Abstract
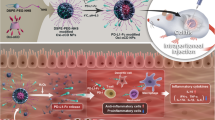
While conventional approaches for inflammatory bowel diseases mainly focus on suppressing hyperactive immune responses, it remains unclear how to address disrupted intestinal barriers, dysbiosis of the gut commensal microbiota and dysregulated mucosal immune responses in inflammatory bowel diseases. Moreover, immunosuppressive agents can cause off-target systemic side effects and complications. Here, we report the development of hyaluronic acid–bilirubin nanomedicine (HABN) that accumulates in inflamed colonic epithelium and restores the epithelium barriers in a murine model of acute colitis. Surprisingly, HABN also modulates the gut microbiota, increasing the overall richness and diversity and markedly augmenting the abundance of Akkermansia muciniphila and Clostridium XIVα, which are microorganisms with crucial roles in gut homeostasis. Importantly, HABN associated with pro-inflammatory macrophages, regulated innate immune responses and exerted potent therapeutic efficacy against colitis. Our work sheds light on the impact of nanotherapeutics on gut homeostasis, microbiome and innate immune responses for the treatment of inflammatory diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the article and its Supplementary Information files. All relevant data can be provided by the authors upon reasonable request.
References
Turner, J. R. Intestinal mucosal barrier function in health and disease. Nat. Rev. Immunol. 9, 799–809 (2009).
Citi, S. Intestinal barriers protect against disease. Science 359, 1097–1098 (2018).
Round, J. L. & Mazmanian, S. K. The gut microbiota shapes intestinal immune responses during health and disease. Nat. Rev. Immunol. 9, 313–323 (2009).
Kostic, A. D., Xavier, R. J. & Gevers, D. The microbiome in inflammatory bowel disease: current status and the future ahead. Gastroenterology 146, 1489–1499 (2014).
Halfvarson, J. et al. Dynamics of the human gut microbiome in inflammatory bowel disease. Nat. Microbiol. 2, 17004 (2017).
Bernstein, C. N. et al. World gastroenterology organization practice guidelines for the diagnosis and management of IBD in 2010. Inflamm. Bowel. Dis. 16, 112–124 (2010).
Lautenschlager, C., Schmidt, C., Fischer, D. & Stallmach, A. Drug delivery strategies in the therapy of inflammatory bowel disease. Adv. Drug Deliv. Rev. 71, 58–76 (2014).
Wilson, D. S. et al. Orally delivered thioketal nanoparticles loaded with TNF-alpha-siRNA target inflammation and inhibit gene expression in the intestines. Nat. Mater. 9, 923–928 (2010).
Zhang, S. et al. An inflammation-targeting hydrogel for local drug delivery in inflammatory bowel disease. Sci. Transl. Med. 7, 300ra128 (2015).
Stallmach, A., Hagel, S. & Bruns, T. Adverse effects of biologics used for treating IBD. Best Pract. Res. Clin. Gastroenterol. 24, 167–182 (2010).
Rayahin, J. E., Buhrman, J. S., Zhang, Y., Koh, T. J. & Gemeinhart, R. A. High and low molecular weight hyaluronic acid differentially influence macrophage activation. ACS Biomater. Sci. Eng. 1, 481–493 (2015).
Hill, D. R., Kessler, S. P., Rho, H. K., Cowman, M. K. & de la Motte, C. A. Specific-sized hyaluronan fragments promote expression of human beta-defensin 2 in intestinal epithelium. J. Biol. Chem. 287, 30610–30624 (2012).
Bollyky, P. L. et al. Intact extracellular matrix and the maintenance of immune tolerance: high molecular weight hyaluronan promotes persistence of induced CD4+CD25+ regulatory T cells. J. Leukoc. Biol. 86, 567–572 (2009).
Zheng, L., Riehl, T. E. & Stenson, W. F. Regulation of colonic epithelial repair in mice by Toll-like receptors and hyaluronic acid. Gastroenterology 137, 2041–2051 (2009).
Xiao, B. et al. Combination therapy for ulcerative colitis: orally targeted nanoparticles prevent mucosal damage and relieve inflammation. Theranostics 6, 2250–2266 (2016).
Petrey, A. C. & de la Motte, C. A. Hyaluronan, a crucial regulator of inflammation. Front. Immunol. 5, 101 (2014).
Kapitulnik, J. Bilirubin: an endogenous product of heme degradation with both cytotoxic and cytoprotective properties. Mol. Pharmacol. 66, 773–779 (2004).
Sedlak, T. W. et al. Bilirubin and glutathione have complementary antioxidant and cytoprotective roles. Proc. Natl Acad. Sci. USA 106, 5171–5176 (2009).
