Abstract
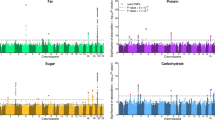
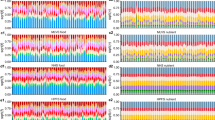
Dietary intake is a major contributor to the global obesity epidemic and represents a complex behavioural phenotype that is partially affected by innate biological differences. Here, we present a multivariate genome-wide association analysis of overall variation in dietary intake to account for the correlation between dietary carbohydrate, fat and protein in 282,271 participants of European ancestry from the UK Biobank (n = 191,157) and Cohorts for Heart and Aging Research in Genomic Epidemiology Consortium (n = 91,114), and identify 26 distinct genome-wide significant loci. Dietary intake signals map exclusively to specific brain regions and are enriched for genes expressed in specialized subtypes of GABAergic, dopaminergic and glutamatergic neurons. We identified two main clusters of genetic variants for overall variation in dietary intake that were differently associated with obesity and coronary artery disease. These results enhance the biological understanding of interindividual differences in dietary intake by highlighting neural mechanisms, supporting functional follow-up experiments and possibly providing new avenues for the prevention and treatment of prevalent complex metabolic diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The summary GWAS statistics will be publicly available at the UKBB website (http://biobank.ctsu.ox.ac.uk/), database of Genotypes and Phenotypes (accession no. phs000930) and Type 2 Diabetes Knowledge Portal (http://www.kp4cd.org/dataset_downloads/t2d). The single-cell expression datasets can be found at https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc= under accession nos. GSE93374, GSE104276 and GSE763816,46,65. The ldsc command line tool can be found at https://github.com/bulik/ldsc20.
Code availability
The code used to reproduce the analyses for this manuscript will be made available on publication at http://sites.bu.edu/fhspl/publications/ and https://github.com/perslab/Merino_2020. The CELLEX precomputed expression specificity files are available at https://github.com/perslab/CELLECT/wiki/Precomputed-CELLEX-datasets.
References
Atasoy, D., Betley, J. N., Su, H. H. & Sternson, S. M. Deconstruction of a neural circuit for hunger. Nature 488, 172–177 (2012).
van der Klaauw, A. A. & Farooqi, I. S. The hunger genes: pathways to obesity. Cell 161, 119–132 (2015).
Andermann, M. L. & Lowell, B. B. Toward a wiring diagram understanding of appetite control. Neuron 95, 757–778 (2017).
Polderman, T. J. C. et al. Meta-analysis of the heritability of human traits based on fifty years of twin studies. Nat. Genet. 47, 702–709 (2015).
Merino, J. et al. Genome-wide meta-analysis of macronutrient intake of 91,114 European ancestry participants from the cohorts for heart and aging research in genomic epidemiology consortium. Mol. Psychiatry 24, 1920–1932 (2019).
Campbell, J. N. et al. A molecular census of arcuate hypothalamus and median eminence cell types. Nat. Neurosci. 20, 484–496 (2017).
Livneh, Y. et al. Homeostatic circuits selectively gate food cue responses in insular cortex. Nature 546, 611–616 (2017).
Farooqi, I. S. et al. Leptin regulates striatal regions and human eating behavior. Science 317, 1355 (2007).
Lowell, B. B. New neuroscience of homeostasis and drives for food, water, and salt. N. Engl. J. Med. 380, 459–471 (2019).
Gaich, G. et al. The effects of LY2405319, an FGF21 analog, in obese human subjects with type 2 diabetes. Cell Metab. 18, 333–340 (2013).
Søberg, S. et al. FGF21 is a sugar-induced hormone associated with sweet intake and preference in humans. Cell Metab. 25, 1045–1053.e6 (2017).
Chu, A. Y. et al. Novel locus including FGF21 is associated with dietary macronutrient intake. Hum. Mol. Genet. 22, 1895–1902 (2013).
Zhong, V. W. et al. A genome-wide association study of bitter and sweet beverage consumption. Hum. Mol. Genet. 28, 2449–2457 (2019).
