulcerative colitis (UC), the two main forms of IBD, is one of the most important developments in the field of IBD in the past decade and provides a wealth of opportunities for the discovery of diagnostic and therapeutic approaches4,5.
A gut microbial dysbiosis exists at the community level in patients with IBD, combining a general decrease in alpha diversity with clade-specific enrichments and depletions6,7. However, microbial taxonomic profiles can be highly divergent among patients, making it difficult to implicate specific microbial species or strains in disease onset and progression. Some broad patterns do apply: taxa from the Enterobacteriaceae family are generally increased, whereas members of the Firmicutes phylum are decreased8,9,10. Furthermore, several bacterial taxa have been suggested to have protective effects in IBD, such as Lactobacillus and Faecalibacterium 11,12,13.
Most such findings are based on samples from cross-sectional cohorts, emphasizing the need for longitudinal studies to explore changes in the gut microbiome in individual patients. Indeed, gut microbiome composition is known to vary over time in individuals, and such variations may be more pronounced in patients with IBD14,15. The links between metagenomic functional potential and realized functional activity (gene expression or other molecular products, such as proteins or metabolites) remain almost completely unexplored in IBD. Although the overall functional potential of the gut microbiome tends to be stable relative to taxonomic composition16, it can vary over time and across phenotypes. For example, short-chain fatty acid (SCFA)-producing bacteria are depleted in IBD; SFCAs are metabolites that are broadly anti-inflammatory in a typical gut17,18. Furthermore, an increase in oxidative stress pathways, and a decrease in carbohydrate metabolism and amino acid biosynthesis have also been consistently reported6, each affecting multiple underlying taxa in different individuals. Notably, the functional potential of an organism (that is, the genes and pathways encoded in its genome) provides only indirect information about the level or extent to which these functions are active. Such measures of functional activity are critical for understanding the mechanisms associating gut microbial dysbioses and aberrant immune responses, which, to date, remain largely unknown. Alterations in transcriptional activity in IBD have been established based on ribosomal RNA expression and indicate that some bacterial populations are active in patients with IBD, whereas other groups are inactive or dormant in disease19. However, the specific bacterial species and metabolic pathways remain to be elucidated.
To close these gaps, we compared the functional potential of gut microbial communities (from shotgun metagenomics) to direct measures of functional activity (from metatranscriptomics) in a longitudinal cohort. Faecal samples were collected biweekly from ~100 patients over the course of 1 year and subjected to shotgun metagenomic (DNA) and metatranscriptomic (RNA) sequencing. Here, we present the results based on 78 paired metagenomes and metatranscriptomes and an additional 222 metagenomes. For many species, functional potential was well correlated with functional activity. However, we were able to identify species-specific shifts in transcript levels, indicating that some organisms (for example, Faecalibacterium prausnitzii) and pathways may have a more central role in maintaining gut health than their genomic abundances indicate. We also detected organisms that were metagenomically present but with low or non-existent gene expression (for example, Dialister invisus), suggesting that the organism is either dead or inactive and therefore of questionable importance in the gut community. In addition, we uncovered disease-specific changes in microbial gene expression that were either more pronounced or only detectable on the RNA level (for example, metabolic pathways that were contributed by Bacteroides vulgatus and Alistipes putredinis). Together, our findings highlight that crucial insight into microbial community dynamics can be gained through integrated analysis of metatranscriptomic and metagenomic profiles of microbial community structure and function. This approach will lead to a better understanding of the underlying mechanisms of gut microbial dysbioses and their role in IBD.
Results
A longitudinal IBD cohort profiled using metagenomic and metatranscriptomic sequencing
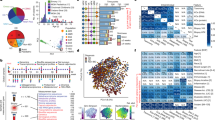
As part of the Integrative Human Microbiome Project (iHMP; also known as HMP2), the goal of the IBD Multi’omics Database (IBDMDB; http://ibdmdb.org) is to assemble longitudinal multi’omic profiles of patients with IBD to gain insight into the mechanisms of microbial dysbioses and their effects on disease onset and progression (Fig. 1). Participants provided biweekly stool samples over the course of 1 year, from which we generated shotgun metagenomic and metatranscriptomic sequencing data. In this one of several resulting data sets and studies, we analysed 78 paired metagenomes/metatranscriptomes and an additional 222 metagenomes from 117 individuals: 59 patients with CD, 34 with UC and 24 non-IBD controls (Fig. 1a). These data sets yielded high-resolution profiles of gut microbial community composition (taxonomy), functional potential and functional activity.
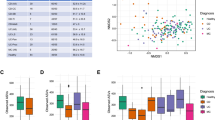
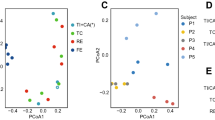
a, Biweekly stool samples were collected from 117 individuals (59 with CD, 34 with UC and 24 non-IBD controls) over the course of 1 year. The resulting data comprised 78 paired stool metagenomes and metatranscriptomes as well as 222 additional metagenomes. While most samples are part of short time courses, two long time courses per disease phenotype (with up to 17 time points) were also included. Each row corresponds to the time course of a subject. b, Principal coordinate analysis (PCoA) on Bray–Curtis distance at the species level for all 300 metagenomic taxonomic profiles from 117 patients. In addition, 19 species are overlaid where their position represents the species’ weighted average score and indicates that samples in close proximity probably contain higher abundances of the respective organism. While the time courses highlight that inter-individual effects dominate the variation in taxonomic composition, longitudinal shifts were also observed, motivating the investigation of inter-personal and intra-personal dynamic patterns of microbial species.
Consistent with previous studies, taxonomic shifts in microbial composition inferred from the metagenomic data accounted for significant, but modest effect size, separation among the three phenotypes (Fig. 1b). Longitudinal profiling further emphasized that variation in microbial community composition is dominated by inter-individual effects, as samples from the same subject tended to cluster tightly. However, we also observed taxonomic shifts in community composition over time that coincided with changes in disease severity (as measured by the Harvey–Bradshaw Index (HBI)) and antibiotic treatment (Supplementary Fig. 1). This highlights the importance of examining longitudinal profiles to establish a better understanding of species dynamics within and across patients.