Chassaing, B., Aitken, J. D., Malleshappa, M. & Vijay-Kumar, M. Dextran sulfate sodium (DSS)-induced colitis in mice. Curr. Protoc. Immunol. 104, 15.25.1–15.25.14 (2014).
Hansberry, D. R., Shah, K., Agarwal, P. & Agarwal, N. Fecal Myeloperoxidase as a Biomarker for Inflammatory Bowel Disease. Cureus 9, e1004 (2017).
Hall, E. D., McCall, J. M., Chase, R. L., Yonkers, P. A. & Braughler, J. M. A nonglucocorticoid steroid analog of methylprednisolone duplicates its high-dose pharmacology in models of central nervous system trauma and neuronal membrane damage. J. Pharmacol. Exp. Ther. 242, 137–142 (1987).
Li, B., Alli, R., Vogel, P. & Geiger, T. L. IL-10 modulates DSS-induced colitis through a macrophage-ROS-NO axis. Mucosal Immunol. 7, 869–878 (2014).
Gibson, G. R. et al. Expert consensus document: the international scientific association for probiotics and prebiotics (ISAPP) consensus statement on the definition and scope of prebiotics. Nat. Rev. Gastroenterol. Hepatol. 14, 491–502 (2017).
Ma, C. et al. Gut microbiome-mediated bile acid metabolism regulates liver cancer via NKT cells. Science 360, eaan5931 (2018).
Cani, P. D. & de Vos, W. M. Next-generation beneficial microbes: the case of Akkermansia muciniphila. Front. Microbiol. 8, 1765 (2017).
Everard, A. et al. Cross-talk between Akkermansia muciniphila and intestinal epithelium controls diet-induced obesity. Proc. Natl Acad. Sci. USA 110, 9066–9071 (2013).
Zhang, Z. et al. Chlorogenic acid ameliorates experimental colitis by promoting growth of Akkermansia in mice. Nutrients 9, 677 (2017).
Furusawa, Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504, 446–450 (2013).
Madsen, K. L., Doyle, J. S., Jewell, L. D., Tavernini, M. M. & Fedorak, R. N. Lactobacillus species prevents colitis in interleukin 10 gene-deficient mice. Gastroenterology 116, 1107–1114 (1999).
Galdeano, C. M. & Perdigon, G. The probiotic bacterium Lactobacillus casei induces activation of the gut mucosal immune system through innate immunity. Clin. Vaccine Immunol. 13, 219–226 (2006).
Geier, M. S., Butler, R. N., Giffard, P. M. & Howarth, G. S. Lactobacillus fermentum BR11, a potential new probiotic, alleviates symptoms of colitis induced by dextran sulfate sodium (DSS) in rats. Int. J. Food Microbiol. 114, 267–274 (2007).
Sartor, R. B. Therapeutic manipulation of the enteric microflora in inflammatory bowel diseases: antibiotics, probiotics, and prebiotics. Gastroenterology 126, 1620–1633 (2004).
Guandalini, S. Use of Lactobacillus-GG in paediatric Crohn’s disease. Dig. Liver Dis. 34, S63–S65 (2002). Suppl 2.
Zocco, M. A. et al. Efficacy of Lactobacillus GG in maintaining remission of ulcerative colitis. Aliment Pharmacol. Ther. 23, 1567–1574 (2006).
Oliva, S. et al. Randomised clinical trial: the effectiveness of Lactobacillus reuteri ATCC 55730 rectal enema in children with active distal ulcerative colitis. Aliment Pharmacol. Ther. 35, 327–334 (2012).
Png, C. W. et al. Mucolytic bacteria with increased prevalence in IBD mucosa augment in vitro utilization of mucin by other bacteria. Am. J. Gastroenterol. 105, 2420–2428 (2010).
Fabia, R. et al. Impairment of bacterial flora in human ulcerative colitis and experimental colitis in the rat. Digestion 54, 248–255 (1993).
Rakoff-Nahoum, S., Paglino, J., Eslami-Varzaneh, F., Edberg, S. & Medzhitov, R. Recognition of commensal microflora by toll-like receptors is required for intestinal homeostasis. Cell 118, 229–241 (2004).
Lee, Y. et al. Bilirubin nanoparticles as a nanomedicine for anti-inflammation therapy. Angew Chem. Int. Ed. Engl. 55, 7460–7463 (2016).
Kim, D. E. et al. Bilirubin nanoparticles ameliorate allergic lung inflammation in a mouse model of asthma. Biomaterials 140, 37–44 (2017).
Lee, S., Lee, Y., Kim, H., Lee, D. Y. & Jon, S. Bilirubin nanoparticle-assisted delivery of a small molecule-drug conjugate for targeted cancer therapy. Biomacromolecules 19, 2270–2277 (2018).
Cho, I. & Blaser, M. J. The human microbiome: at the interface of health and disease. Nat. Rev. Genet. 13, 260–270 (2012).