Meddens, S. F. W. et al. Genomic analysis of diet composition finds novel loci and associations with health and lifestyle. Mol. Psychiatry https://doi.org/10.1038/s41380-020-0697-5 (2020).
Vroom, C.-R., de Leeuw, C., Posthuma, D., Dolan, C. V. & van der Sluis, S. The more the merrier? Multivariate approaches to genome-wide association analysis. Preprint at bioRxiv https://doi.org/10.1101/610287 (2019).
Lane, J. M. et al. Genome-wide association analyses of sleep disturbance traits identify new loci and highlight shared genetics with neuropsychiatric and metabolic traits. Nat. Genet. 49, 274–281 (2017).
Turley, P. et al. Multi-trait analysis of genome-wide association summary statistics using MTAG. Nat. Genet. 50, 229–237 (2018).
Sudlow, C. et al. UK Biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
Zhu, X. et al. Meta-analysis of correlated traits via summary statistics from GWASs with an application in hypertension. Am. J. Hum. Genet. 96, 21–36 (2015).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Liu, M. et al. Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat. Genet. 51, 237–244 (2019).
Nagel, M., Watanabe, K., Stringer, S., Posthuma, D. & van der Sluis, S. Item-level analyses reveal genetic heterogeneity in neuroticism. Nat. Commun. 9, 905 (2018).
Karlsson Linnér, R. et al. Genome-wide association analyses of risk tolerance and risky behaviors in over 1 million individuals identify hundreds of loci and shared genetic influences. Nat. Genet. 51, 245–257 (2019).
Choi, K. W. et al. Assessment of bidirectional relationships between physical activity and depression among adults: a 2-sample Mendelian randomization study. JAMA Psychiatry 76, 399–408 (2019).
Choi, K. W. et al. An exposure-wide and Mendelian randomization approach to identifying modifiable factors for the prevention of depression. Am. J. Psychiatry 177, 944–954 (2020).
Ardlie, K. G. et al. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Ramasamy, A. et al. Genetic variability in the regulation of gene expression in ten regions of the human brain. Nat. Neurosci. 17, 1418–1428 (2014).
Jaffe, A. E. et al. Developmental and genetic regulation of the human cortex transcriptome illuminate schizophrenia pathogenesis. Nat. Neurosci. 21, 1117–1125 (2018).
Schumann, G. et al. KLB is associated with alcohol drinking, and its gene product β-Klotho is necessary for FGF21 regulation of alcohol preference. Proc. Natl Acad. Sci. USA 113, 14372–14377 (2016).
Liu, Z. et al. Association of corticotropin-releasing hormone receptor1 gene SNP and haplotype with major depression. Neurosci. Lett. 404, 358–362 (2006).
Schaum, N. et al. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562, 367–372 (2018).
Zeisel, A. et al. Molecular architecture of the mouse nervous system. Cell 174, 999–1014.e22 (2018).
Tan, V. Y. F. & Févotte, C. Automatic relevance determination in nonnegative matrix factorization with the β-divergence. IEEE Trans. Pattern Anal. Mach. Intell. 35, 1592–1605 (2013).
Locke, A. E. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Scott, R. A. et al. An expanded genome-wide association study of type 2 diabetes in Europeans. Diabetes 66, 2888–2902 (2017).
Nikpay, M. et al. A comprehensive 1000 Genomes-based genome-wide association meta-analysis of coronary artery disease. Nat. Genet. 47, 1121–1130 (2015).
Karlson, E. W., Boutin, N. T., Hoffnagle, A. G. & Allen, N. L. Building the Partners HealthCare Biobank at Partners Personalized Medicine: informed consent, return of research results, recruitment lessons and operational considerations. J. Pers. Med. 6, 2 (2016).
Blouet, C. & Schwartz, G. J. Brainstem nutrient sensing in the nucleus of the solitary tract inhibits feeding. Cell Metab. 16, 579–587 (2012).
Hayes, M. R. et al. Endogenous leptin signaling in the caudal nucleus tractus solitarius and area postrema is required for energy balance regulation. Cell Metab. 23, 744 (2016).