Inter-personal and intra-personal dynamic patterns of microbial species
To better understand the variability in microbial species composition within and across individuals, we first examined the taxonomic profiles from six long time courses (that is, 2 patients with CD, 2 with UC and 2 non-IBD controls, with at least 12 samples each over the 1-year sampling period; Supplementary Fig. 2a). Three general patterns were observed: (1) intra-personal stability, (2) global stability, and (3) inter-personal and intra-personal variability. Intra-personal stability refers to species that were only encountered in individual patients or in a subset of the patients and represented permanent members of their gut microbial community (Supplementary Fig. 2b). The relative abundances of these species often remained fairly stable over the course of the year. Among these patient-specific microbial organisms were several Bacteroides species, suggesting that these closely related organisms can contribute similar functions in different patients.
By contrast, we also encountered many examples of microorganisms that were universally present in all patients at high abundance (1–10%), including F. prausnitzii and B. vulgatus, two species that are implicated in gut inflammation and IBD specifically13,17,20,21 (Supplementary Fig. 2c). Their tendency to be present in all patients irrespective of disease phenotype suggests that their abundance or transcriptional activity rather than presence/absence has a role in gut inflammation. Finally, some species displayed inter-personal and intra-personal variability patterns, intermittently disappearing and reappearing in all six patients over time (Supplementary Fig. 2d). Among these were prominent IBD-associated organisms, such as Ruminococcus gnavus and Roseburia intestinalis, highlighting that taxonomic variability is not only observed between patients but also within a patient over time22,23. Furthermore, spikes in the relative abundance of R. gnavus were observed in some patients with IBD in the overall cohort, with relative abundances of up to 19%.
Functional potential is often, but not always, proportional to metatranscriptomic expression in the gut microbiome
To compare and contrast community functional potential and functional activity, we profiled the 78 paired metagenomes and metatranscriptomes with HUMAnN2 (ref. 24), which outputs per-sample pathway abundance, stratified according to individual species’ contributions (see Methods). Averaging first within and then across patients, we found that species contributing more pathway copies to the total pool of microbial genomic DNA (that is, more metagenomically abundant species) also tended to contribute more pathway transcripts (Fig. 2a).
a, Summary of metagenomic and metatranscriptomic activities across all analysed pathways as assigned to individual species (based on 78 datasets). The activity of each species is first averaged within and then across participants for DNA and RNA samples, respectively; only species that were detected in at least 10% of the samples are shown. b,c, Each point represents the cummulative contribution of a species across all pathways on the DNA and RNA level. Detailed examples are shown for Parabacteriodes merdae (b, high DNA–RNA correlation; Spearman r = 0.85, P = 0) and F. prausnitzii (c, low correlation; Spearman r = 0.35, P = 0.0019). Here, each point represents one sample (n = 78), indicating the species’ overall contribution to all pathways in its metagenome and metatranscriptome, including a linear fit with a 95% confidence interval. d,e Species that exhibited the largest (d) and smallest (e) disease-specific changes after repeating the analysis described in a for each disease group separately. Results are summarized as triangles for each species connecting vertices, representing the values for each disease group (CD, n = 46; UC, n = 21; non-IBD, n = 11). Overall, although functional potential is indicative of functional activity for many microorganisms, others exhibited low DNA–RNA correlation or disease-specific differences in functional activity. For a complete list of taxa labelled in a,d and e, see Supplementary Table 1.
Moreover, among metagenomically abundant species (which are less sensitive to undersampling), mean pathway abundance at the DNA and RNA levels tended to correlate well across samples. This correlation was particularly strong for Parabacteroides merdae, a mucin-degrading, SCFA producer25,26 (Spearman r = 0.85; Fig. 2b). Such a strong correlation suggests that the total transcript output of P. merdae is relatively constant across samples, and hence, samples with a larger P. merdae cell fraction (that is, metagenomic relative abundance) coincide with an increase in contributions from P. merdae in the total transcript pool. This behaviour is in contrast to that of D. invisus: a bacterium that is predominantly associated with the human oral cavity, but has also been detected in the gut and has been implicated in diseases such as IBD and type 1 diabetes mellitus22,27,28. Although the average DNA abundance of D. invisus was comparable to that of P. merdae, and varied across samples, D. invisus was largely absent from the metatranscriptomic data (Supplementary Fig. 3a). This suggests that D. invisus is not actively transcribing in the gastrointestinal tract, which is consistent with a dead or non-growing population. Although we infer that organisms such as D. invisus are not transcriptionally active when we do not observe any transcripts from those organisms, it is technically possible that different microbial species might have different RNA stability, resulting in faster degradation of RNA from one organism relative to another. Nevertheless, such dramatic differences in transcriptional behaviour between metagenomically similar species underscore the importance of measuring functional potential and activity in tandem.
F. prausnitzii exhibited a third distinct pattern of behaviour: poor correlation in total DNA and RNA abundances across samples despite being abundant on average across both data types (Fig. 2c). F. prausnitzii is a known producer of butyrate, a SCFA that has a role in maintaining gut health and barrier function, and previous studies have found the species to be depleted in patients with IBD13,17,20. Although F. prausnitzii remained fairly stably abundant in patients with IBD in the long time courses (Supplementary Fig. 2c), the metatranscriptomic data indicate that the metagenomic abundance of this species is not predictive of its relative transcriptional activity (Fig. 2c).
Disease-specific differences in functional activity of microorganisms
The patterns of species-specific microbial transcription introduced above can be further stratified to identify interactions with IBD phenotype (Supplementary Fig. 4). More specifically, if we define a dysbiosis as a shift in a species’ mean pathway contributions between IBD phenotypes (UC or CD) and non-IBD controls, a species can be dysbiotic at the DNA level, the RNA level, or both, and in any combination of directions. The seven species that exhibited the largest such dysbioses are summarized in Fig. 2d (for a complete list see Supplementary Table 1). Of these, two exhibited a more pronounced dysbiosis in their functional activity than their functional potential. R. gnavus exhibited the largest amplification of disease-specific dysbiosis on the RNA level, with greatly increased RNA abundance in both patients with CD and patients with UC compared with non-IBD controls (about three orders of magnitude) in a background of a smaller increase in DNA abundance (about one order of magnitude). Hence, small changes in the abundance of R. gnavus may be more consequential than previously assumed.
