Abstract
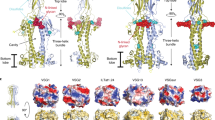
The most prominent defence of the unicellular parasite Trypanosoma brucei against the host immune system is a dense coat that comprises a variant surface glycoprotein (VSG). Despite the importance of the VSG family, no complete structure of a VSG has been reported. Making use of high-resolution structures of individual VSG domains, we employed small-angle X-ray scattering to elucidate the first two complete VSG structures. The resulting models imply that the linker regions confer great flexibility between domains, which suggests that VSGs can adopt two main conformations to respond to obstacles and changes of protein density, while maintaining a protective barrier at all times. Single-molecule diffusion measurements of VSG in supported lipid bilayers substantiate this possibility, as two freely diffusing populations could be detected. This translates into a highly flexible overall topology of the surface VSG coat, which displays both lateral movement in the plane of the membrane and variation in the overall thickness of the coat.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Ziegelbauer, K. & Overath, P. Identification of invariant surface glycoproteins in the bloodstream stage of Trypanosoma brucei. J. Biol. Chem. 267, 10791–10796 (1992).
Grünfelder, C. G. et al. Accumulation of a GPI-anchored protein at the cell surface requires sorting at multiple intracellular levels. Traffic 3, 547–559 (2002).
Cross, G. A. M. Identification, purification and properties of clone-specific glycoprotein antigens constituting the surface coat of Trypanosoma brucei. Parasitology 71, 393–417 (1975).
Cardoso de Almeida, M. L. & Turner, M. J. The membrane form of variant surface glycoproteins of Trypanosoma brucei. Nature 302, 349–352 (1983).
Ziegelbauer, K. & Overath, P. Organization of two invariant surface glycoproteins in the surface coat of Trypanosoma brucei. Infect. Immun. 61, 4540–4545 (1993).
Sullivan, L., Wall, S. J., Carrington, M. & Ferguson, M. A. J. Proteomic selection of immunodiagnostic antigens for human African trypanosomiasis and generation of a prototype lateral flow immunodiagnostic device. PLoS Negl. Trop. Dis. 7, e2087 (2013).
Macaskill, J. A., Holmes, P. H., Jennings, F. W. & Urquhart, G. M. Immunological clearance of 75Se-labelled Trypanosoma brucei in mice. III. Studies in animals with acute infections. Immunology 43, 691–698 (1981).
McLintock, L. M., Turner, C. M. & Vickerman, K. Comparison of the effects of immune killing mechanisms on Trypanosoma brucei parasites of slender and stumpy morphology. Parasite Immunol. 15, 475–480 (1993).
Cross, G. A. M., Kim, H.-S. & Wickstead, B. Capturing the variant surface glycoprotein repertoire (the VSGnome) of Trypanosoma brucei Lister 427. Mol. Biochem. Parasitol. 195, 59–73 (2014).
Engstler, M. et al. Hydrodynamic flow-mediated protein sorting on the cell surface of trypanosomes. Cell 131, 505–515 (2007).
Freymann, D. M., Metcalf, P., Turner, M. & Wiley, D. C. 6 Å-resolution X-ray structure of a variable surface glycoprotein from Trypanosoma brucei. Nature 311, 167–169 (1984).
Metcalf, P., Down, J. A., Turner, M. J. & Wiley, D. C. Crystallization of amino-terminal domains and domain fragments of variant surface glycoproteins from Trypanosoma brucei brucei. J. Biol. Chem. 263, 17030–17033 (1988).
Carrington, M. et al. Variant specific glycoprotein of Trypanosoma brucei consists of two domains each having an independently conserved pattern of cysteine residues. J. Mol. Biol. 221, 823–835 (1991).
Marcello, L. & Barry, J. D. Analysis of the VSG gene silent archive in Trypanosoma brucei reveals that mosaic gene expression is prominent in antigenic variation and is favored by archive substructure. Genome Res. 17, 1344–1352 (2007).