D’Agostino, G. et al. Nucleus of the solitary tract serotonin 5-HT2C receptors modulate food intake. Cell Metab. 28, 619–630.e5 (2018).
Holt, M. K. et al. Preproglucagon neurons in the nucleus of the solitary tract are the main source of brain GLP-1, mediate stress-induced hypophagia, and limit unusually large intakes of food. Diabetes 68, 21–33 (2019).
Timshel, P. N., Thompson, J. J. & Pers, T. H. Genetic mapping of etiologic brain cell types for obesity. eLife 9, e55851 (2020).
Shai, I. et al. Weight loss with a low-carbohydrate, Mediterranean, or low-fat diet. N. Engl. J. Med. 359, 229–241 (2008).
Ludwig, D. S., Willett, W. C., Volek, J. S. & Neuhouser, M. L. Dietary fat: from foe to friend? Science 362, 764–770 (2018).
Gibson, E. L. Emotional influences on food choice: sensory, physiological and psychological pathways. Physiol. Behav. 89, 53–61 (2006).
La Manno, G. et al. Molecular diversity of midbrain development in mouse, human, and stem cells. Cell 167, 566–580.e19 (2016).
Liu, B. et al. Development and evaluation of the Oxford WebQ, a low-cost, web-based method for assessment of previous 24 h dietary intakes in large-scale prospective studies. Public Health Nutr. 14, 1998–2005 (2011).
Bradbury, K. E., Young, H. J., Guo, W. & Key, T. J. Dietary assessment in UK Biobank: an evaluation of the performance of the touchscreen dietary questionnaire. J. Nutr. Sci. 7, e6 (2018).
Greenwood, D. C. et al. Validation of the Oxford WebQ online 24-hour dietary questionnaire using biomarkers. Am. J. Epidemiol. 188, 1858–1867 (2019).
McCance, R. A. & Widdowson, E. M. McCance and Widdowson’s the Composition of Foods (Royal Society of Chemistry, 2002).
Mifflin, M. D. et al. A new predictive equation for resting energy expenditure in healthy individuals. Am. J. Clin. Nutr. 51, 241–247 (1990).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Li, Y., Willer, C. J., Ding, J., Scheet, P. & Abecasis, G. R. MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet. Epidemiol. 34, 816–834 (2010).
Howie, B., Fuchsberger, C., Stephens, M., Marchini, J. & Abecasis, G. R. Fast and accurate genotype imputation in genome-wide association studies through pre-phasing. Nat. Genet. 44, 955–959 (2012).
Zhou, W. et al. Efficiently controlling for case-control imbalance and sample relatedness in large-scale genetic association studies. Nat. Genet. 50, 1335–1341 (2018).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Higgins, J. P. T. & Thompson, S. G. Quantifying heterogeneity in a meta-analysis. Stat. Med. 21, 1539–1558 (2002).
Watanabe, K., Taskesen, E., van Bochoven, A. & Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 8, 1826 (2017).
Yang, H. & Wang, K. Genomic variant annotation and prioritization with ANNOVAR and wANNOVAR. Nat. Protoc. 10, 1556–1566 (2015).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Boyle, A. P. et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 22, 1790–1797 (2012).
Ernst, J. & Kellis, M. ChromHMM: automating chromatin-state discovery and characterization. Nat. Methods 9, 215–216 (2012).
Kundaje, A. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
de Leeuw, C. A., Mooij, J. M., Heskes, T. & Posthuma, D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput. Biol. 11, e1004219 (2015).
Zhong, S. et al. A single-cell RNA-seq survey of the developmental landscape of the human prefrontal cortex. Nature 555, 524–528 (2018).
Hodge, R. D. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019).
Wang, D. et al. Comprehensive functional genomic resource and integrative model for the human brain. Science 362, eaat8464 (2018).
Eastwood, S. V. et al. Algorithms for the capture and adjudication of prevalent and incident diabetes in UK Biobank. PLoS ONE 11, e0162388 (2016).