The two species that exhibited the largest difference in community DNA compared to RNA were Clostridium symbiosum and Bacteroides faecis. One possible implication may be that the effect of these species on disease progression is less critical than suggested by the metagenomic data alone. Although the preceding examples focused on dysbioses that were consistent across the two IBD subtypes, this was not always the case. For example, Bacteroides fragilis was less abundant in DNA from patients with UC than in DNA from non-IBD controls, whereas it was more abundant in patients with CD. Conversely, the species' mean RNA abundance was similar between patients with CD and non-IBD controls, but markedly lower in patients with UC. Furthermore, we observed many examples of species whose overall expression was comparable across disease phenotypes with similar DNA and RNA pathway abundances (Fig. 2e). These organisms are therefore unlikely to have a role in disease onset and progression.
Contrasting metabolic functional potential and functional activity
To compare the functional potential and activity of the entire microbial community, we next investigated overall metabolic pathway abundances in both data types (Fig. 3). We used contributional alpha diversity as a measurement to compare the diversity of organisms contributing metagenomically and metatranscriptomically to each pathway (Methods). This allowed us to distinguish pathways that were contributed by a single or few microorganisms, representing specialized metabolic processes, from pathways that are contributed by a multitude of organisms, representing more essential metabolic processes (Fig. 3a).
a, Contributional alpha diversity of species represented in DNA (metagenomic) and RNA (metatranscriptomic) pathway profiles, calculated across all 78 paired samples. The mean, first and third quartiles are displayed. Black dots represent outliers, values outside 1.5 times the interquartile range. Pathways are ordered by the sum of their median DNA-level and RNA-level diversity measures. Although diversity was generally lower at the transcriptional level, as expected, we also observed that many specialized pathways were contributed by only one or few microorganisms. biosyn., biosynthesis; degr., degradation. b, Differences in per-pathway DNA versus RNA contributions of microbial species, each point representing one pathway averaged first within samples from the same patient and subsequently across patients. The mean, first and third quartiles are displayed. Only pathways detected in at least 20% of metagenomic and metatranscriptomic samples (n = 78) were included for a particular species, and only species contributing to at least 8 pathways across samples are displayed. Some microorganisms exhibited a general tendency for overtranscription or undertranscription, whereas others displayed pathway-specific activity patterns. c, Example where pathway transcription is dominated by a single species. Relative contributions of the top 20 species in metagenomes (DNA) versus metatranscriptomes (RNA) for the dTDP-l-rhamnose biosynthesis I pathway (DTDPRHAMSYN–PWY) (n = 78). This pathway was one of the most overtranscribed pathways of F. prausnitzii (b), the species with the largest range of pathway-specific overtranscription and undertranscription.
The two pathways with the lowest contributional diversity on the DNA and RNA levels were almost entirely contributed by F. prausnitzii: (1) GALACT−GLUCUROCAT−PWY, the superpathway of galacturonate and glucuronate degradation (also known as the superpathway of hexuronide and hexuronate degradation (Supplementary Fig. 5a)), and (2) GLUCUROCAT−PWY, the superpathway of β-d-glucuronide and d-glucuronate degradation. A subset of samples broke this trend, with Escherichia coli appearing as the dominant transcriber of these pathways. E. coli has been previously shown to be able to use β-d-glucuronides and the hexuronates D-glucuronate and D-fructuronate as the sole carbon source for growth. Our data indicate that F. prausnitzii is the main utilizer of these sugars, even in samples where E. coli was present (Supplementary Fig. 5a).
By contrast, the most ubiquitous pathways with the greatest contributional alpha diversities were two biosynthesis pathways for the ribonucleotides adenosine and guanosine (PWY−7219 and PWY−7221), which are involved in numerous basic biochemical processes (including functioning as extracellular signalling molecules). These essential functions were contributed by a multitude of organisms, and generally all organisms that encoded the pathway were also expressing it (Supplementary Figs. 5b and 6a). Adenosine is an important modulator of inflammation with anti-inflammatory effects and therefore a potential therapeutic target in IBD29,30. Furthermore, guanosine can inhibit lipopolysaccharide-induced pro-inflammatory responses in the context of neuroinflammatory-related diseases31.
Many pathways exhibited similar alpha diversity patterns in DNA and RNA, as illustrated by the examples above. However, for a subset of pathways, a lower diversity of contributing species was detected on the RNA level, with transcription often dominated by a single species. The species with the greatest discrepancies in DNA and RNA transcriptional profiles included four Bacteroides species (B. vulgatus, B. uniformis, B. ovatus and B. xylanisolvens), F. prausnitzii, Parabacteroides distasonis and A. putredinis (Fig. 3b). As an example, F. prausnitzii showed the highest degree of variation in DNA–RNA differences and also contributed to the largest number of pathways.
One of the pathways where transcription was dominated by F. prausnitzii was dTDP-l-rhamnose biosynthesis I (Fig. 3c). The resulting deoxysugar β-l-rhamnopyranose functions as a building block of the glycan component of the O-antigens, which are a major target for the immune systems and the target of many vaccine development studies32,33,34,35. Regulation of this biosynthetic pathway has been previously studied in the context of Pseudomonas aeruginosa, in which it is transcriptionally regulated specifically by quorum-sensing systems36. Quorum sensing is a mechanism by which regulation (within or among species) can achieve multi-stability, that is, activate or deactivate expression only after a certain concentration of signalling molecule is achieved. As this type of multi-stability is exhibited by many pathways in the metatranscriptome—that is, only one of several possible organisms upregulated, and others downregulated—we hypothesize that these, and specifically the expression of F. prausnitzii for this pathway, may be regulated through quorum-sensing-like mechanisms, constituting a potential example of inter-species interactions in the gut.