Freymann, D. et al. 2.9 Å resolution structure of the N-terminal domain of a variant surface glycoprotein from Trypanosoma brucei. J. Mol. Biol. 216, 141–160 (1990).
Blum, M. L. et al. A structural motif in the variant surface glycoproteins of Trypanosoma brucei. Nature 362, 603–609 (1993).
Berriman, M. et al. The genome of the African trypanosome Trypanosoma brucei. Science 309, 416–422 (2005).
Chattopadhyay, A. et al. Structure of the C-terminal domain from Trypanosoma brucei variant surface glycoprotein MITat1.2. J. Biol. Chem. 280, 7228–7235 (2005).
Jones, N. G. et al. Structure of a glycosylphosphatidylinositol-anchored domain from a trypanosome variant surface glycoprotein. J. Biol. Chem. 283, 3584–3593 (2008).
Allen, G. & Gurnett, L. Locations of the six disulphide bonds in a variant surface glycoprotein (VSG 117) from Trypanosoma brucei. Biochem. J. 209, 481–487 (1983).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Zamze, S. E. et al. Characterisation of the asparagine-linked oligosaccharides from Trypanosoma brucei type-I variant surface glycoproteins. Eur. J. Biochem. 187, 657–663 (1990).
Bangs, J., Doering, T., Englund, P. & Hart, G. Biosynthesis of a variant surface glycoprotein of Trypanosoma brucei. Processing of the glycolipid membrane anchor and N-linked oligosaccharides. J. Biol. Chem. 263, 17697–17705 (1988).
Strang, A. M., Allen, A. K., Holder, A. A. & van Halbeek, H. The carbohydrate structures of Trypanosoma brucei brucei MITat 1.6 variant surface glycoprotein. A re-investigation of the C-terminal glycan. Biochem. Biophys. Res. Commun. 196, 1430–1439 (1993).
Jackson, D. G., Owen, M. J. & Voorheis, H. P. A new method for the rapid purification of both the membrane-bound and released forms of the variant surface glycoprotein from Trypanosoma brucei. Biochem. J. 230, 195–202 (1985).
Grünfelder, C. G. et al. Endocytosis of a glycosylphosphatidylinositol-anchored protein via clathrin-coated vesicles, sorting by default in endosomes, and exocytosis via RAB11-positive carriers. Mol. Biol. Cell 14, 2029–2040 (2003).
Overath, P. & Engstler, M. Endocytosis, membrane recycling and sorting of GPI-anchored proteins: Trypanosoma brucei as a model system. Mol. Microbiol. 53, 735–744 (2004).
Hartel, A. J. W. et al. The molecular size of the extra-membrane domain influences the diffusion of the GPI-anchored VSG on the trypanosome plasma membrane. Sci. Rep. 5, 10394 (2015).
Hartel, A. J. W. et al. N-glycosylation enables high lateral mobility of GPI-anchored proteins at a molecular crowding threshold. Nat. Commun. 7, 12870 (2016).
Batram, C., Jones, N. G., Janzen, C. J., Markert, S. M. & Engstler, M. Expression site attenuation mechanistically links antigenic variation and development in Trypanosoma brucei. eLife 3, e02324 (2014).
Salmon, D. et al. Characterization of the ligand-binding site of the transferrin receptor in Trypanosoma brucei demonstrates a structural relationship with the N-terminal domain of the variant surface glycoprotein. EMBO J. 16, 7272–7278 (1997).
Lane-Serff, H. et al. Structural basis for ligand and innate immunity factor uptake by the trypanosome haptoglobin–haemoglobin receptor. eLife 3, e05553 (2014).
Bargul, J. L. et al. Species-specific adaptations of trypanosome morphology and motility to the mammalian host. PLoS Pathog. 12, e1005448 (2016).
Muñoz-Jordán, J. L., Davies, K. P. & Cross, G. A. Stable expression of mosaic coats of variant surface glycoproteins in Trypanosoma brucei. Science 272, 1795–1797 (1996).