Acknowledgements
This study was designed and carried out by the CHARGE Consortium Nutrition Working Group. Part of this work was conducted using the UKBB resource under application no. 27892. This project has received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Sklodowska-Curie grant agreement No 703787. J.M. is supported by the National Institutes of Health (NIH) grant no. P30 DK040561. H.S.D. and R.S. are supported by NIH grant nos. R01 DK105072 and DK107859. R.S. is supported by NIH grant no. R01 DK102696 and the MGH Research Scholar Fund. J.M.L. is supported by NIH grant nos. F32 DK102323 and T32 HL007901. C.S., J.C.F. and J.D. are supported by NIH grant no. U01 DK078616. J.C.F. is supported by NIH grant no. K24 DK110550. J.C. is supported by the American Diabetes Association Pathway to Stop Diabetes award no. 1-18-INI-14. T.H.P. and P.V.T. acknowledge the Novo Nordisk Foundation (no. NNF16OC0021496). T.H.P. acknowledges the Lundbeck Foundation (no. R19020143904). This research was supported in part by the Intramural Research Program of the National Institute on Aging. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
J.M., H.S.D., C.S., A.Y.C., D.I.C., J.C.F. and R.S. conceived and designed the study. J.D., J.C.F. and R.S. oversaw the study. H.S.D., J.M., C.S., J.M.L. and Y.S. served as analysts. Phenotype definitions were developed by J.M., H.S.D., C.S., J.D. and R.S. C.S. performed the quality control and meta-analyses. Heritability and genetic correlation were performed by H.S.D. The bioinformatic analyses were performed and interpreted by J.M., H.S.D., C.S., J.M.L., Y.S., H.W., J.K., C.T., T.T., D.I.C., J.C.F. and R.S. The single-cell expression analyses were conducted and interpreted by J.M., P.V.T., T.H.P., J.C., L.T. and J.C.F. The figures were created by J.M., H.S.D., C.S., M.S.U., Y.S., P.V.T., T.H.P., J.D. and D.I.C. M.K.R. provided helpful advice and feedback on study design and manuscript writing. J.M., H.S.D., C.S., J.D., J.C.F. and R.S., made major contributions to manuscript writing and editing. All authors contributed to and critically reviewed the manuscript and approved its final version.
Corresponding authors
Ethics declarations
Competing interests
A.Y.C. is currently employed by Merck Research Laboratories. M.K.R. reports receiving research funding from Novo Nordisk, consultancy fees from Novo Nordisk and Roche Diabetes Care and modest owning of shares in GlaxoSmithKline. All other authors declare no competing interests.
Additional information
Peer review information Nature Human Behaviour thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 UK Biobank sample selection.
A total of 502,536 participants were available in UK Biobank at the beginning of this study. Thirty participants withdrew consent during the implementation of the analysis plan. We excluded 892 participants with invalid diet data based on previously defined quality control filtering criteria for diet data. A total of 16,034 participants did not pass quality control criteria based on UK Biobank quality control definitions (high heterozygosity & high missing rate, sex aneuploidy, submitted sex different from inferred sex). We excluded 27,965 participants based on a non-European self-reported ancestry, and 5,955 European ancestry outliers based on + /-6SD from the mean in the subset of 192,025 participants based on the first 4 PCs. Among 451,660 participants remaining for the discovery of genetic variants for dietary intake, 260,503 had missing diet data (n = 258,393) or invalid values (n = 2,110). Due to the skewed distribution of macronutrient intake, we winsorized at mean + /− 5 SD for each phenotype.
Extended Data Fig. 2 Schematic of the study design in the multi-trait genome-wide association meta-analysis for dietary intake in 282,271 individuals.
The genome-wide association meta-analysis of dietary intake comprised data from 191,157 participants from the UK Biobank and 91,114 participants from the CHARGE Consortium. Single-trait macronutrient GWAS from the UK Biobank and CHARGE Consortium were meta-analyzed using METAL and then combined into a multi-trait GWAS using the multi-trait CPASSOC method. Downstream in silico analyses were conducted to identify biological features of identified loci including functional annotation, tissue and pathway enrichment, and single-cell RNA expression analyses. The Bayesian nonnegative matrix factorization clustering algorithm was used to classify dietary intake genetic loci into subgroups based on potential functional and clinical similarities. Cluster-based polygenic risk scores were built to investigate patterns of metabolic risk.