Disease-specific transcriptional microbial signatures
Differences between pathway encoding versus transcription were particularly evident in the pathway contributions of A. putredinis and B. vulgatus, and these transcriptional effects were also associated with disease (Fig. 4). A. putredinis exhibited the highest discrepancy between functional potential and functional activity across all of its pathways (Fig. 3b). One example is the methylerythritol phosphate (MEP) pathway, which was consistently overtranscribed by A. putredinis (Fig. 4a). The product of this pathway, isopentenyl diphosphate (IPP), is used by organisms in the biosynthesis of terpenes and terpenoids, a group of natural products that have been increasingly mined for drug discovery, in particular, for cancer. In E. coli, the MEP pathway is also involved in the production of phosphate-containing antigens recognized by human γδ T lymphocytes, which are suggested to have an important role in the immune response to microorganisms37. Furthermore, IPP can be converted to the more-reactive electrophile dimethylallyl pyrophosphate (DMAPP), which has been shown to induce acute inflammation38.
a, Metagenomic (DNA) and metatranscriptomic (RNA) analysis showed relative contribution of organisms to MEP pathway I (NONMEVIPP–PWY), including all time points and individuals (n = 78). Overtranscription is visible for A. putredinis in the absence of metagenomic disruption. b, Relative contribution of organisms to the MEP pathway, as in a, restricted to the samples from one patient with CD (M2021) over time (days since first sample collection). A. putredinis consistently overtranscribed this pathway in this individual. Increases in the HBI of this patient were observed at time points 2 and 4 (30 and 70 days after the first sample collection, respectively), uniquely where pathway transcription was partially contributed by B. vulgatus. c, Differences in per-pathway DNA versus RNA contributions of microbial species across disease phenotypes, each point representing the perimeter of the disease-specific pathway triangles described in Fig. 2d,e. The mean, first and third quartiles are displayed. Only pathways detected in at least 20% of metagenomic and metatranscriptomic samples (n = 78) were included for a particular species, and only species contributing to at least 8 pathways across samples are displayed. On average, B. vulgatus exhibited the largest disease-specific difference. d, Changes in the relative contribution of B. vulgatus to its transcribed pathways stratified by IBD phenotype. For this organism, IBD-specific dysbioses were particularly pronounced at the transcriptional level, with many pathways upregulated both in UC and in CD.
Finally, disease-associated transcriptional effects became evident when examining patient time courses. For one patient with CD, for example (Fig. 4b), we observed fairly constant proportions of all microbial species contributing the MEP pathway at the DNA level. However, among RNA data, A. putredinis monopolized MEP pathway transcription, and it was strikingly the sole contributor at time points 1 and 3. At time points 2 and 4, B. vulgatus contributed transcriptional activity for the MEP pathway. Interestingly, this coincided with changes in disease severity for this patient, with HBI scores increased at both time points in which B. vulgatus was a pathway contributor (Supplementary Fig. 3b). Both species exhibited an overall correlation with disease severity, with A. putredinis negatively correlated with disease severity and B. vulgatus positively correlated (Supplementary Fig. 6b,c). We hypothesize that this is probably a sufficient but not necessary covariation with disease. Indeed, this correlation was not induced on a population level. This example highlights the importance of examining changes in metatranscription over time within individual patients; such changes may correlate with variation in disease severity, thus suggesting a mechanistic relationship that would be masked in DNA data alone.
We further examined disease-associated transcriptional changes that generalized across patients with IBD. The most pronounced IBD-specific transcriptional changes were observed for B. vulgatus (Fig. 4c). More specifically, most B. vulgatus pathways were more DNA abundant in non-IBD patients, but many of these pathways were considerably more RNA abundant among patients with UC and CD (Fig. 4d). This suggests that B. vulgatus follows a different transcriptional programme in patients with IBD, possibly triggered by disease-specific environmental changes in the gut (for example, inflammation or increased levels of oxygen).
Discussion
Our findings highlight that directly measuring functional activity of the gut microbiome through metatranscriptomics reveals important insights that are only partially observable in metagenomic functional potential, including disease-linked observations. For some pathways, a dominant transcribing organism was identified in a background of mixed metagenomic contributions. Several striking examples of this phenomenon involved, for example, F. prausnitzii, B. vulgatus or A. putredinis, which often dominated pathway transcription in IBD even when they were not the most abundant organism in a sample. Furthermore, several species displayed altered expression profiles in IBD.
Importantly, many IBD-specific signals were either more pronounced or only detectable on the RNA level, such as pathways that were substantially upregulated in disease and species that displayed altered expression profiles in patients with IBD. These altered expression profiles are potentially the result of changes in the gut environment in patients with IBD, which include increased levels of inflammation (resulting in an aggravated immune response), higher concentrations of oxygen (which may be toxic to obligate anaerobes) and a diminished mucous layer39. Metatranscriptomics circumvents the challenges of assaying diverse biochemical products dynamically in situ (for example, mucus40,41 or oxygen6,42) and enables us to study the effects of environmental changes on microbial expression patterns in vivo for large human populations.
In addition to the direct benefits of measuring community functional activity, coupling such measurements with longitudinal sampling enables association of modulated activity with disease progression. This longitudinal design resulted in most samples corresponding to a minority of patients, thus not appropriate for most cross-sectional analyses. For example, our data highlight cases in which microbial genomic contributions to a particular pathway remained stable over time, whereas the corresponding expression patterns varied with disease severity. Thus, microbial dysbioses affecting disease progression and severity may be mechanistically related to changes in the transcriptional programmes of an otherwise stable community, thus making metatranscriptomic profiling an important tool for understanding such mechanisms. Although both RNA and DNA abundances can change in microbial communities, they of course do so at very different time scales (minutes or less, versus hours or more). This underlying biological difference represents another way in which the measurement types may capture complementary microbial processes as they relate to host phenotypes, such as disease flares or changes in inflammation.