Böhme, U. & Cross, G. A. M. Mutational analysis of the variant surface glycoprotein GPI-anchor signal sequence in Trypanosoma brucei. J. Cell Sci. 115, 805–816 (2002).
Cross, G. A. M. Release and purification of Trypanosoma brucei variant surface glycoprotein. J. Cell. Biochem. 24, 79–90 (1984).
Ferguson, M. A., Haldar, K. & Cross, G. A. Trypanosoma brucei variant surface glycoprotein has a sn-1,2-dimyristyl glycerol membrane anchor at its COOH terminus. J. Biol. Chem. 260, 4963–4968 (1985).
Hirumi, H. & Hirumi, K. Continuous cultivation of Trypanosoma brucei blood stream forms in a medium containing a low concentration of serum protein without feeder cell layers. J. Parasitol. 75, 985–989 (1989).
Gerlach, M., Mueller, U. & Weiss, M. S. The MX Beamlines BL14.1-3 at BESSY II. J. Large-scale Res. Fac. 2, A47 (2016).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011).
Battye, T. G. G., Kontogiannis, L., Johnson, O., Powell, H. R. & Leslie, A. G. W. iMOSFLM: a new graphical interface for diffraction-image processing with MOSFLM. Acta Crystallogr. D 67, 271–281 (2011).
Evans, P. Scaling and assessment of data quality. Acta Crystallogr. D 62, 72–82 (2006).
Evans, P. R. An introduction to data reduction: space-group determination, scaling and intensity statistics. Acta Crystallogr. D 67, 282–292 (2011).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Langer, G., Cohen, S. X., Lamzin, V. S. & Perrakis, A. Automated macromolecular model building for X-ray crystallography using ARP/wARP version 7. Nat. Protoc. 3, 1171–1179 (2008).
Cowtan, K. The Buccaneer software for automated model building. 1. Tracing protein chains. Acta Crystallogr. D 62, 1002–1011 (2006).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997).
Needleman, S. B. & Wunsch, C. D. A general method applicable to the search for similarities in the amino acid sequence of two proteins. J. Mol. Biol. 48, 443–453 (1970).
Henikoff, S. & Henikoff, J. G. Amino acid substitution matrices from protein blocks. Proc. Natl Acad. Sci. USA 89, 10915–10919 (1992).
Kraulis, P., Domaille, P., Campbell-Burk, S., Van Aken, T. & Laue, E. Solution structure and dynamics of ras p21.GDP determined by heteronuclear three- and four-dimensional NMR spectroscopy. Biochemistry 33, 3515–3531 (1994).
Linge, J., Habeck, M., Rieping, W. & Nilges, M. ARIA: automated NOE assignment and NMR structure calculation. Bioinformatics 19, 315–316 (2003).
Brunger, A. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Brennich, M. E. et al. Online data analysis at the ESRF bioSAXS beamline, BM29. J. Appl. Crystallogr. 49, 203–212 (2016).
Pernot, P. et al. Upgraded ESRF BM29 beamline for SAXS on macromolecules in solution. J. Synchrotron. Rad. 20, 660–664 (2013).
Round, A. R. et al. Automated sample-changing robot for solution scattering experiments at the EMBL Hamburg SAXS station X33. J. Appl. Crystallogr. 41, 913–917 (2008).
Konarev, P. V., Volkov, V. V., Sokolova, A. V., Koch, M. & Svergun, D. I. PRIMUS: a Windows PC-based system for small-angle scattering data analysis. J. Appl. Crystallogr. 36, 1277–1282 (2003).
Petoukhov, M., Konarev, P. V., Kikhney, A. G. & Svergun, D. I. ATSAS 2.1—towards automated and web-supported small-angle scattering data analysis. J. Appl. Crystallogr. 40, S223–S228 (2007).
Svergun, D. I. Determination of the regularization parameter in indirect-transform methods using perceptual criteria. J. Appl. Crystallogr. 25, 495–503 (1992).