Lynch, S. V. & Pedersen, O. The human intestinal microbiome in health and disease. N. Engl. J. Med. 375, 2369–2379 (2016).
Pietroiusti, A., Magrini, A. & Campagnolo, L. New frontiers in nanotoxicology: gut microbiota/microbiome-mediated effects of engineered nanomaterials. Toxicol. Appl. Pharmacol. 299, 90–95 (2016).
Javurek, A. B. et al. Gut dysbiosis and neurobehavioral alterations in rats exposed to silver nanoparticles. Sci. Rep. 7, 2822 (2017).
Qiu, K., Durham, P. G. & Anselmo, A. C. Inorganic nanoparticles and the microbiome. Nano Res. 11, 4936–4954 (2018).
Mosquera, M. J. et al. Immunomodulatory nanogels overcome restricted immunity in a murine model of gut microbiome-mediated metabolic syndrome. Sci. Adv. 5, eaav9788 (2019).
Lee, Y., Lee, S. & Jon, S. Biotinylated Bilirubin nanoparticles as a tumor microenvironment-responsive drug delivery system for targeted cancer therapy. Adv. Sci. (Weinh.) 5, 1800017 (2018).
Reissig, J. L., Storminger, J. L. & Leloir, L. F. A modified colorimetric method for the estimation of N-acetylamino sugars. J. Biol. Chem. 217, 959–966 (1955).
McCoy, K. D., Geuking, M. B. & Ronchi, F. Gut microbiome standardization in control and experimental mice. Curr. Protoc. Immunol. 117, 23.1.1–23.1.13 (2017).
Seekatz, A. M. et al. Fecal microbiota transplantation eliminates Clostridium difficile in a murine model of relapsing disease. Infect. Immun. 83, 3838–3846 (2015).
Schloss, P. D. et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75, 7537–7541 (2009).
Kozich, J. J., Westcott, S. L., Baxter, N. T., Highlander, S. K. & Schloss, P. D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl. Environ. Microbiol. 79, 5112–5120 (2013).
Acknowledgements
This work was supported in part by the NIH (grant nos. R01EB022563, R01AI127070, R01CA210273, R01CA223804, U01CA210152, R01DK108901), the MTRAC for Life Sciences Hub and the Emerald Foundation. J.J.M. is a Young Investigator supported by the Melanoma Research Alliance (grant no. 348774), the DoD/CDMRP Peer Reviewed Cancer Research Program (grant no. W81XWH-16-1-0369) and a NSF CAREER Award (no. 1553831). Opinions interpretations, conclusions, and recommendations are those of the author and are not necessarily endorsed by the Department of Defense. The authors thank H. Atsushi for his technical help with LPMC isolation and flow cytometric analysis; the University of Michigan Medical School Host Microbiome Initiative for microbial community analysis; the University of Michigan Cancer Center Immunology Core for ELISA analysis; the ULAM In Vivo Animal Core for tissue sectioning and histological analysis of colon samples; the ULAM Pathology Core for blood analysis; and the College of Pharmacy Biochemical NMR Core at the University of Michigan.
Author information
Authors and Affiliations
Contributions
Y.L. and J.M. designed the experiments. Y.L. performed all experiments. K.S. and N.K. contributed technical expertise, including qPCR analysis, LPMC isolation and flow cytometry analysis. Y.L. and J.M. analysed the data. M.G. aided with interpretation of data on gut microbiome analysis. S.J. contributed the initial design of bilirubin conjugates. Y.L. and J.M. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–22
Rights and permissions
About this article
Cite this article
Lee, Y., Sugihara, K., Gillilland, M.G. et al. Hyaluronic acid–bilirubin nanomedicine for targeted modulation of dysregulated intestinal barrier, microbiome and immune responses in colitis. Nat. Mater. 19, 118–126 (2020). https://doi.org/10.1038/s41563-019-0462-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41563-019-0462-9
This article is cited by
-
Attenuation of inflammatory bowel disease by oral administration of mucoadhesive polydopamine-coated yeast β-glucan via ROS scavenging and gut microbiota regulation
Journal of Nanobiotechnology (2024)
-
The therapeutic potential of immunoengineering for systemic autoimmunity
Nature Reviews Rheumatology (2024)
-
Effects of radiofrequency field from 5G communication on fecal microbiome and metabolome profiles in mice
Scientific Reports (2024)
-
ROS-responsive nanoparticles targeting inflamed colon for synergistic therapy of inflammatory bowel disease via barrier repair and anti-inflammation
Nano Research (2024)
-
Reactive oxygen species (ROS) scavenging biomaterials for anti-inflammatory diseases: from mechanism to therapy
Journal of Hematology & Oncology (2023)