We hypothesize that behaviour such as that of F. prausnitzii involves multi-species bistability (or more accurately multi-stability), in which inter-microbial interactions converge on a single dominant transcriber for some functions that can differ between individuals. This type of behaviour in microbial communities is best known from quorum sensing, which itself has been mostly studied in the context of biofilm formation and pathogenic bacteria. For example, the quorum sensor gene lasR of P. aeruginosa (an opportunistic pathogen) produces the molecule N-3-oxo-dodecanoyl-l-homoserine lactone (C12), which allows a microbial subpopulation to bistably activate (or deactivate) regulation after reaching a critical threshold43,44. Other examples include Staphylococcus epidermis, a bacterium that uses quorum sensing to evade human innate immune defence mechanisms45. Furthermore, quorum-sensing molecules have also been shown to affect gut microbial community composition in mice, where increased levels of the quorum-sensing signal autoinducer-2 favoured an expansion of Firmicutes following antibiotic treatment46. As many transcriptional systems in the human gut seem to be regulated in a manner that is multi-stable among microorganisms and individuals, it remains to be determined whether formal quorum-sensing molecules or other regulatory mechanisms are responsible, particularly in the context of IBD.
Some technical limitations apply to RNA-based measurements in stool. Faecal metatranscriptomics captures RNA that is extractable, not degraded during the extraction procedure or in the cells beforehand, and restricted to the organisms that are present in stool samples. Although this is a subset of total biological regulatory activity, the same kind of caveats and technical limitations apply to any kind of RNA-based measurements of transcriptional regulation in other systems. Some of these technical limitations also apply to faecal metagenomics. Although biopsies may be more representative of microbial abundance and expression at the colonic mucosa, frequent longitudinal sampling is implausible owing to the invasive nature of this procedure, and extracting sufficient amounts of bacterial nucleotides for metagenomics or metatranscriptomics is challenging owing to the predominance of host tissue. Differences may also arise due to variation in transit time among subjects. Furthermore, in this and most studies, samples were processed uniformly, ensuring that the same technical limitations apply to all phenotypes and that disease-specific differential expression is likely to reflect underlying biological differences.
In conclusion, metagenomics and metatranscriptomics can provide complementary insights into community interactions and disease-specific alterations in population-scale human microbiomes, here, demonstrated in the IBD gut microbial community. In particular, disease-related changes in the gut environment may specifically affect microbial expression patterns, in different organisms and pathways among individuals, and in some cases without altering metagenomically measured functional potential. To understand the underlying mechanisms associating microbial dysbiosis with aberrant immune responses, we need to understand how the behaviour of individual organisms, as well as the gut community as a whole, changes in disease. Furthermore, disease-specific changes may be patient-specific and the specific microorganisms in a patient’s gut may react differently to environmental changes, resulting in different short-term expression dynamics. Thus, longitudinal, multi’omic, patient-focused studies will provide an important step towards understanding microbiome-related diseases and their roles in personalized medicine.
Methods
Experimental model and subject details
Human cohort
Patients at Massachusetts General Hospital (as part of the Prospective Registry in IBD Study (PRISM)), Emory University and Cincinnati Children’s Hospital Medical Center, with a suspected diagnosis of IBD were approached for participation in the new-onset and paediatric portion of the study. Patients were consented prior to a screening colonoscopy, which separated them into patients with confirmed IBD and non-IBD controls. Sampling and data gathering began at a later 'baseline' visit no more than 6 months after their diagnosis was confirmed. New-onset patients were excluded if they were on an anti-tumour necrosis factor (TNF) inhibitor. Established disease patients were recruited from the MLI (Mucosal Luminal Interface) cohort at the Cedars-Sinai IBD Center, and were required to have had a diagnosis of IBD for >5 years. Participants in all groups were excluded if they were pregnant, had a known bleeding disorder, had taken antibiotics within the month preceding the screening visit, were actively being treated for a malignancy with chemotherapy, had an acute gastrointestinal infection, were diagnosed with an indeterminate colitis, or had had bowel/intestinal surgery other than an appendectomy or cholecystectomy. Non-IBD controls were further required to have no known immune-mediated disease (rheumatoid arthritis, lupus or type 1 diabetes mellitus).
In total, 117 patients participated in the study: 59 patients with CD, 34 patients with UC and 24 non-IBD controls. This included 55 paediatric patients (≤17 years of age; 13 non-IBD patients) with new-onset disease (13 UC and 29 CD), and 62 adults (≥18 years of age; 11 non-IBD patients), who were divided into new-onset (9 UC and 13 CD) and those with established disease (12 UC and 17 CD). Gender was balanced across all cohorts, with 57 male and 60 female patients overall, and no more than a difference of 2 patients between genders for any disease type. Stool samples were self-collected biweekly for 1 year from each patient according to the protocol established in ref.47, starting from the baseline visit. Disease severity was monitored using the HBI48 for patients with CD and the Simple Clinical Colitis Activity Index49 (SCCAI) for patients with UC.
Ethics statement
Subject recruitment and study procedures were approved by and carried out in accordance with the Research Ethics Boards of Massachusetts General Hospital (institutional review board (IRB) for adult cohort: 2013P002215, and IRB for paediatric cohort: 2014P001115), Cincinnati Children’s Hospital Medical Center (IRB: 2013-7586), Emory University (IRB: IRB00071468), and Cedars-Sinai Medical Center (IRB: 3358). In compliance with the Research Ethics Board study approval, informed consent was obtained from all study participants immediately before the initial sample collection. Furthermore, all experimental methods are compliant with the Helsinki Declaration.