Petoukhov, M. et al. New developments in the ATSAS program package for small-angle scattering data analysis. J. Appl. Crystallogr. 45, 342–350 (2012).
Svergun, D. I. Restoring low resolution structure of biological macromolecules from solution scattering using simulated annealing. Biophys. J. 76, 2879–2886 (1999).
Volkov, V. V. & Svergun, D. I. Uniqueness of ab initio shape determination in small-angle scattering. J. Appl. Crystallogr. 36, 860–864 (2003).
Wriggers, W. Using Situs for the integration of multi-resolution structures. Biophys. Rev. 2, 21–27 (2010).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graphics 14, 33–38 (1996).
Svergun, D., Barberato, C. & Koch, M. CRYSOL—a program to evaluate X-ray solution scattering of biological macromolecules from atomic coordinates. J. Appl. Crystallogr. 28, 768–773 (1995).
Petoukhov, M. & Svergun, D. I. Global rigid body modeling of macromolecular complexes against small-angle scattering data. Biophys. J. 89, 1237–1250 (2005).
Schmidt, T., Schütz, G. J., Baumgartner, W., Gruber, H. J. & Schindler, H. Imaging of single molecule diffusion. Proc. Natl Acad. Sci. USA 93, 2926–2929 (1996).
Acknowledgements
This work was supported by the Deutsche Forschungsgemeinschaft (DFG, grants EN 305, GRK 1114 to M.E. and SFB 630 to M.E. and C.K.), and the Wellcome Trust (grant 022758/Z/03/Z to M.Ca.). A.-S.S. and M.Cv. were funded from grant ERC StG 2013-337283 of the European Research Council and supported by the DFG GRK 1962. M.E. is a member of the Wilhelm Conrad Röntgen-Center for Complex Material Systems. We thank the Helmholtz-Zentrum Berlin for the allocation of synchrotron radiation beamtime and the staff of the BESSY at beamline 14.1 for technical assistance. The SAXS experiments were performed on beamline BM29 at ESRF. We are grateful to A. Round at the ESRF for providing assistance in using beamline BM29 and for invaluable tips concerning data analysis. We thank D. Nietlispach for the acquisition of NMR data and B. Morriswood for critical reading of the manuscript.
Author information
Authors and Affiliations
Contributions
T.B., N.G.J. and M.E. conceived the study, T.B., N.G.J., M.Ca., A.-A.S., S.F., C.K. and M.E. designed the research; T.B., N.G.J., M.G. and S.F. performed the experiments; T.B., N.G.J., C.S., M.Cv., M.G., H.R.M., J.K., M.B., A.-S.S., S.F. and M.E. analysed the data; T.B., N.G.J. and M.E. wrote the paper with contributions from M.Cv., A.-S.S. and S.F. during manuscript editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Information
Supplementary Figures 1–9, Supplementary Tables 1–4.
Rights and permissions
About this article
Cite this article
Bartossek, T., Jones, N.G., Schäfer, C. et al. Structural basis for the shielding function of the dynamic trypanosome variant surface glycoprotein coat. Nat Microbiol 2, 1523–1532 (2017). https://doi.org/10.1038/s41564-017-0013-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-017-0013-6
This article is cited by
-
Neutrophil metalloproteinase driven spleen damage hampers infection control of trypanosomiasis
Nature Communications (2023)
-
Cryo-EM structures of Trypanosoma brucei gambiense ISG65 with human complement C3 and C3b and their roles in alternative pathway restriction
Nature Communications (2023)
-
Quantitative proteomic analysis reveals differential modulation of crucial stage specific proteins during promastigote to amastigote differentiation in Leishmania donovani
Journal of Proteins and Proteomics (2022)
-
Structure of trypanosome coat protein VSGsur and function in suramin resistance
Nature Microbiology (2021)
-
A receptor for the complement regulator factor H increases transmission of trypanosomes to tsetse flies
Nature Communications (2020)