Extended Data Fig. 3 Quantile-quantile plot of the SNP-based associations with single-trait and multi-trait genome-wide association meta-analyses of 282,271 individuals.
Quantile–quantile plot of the SNP-based associations with multivariate (a), carbohydrate (b), fat (c), and protein intake (d). SNP P values were computed in METAL by weighting effect size estimates using the inverse of the corresponding standard errors.
Extended Data Fig. 4 Associations between environmental factors and macronutrient intake.
Shown are effect estimates (betas) and 95% confidence intervals for the association between environmental factors and macronutrient intake among UK Biobank participants. Environmental factors were all added to the initial model used for the primary main UK Biobank analyses. A null model was first run using SAIGE to evaluate the association of each environmental factor with each macronutrient intake. Only significant factors were retained in the model. Genetic association analyses were then conducted, similarly to the primary main UK Biobank analyses, using SAIGE.
Extended Data Fig. 5 Schematic overview of the Bayesian nonnegative matrix factorization clustering algorithm.
The input for the Bayesian nonnegative matrix factorization clustering algorithm (bNMF) was the set of 31 genetic variants reaching nominal significance association with proportion fat intake. Next summary association statistics for 22 dietary intake traits from the UK Biobank were aggregated for each dietary intake variant. Our analyses involved variants aligned by their alleles associated with increased fat intake. We generated standardized effect sizes for variant trait associations from GWAS by dividing the estimated regression coefficient beta by the standard error, using the UK Biobank summary statistic results (variant-trait association matrix (31 by 22)). The defining features of each cluster were determined by the most highly associated traits, which is a natural output of the bNMF approach. bNMF algorithm was performed in R for 1,000 iterations with different initial conditions, and the maximum posterior solution at the most probable number of clusters was selected for downstream analysis.
Extended Data Fig. 6 Trait and loci association to clusters.
Clustering of variant-trait associations was performed for 31 genetic variants reaching nominal significance association with proportion fat intake and 22 nutritional traits derived from GWAS using the Bayesian nonnegative matrix factorization clustering algorithm, with identification of two clusters present on 80% of iterations. Loci and traits defining each cluster were based on a cut-off of weighting of 0.94 (Methods). a) trait association to cluster, b) loci association to cluster: 1 NEGR1, RARB, RP11.161I6.2, RP11.22P4.1, TMEM108. Cluster 2: ADH1A, DNMT3A, FGF21, FUT1, FUT2, GCKR, KLB, PLEKHM1, PPP1R3B, RP11.696N14.1, TSPAN5.
Supplementary information
Supplementary Information
Bayesian non-negative matrix factorization.
Rights and permissions
About this article
Cite this article
Merino, J., Dashti, H.S., Sarnowski, C. et al. Genetic analysis of dietary intake identifies new loci and functional links with metabolic traits. Nat Hum Behav 6, 155–163 (2022). https://doi.org/10.1038/s41562-021-01182-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41562-021-01182-w
This article is cited by
-
Genetic predisposition to macronutrient preference and workplace food choices
Molecular Psychiatry (2023)
-
High sucrose consumption decouples intrinsic and synaptic excitability of AgRP neurons without altering body weight
International Journal of Obesity (2023)
-
Large-scale GWAS of food liking reveals genetic determinants and genetic correlations with distinct neurophysiological traits
Nature Communications (2022)
-
Bidirectional two-sample Mendelian randomization analysis identifies causal associations between relative carbohydrate intake and depression
Nature Human Behaviour (2022)
-
Genetically regulated multi-omics study for symptom clusters of posttraumatic stress disorder highlights pleiotropy with hematologic and cardio-metabolic traits
Molecular Psychiatry (2022)