Shotgun sequencing
For metagenomic sequencing, the total genomic DNA content of the sample was sequenced, allowing us to infer the functional potential of the community and taxonomic composition at the species level. For metatranscriptomics, messenger RNA was extracted, reverse transcribed into complementary DNA and subsequently sequenced. DNA was extracted from 300 samples spanning all 117 participants, and RNA from a subset of 78 samples spanning 28 participants. Illumina HiSeq sequencing yielded a total of 4.59 giga nucleotides (Gnt) and 1.06 Gnt of paired-end reads (2 × 100 nt) of metagenomic and metatranscriptomic sequencing, respectively. Metagenomes averaged 30,581,993 reads ± 12,567,915 reads (mean ± s.d.) per sample before quality filtering (see below) and 28,242,423 reads ± 12,437,200 reads afterward. Meanwhile, metatranscriptomes averaged 27,211,997 reads ± 21,831,783 reads and 20,050,758 reads ± 16,301,242 reads before and after quality control, respectively.
Quantification and statistical analysis
Preprocessing and quality control
Sequence reads were processed with the KneadData v0.5.1 quality control pipeline (http://huttenhower.sph.harvard.edu/kneaddata), which uses the Trimmomatic50 and BMTagger51 filtering and decontamination algorithms to remove low-quality read bases and host (human) reads, respectively. Trimmomatic was run with parameters MAXINFO:80:0.5, and Phred quality scores were thresholded at <20. Trimmed non-human reads shorter than 50 nt were discarded. Potential human contamination was filtered by removing reads that aligned to the human genome (reference genome hg19). In addition, metatranscriptomic reads were filtered against the human transcriptome and the SILVA database52. After quality control, samples averaged 28 million and 20 million reads in metagenomic and metatranscriptomic data, respectively (minimum 2 million).
Taxonomic and functional profiling
Taxonomic profiling was performed using the MetaPhlAn2 classifier53, which unambiguously classifies metagenomic reads to taxonomies based on a database of clade-specific marker genes derived from 17,000 microbial genomes (corresponding to >7,500 bacterial, viral, archaeal and eukaryotic species). Functional profiling of metagenomes and metatranscriptomes was performed using HUMAnN2 (ref. 24) version 0.9.6 (http://huttenhower.sph.harvard.edu/humann2). Briefly, the MetaPhlAn2 taxonomic profile generated from a metagenome is used to identify the set of organisms present in a sample. Metagenomic and metatranscriptomic reads are then mapped using Bowtie2 (ref. 54) to sample-specific pangenomes, including all gene families in any microorganism present. A translated search using DIAMOND55 then maps unmapped reads against UniRef90 (ref. 56). Hits are counted per gene family and normalized for length and alignment quality. Gene family abundances from both the nucleotide and the translated searches are then combined into structured pathways from MetaCyc57 and sum-normalized to relative abundances. We ran HUMAnN2 with the MinPath58 and gap filling options. As a result, 385 pathways had non-zero abundance in at least 1 metagenome, and 331 pathways had non-zero abundance in at least 1 metatranscriptome. The nucleotide search identified 182 species contributing these pathways in metagenomes, and 134 species in metatranscriptomes (a subset of the metagenomic species).
All of our data sets consisted of at least 2 million reads (corresponding to at least 20 observed reads per species), and the majority of them were in excess of 10 million reads (corresponding to at least 100 observed reads per species), ensuring that species calls were well supported (Supplementary Fig. 2e).
Measuring activity of microbial species
For Figs. 2a,d,e, 4d and Supplementary Fig. 4, the total contribution of each species to the functional profile was computed by summing their contributions over all pathways. Only the 51 species that contributed to at least one pathway at the DNA and RNA level in >10% of samples (8 samples) were considered. Species contributions were first averaged over all samples where the species was detected within each patient and then across patients. For Fig. 2b,c, for all species, we computed Spearman correlation coefficients between their total pathway contributions to the metagenomes and metatranscriptomes across all samples.
Sample order in stacked bar plots
Stacked bar plots presented in Figs. 3 and 4, and Supplementary Figs. 5 and 6 were ordered to maximize the similarity of species contributions to the pathway’s abundance between adjacent samples. For this, we calculated Bray–Curtis dissimilarities between the sum-normalized species contributions to the pathway in a given sample for both the metatranscriptomic and metagenomic data. The two dissimilarity matrices were combined by a weighted mixture, with metagenomic dissimilarities weighted at 1/100th of metatranscriptomic dissimilarities. The final sample order was determined by running solve_TSP from the R package TSP on the mixture dissimilarity matrix. To determine which sample to place first, we included a 'fence' sample with zero dissimilarity to all other samples in the above procedure. The fence sample was then placed in the first position by rotating the final sample order appropriately, before finally removing it.
Contributional alpha diversity of pathways
We quantified the contributional alpha diversity of species contributing to the abundance of a MetaCyc metabolic pathway in DNA or RNA by the Gini–Simpson index of alpha diversity. Pathways were first excluded if they had non-zero abundance in DNA in <95% of the samples, or if >25% of the pathway was attributed to unclassified organisms (from HUMAnN2’s translated search) in >25% of the samples. For each of the 58 remaining pathways, we then computed the Gini–Simpson index from the relative contribution of each species to the pathway for each sample (excluding unclassified organisms). The pathway’s alpha diversity was then defined as the mean alpha diversity of samples with non-zero abundance.
Life Sciences Reporting Summary
Further information on experimental design is available in the Life Sciences Reporting Summary.
Data availability
All sequencing data and metadata are available on www.ibdmdb.org. The metagenomic and metatranscriptomic sequencing data are also available on the Sequence Read Archive (BioProject: PRJNA389280).
References
Burisch, J., Jess, T., Martinato, M. & Lakatos, P. L. The burden of inflammatory bowel disease in Europe. J. Crohns Colitis 7, 322–337 (2013).
IBD Working Group of the European Society for Paediatric Gastroenterology, Hepatology and Nutrition. Inflammatory bowel disease in children and adolescents: recommendations for diagnosis—the Porto criteria. J. Pediatr. Gastr. Nutr. 41, 1–7 (2005).
Kaplan, G. G. The global burden of IBD: from 2015 to 2025. Nat. Rev. Gastro. Hepat. 12, 720–727 (2015).
Fava, F. & Danese, S. Intestinal microbiota in inflammatory bowel disease: friend of foe? World J. Gastroentero. 17, 557–566 (2011).
Hold, G. L. et al. Role of the gut microbiota in inflammatory bowel disease pathogenesis: what have we learnt in the past 10 years? World J. Gastroentero. 20, 1192–1210 (2014).
Morgan, X. C. et al. Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome Biol. 13, R79 (2012).
Gevers, D. et al. The treatment-naive microbiome in new-onset Crohn’s disease. Cell Host Microbe 15, 382–392 (2014).
Lupp, C. et al. Host-mediated inflammation disrupts the intestinal microbiota and promotes the overgrowth of Enterobacteriaceae. Cell Host Microbe 2, 119–129 (2007).
Frank, D. N. et al. Disease phenotype and genotype are associated with shifts in intestinal-associated microbiota in inflammatory bowel diseases. Inflamm. Bowel Dis. 17, 179–184 (2011).
Kostic, A. D., Xavier, R. J. & Gevers, D. The microbiome in inflammatory bowel disease: current status and the future ahead. Gastroenterology 146, 1489–1499 (2014).
Llopis, M. et al. Lactobacillus casei downregulates commensals’ inflammatory signals in Crohn’s disease mucosa. Inflamm. Bowel Dis. 15, 275–283 (2009).
Sokol, H. et al. Low counts of Faecalibacterium prausnitzii in colitis microbiota. Inflamm. Bowel Dis. 15, 1183–1189 (2009).
Sokol, H. et al. Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of Crohn disease patients. Proc. Natl Acad. Sci. USA 105, 16731–16736 (2008).
Halfvarson, J. et al. Dynamics of the human gut microbiome in inflammatory bowel disease. Nat. Microbiol. 2, 17004 (2017).
Lewis, J. D. et al. Inflammation, antibiotics, and diet as environmental stressors of the gut microbiome in pediatric Crohn’s disease. Cell Host Microbe 18, 489–500 (2015).
Zhernakova, A. et al. Population-based metagenomics analysis reveals markers for gut microbiome composition and diversity. Science 352, 565–569 (2016).
Machiels, K. et al. A decrease of the butyrate-producing species Roseburia hominis and Faecalibacterium prausnitzii defines dysbiosis in patients with ulcerative colitis. Gut 63, 1275–1283 (2014).
Nagao-Kitamoto, H. & Kamada, N. Host–microbial cross-talk in inflammatory bowel disease. Immune Netw. 17, 1–12 (2017).
Rehman, A. et al. Transcriptional activity of the dominant gut mucosal microbiota in chronic inflammatory bowel disease patients. J. Med. Microbiol. 59, 1114–1122 (2010).
Dorffel, Y., Swidsinski, A., Loening-Baucke, V., Wiedenmann, B. & Pavel, M. Common biostructure of the colonic microbiota in neuroendocrine tumors and Crohn’s disease and the effect of therapy. Inflamm. Bowel Dis. 18, 1663–1671 (2012).
Bloom, S. M. et al. Commensal Bacteroides species induce colitis in host-genotype-specific fashion in a mouse model of inflammatory bowel disease. Cell Host Microbe 9, 390–403 (2011).
Joossens, M. et al. Dysbiosis of the faecal microbiota in patients with Crohn’s disease and their unaffected relatives. Gut 60, 631–637 (2011).
Hoffmann, T. W. et al. Microorganisms linked to inflammatory bowel disease-associated dysbiosis differentially impact host physiology in gnotobiotic mice. ISME J. 10, 460–477 (2016).
Abubucker, S. et al. Metabolic reconstruction for metagenomic data and its application to the human microbiome. PLoS Comput. Biol. 8, e1002358 (2012).
Feng, Q. et al. Gut microbiome development along the colorectal adenoma-carcinoma sequence. Nat. Commun. 6, 6528 (2015).
Zitomersky, N. L. et al. Characterization of adherent bacteroidales from intestinal biopsies of children and young adults with inflammatory bowel disease. PLoS ONE 8, e63686 (2013).
Downes, J., Munson, M. & Wade, W. G. Dialister invisus sp. nov., isolated from the human oral cavity. Int. J. Syst. Evol. Micr. 53, 1937–1940 (2003).
Maffeis, C. et al. Association between intestinal permeability and faecal microbiota composition in Italian children with beta cell autoimmunity at risk for type 1 diabetes. Diabetes Metab. Res. Rev. 32, 700–709 (2016).
Ye, J. H. & Rajendran, V. M. Adenosine: an immune modulator of inflammatory bowel diseases. World J. Gastroentero. 15, 4491–4498 (2009).
Antonioli, L., Blandizzi, C., Pacher, P. & Hasko, G. Immunity, inflammation and cancer: a leading role for adenosine. Nat. Rev. Cancer 13, 842–857 (2013).
Bellaver, B. et al. Guanosine inhibits LPS-induced pro-inflammatory response and oxidative stress in hippocampal astrocytes through the heme oxygenase-1 pathway. Purinergic Signal. 11, 571–580 (2015).
Pier, G. B. Pseudomonas aeruginosa lipopolysaccharide: a major virulence factor, initiator of inflammation and target for effective immunity. Int. J. Med. Microbiol. 297, 277–295 (2007).
Santos, M. F. et al. Lipopolysaccharide as an antigen target for the formulation of a universal vaccine against Escherichia coli O111 strains. Clin. Vaccine Immunol. 17, 1772–1780 (2010).
Wang, L., Wang, Q. & Reeves, P. R. The variation of O antigens in Gram-negative bacteria. Subcell. Biochem. 53, 123–152 (2010).
Kintz, E. et al. Salmonella enterica Serovar Typhi lipopolysaccharide O-antigen modification impact on serum resistance and antibody recognition. Infect. Immun. 85, e01021-16 (2017).
Aguirre-Ramirez, M., Medina, G., Gonzalez-Valdez, A., Grosso-Becerra, V. & Soberon-Chavez, G. The Pseudomonas aeruginosa rmlBDAC operon, encoding dTDP-l-rhamnose biosynthetic enzymes, is regulated by the quorum-sensing transcriptional regulator RhlR and the alternative sigma factor σS. Microbiology 158, 908–916 (2012).
Feurle, J. et al. Escherichia coli produces phosphoantigens activating human γδ T cells. J. Biol. Chem. 277, 148–154 (2002).
Bang, S., Yoo, S., Yang, T. J., Cho, H. & Hwang, S. W. Nociceptive and pro-inflammatory effects of dimethylallyl pyrophosphate via TRPV4 activation. Br. J. Pharmacol. 166, 1433–1443 (2012).
Naughton, J., Duggan, G., Bourke, B. & Clyne, M. Interaction of microbes with mucus and mucins: recent developments. Gut Microbes 5, 48–52 (2014).
Skoog, E. C. et al. Human gastric mucins differently regulate Helicobacter pylori proliferation, gene expression and interactions with host cells. PLoS ONE 7, e36378 (2012).
Tu, Q. V., McGuckin, M. A. & Mendz, G. L. Campylobacter jejuni response to human mucin MUC2: modulation of colonization and pathogenicity determinants. J. Med. Microbiol. 57, 795–802 (2008).
Albenberg, L. et al. Correlation between intraluminal oxygen gradient and radial partitioning of intestinal microbiota. Gastroenterology 147, 1055–1063 (2014).
LaFayette, S. L. et al. Cystic fibrosis-adapted Pseudomonas aeruginosa quorum sensing lasR mutants cause hyperinflammatory responses. Sci. Adv. 1, e1500199 (2015).
Glucksam-Galnoy, Y. et al. The bacterial quorum-sensing signal molecule N-3-oxo-dodecanoyl-l-homoserine lactone reciprocally modulates pro- and anti-inflammatory cytokines in activated macrophages. J. Immunol. 191, 337–344 (2013).
Yao, Y. et al. Characterization of the Staphylococcus epidermidis accessory-gene regulator response: quorum-sensing regulation of resistance to human innate host defense. J. Infect. Dis. 193, 841–848 (2006).
Thompson, J. A., Oliveira, R. A., Djukovic, A., Ubeda, C. & Xavier, K. B. Manipulation of the quorum sensing signal AI-2 affects the antibiotic-treated gut microbiota. Cell Rep. 10, 1861–1871 (2015).
Franzosa, E. A. et al. Relating the metatranscriptome and metagenome of the human gut. Proc. Natl Acad. Sci. USA 111, E2329–E2338 (2014).
Harvey, R. F. & Bradshaw, J. M. A simple index of Crohn’s-disease activity. Lancet 1, 514 (1980).
Walmsley, R. S., Ayres, R. C., Pounder, R. E. & Allan, R. N. A simple clinical colitis activity index. Gut 43, 29–32 (1998).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Human Microbiome Project Consortium. A framework for human microbiome research. Nature 486, 215–221 (2012).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Truong, D. T. et al. MetaPhlAn2 for enhanced metagenomic taxonomic profiling. Nat. Methods 12, 902–903 (2015).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2015).
Suzek, B. E., Huang, H., McGarvey, P., Mazumder, R. & Wu, C. H. UniRef: comprehensive and non-redundant UniProt reference clusters. Bioinformatics 23, 1282–1288 (2007).
Caspi, R. et al. The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of pathway/genome databases. Nucleic Acids Res. 40, D742–D753 (2012).
Ye, Y. & Doak, T. G. A parsimony approach to biological pathway reconstruction/inference for genomes and metagenomes. PLoS Comput. Biol. 5, e1000465 (2009).
Acknowledgements
We thank the participants from Massachusetts General Hospital, Emory University, Cedars-Sinai IBD Center and Cincinnati Children’s Hospital Medical Center, who made this study possible. Furthermore, we acknowledge B. Sayoldin for making the data available through the Sequence Read Archive and our collaborators throughout the Integrative Human Microbiome Consortium. This work was supported by the US National Institutes of Health (NIH) grants U54DK102557 (C.H. and R.J.X.), STARR Cancer Consortium (C.H.), CCFA 20144126 (R.J.X.) and R01DK92405 (R.J.X.), U01DK062413 (D.P.B.M.), P01DK046763 (D.P.B.M.), UL1TR001881 (J.B.), and The Leona M. and Harry B. Helmsley Charitable Trust (D.P.B.M.).
Author information
Authors and Affiliations
Contributions
M.S., C.H., R.J.X. and H.V. conceived and designed the experiments. A.N.A., E.A., G.B., K.L., M.P., J.S., B.S. and R.G.W. performed the experiments. M.S., C.H. and E.A.F. analysed the data. M.S., C.H., E.A.F., J.L.-P., L.J.M., R.S., T.W.P., E.A., J.B., L.A.D., S.K. and D.P.B.M. contributed materials/analysis tools. M.S., C.H., R.J.X., J.L.-P. and H.V. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
D.P.B.M. is consulting for Cidara. The authors declare no other competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–6, Supplementary Table 1.
Supplementary Table 2
Description of pathways.
Rights and permissions
About this article
Cite this article
Schirmer, M., Franzosa, E.A., Lloyd-Price, J. et al. Dynamics of metatranscription in the inflammatory bowel disease gut microbiome. Nat Microbiol 3, 337–346 (2018). https://doi.org/10.1038/s41564-017-0089-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-017-0089-z
This article is cited by
-
Microbiota-derived indoles alleviate intestinal inflammation and modulate microbiome by microbial cross-feeding
Microbiome (2024)
-
One-pot method for preparing DNA, RNA, and protein for multiomics analysis
Communications Biology (2024)
-
Dubosiella newyorkensis modulates immune tolerance in colitis via the L-lysine-activated AhR-IDO1-Kyn pathway
Nature Communications (2024)
-
Comparison of the effectiveness of different normalization methods for metagenomic cross-study phenotype prediction under heterogeneity
Scientific Reports (2024)
-
Resistome expansion in disease-associated human gut microbiomes
Microbiome (2023)